Genome-Wide Association Study for Ultraviolet-B Resistance in Soybean (Glycine max L.)
Abstract
1. Introduction
2. Results
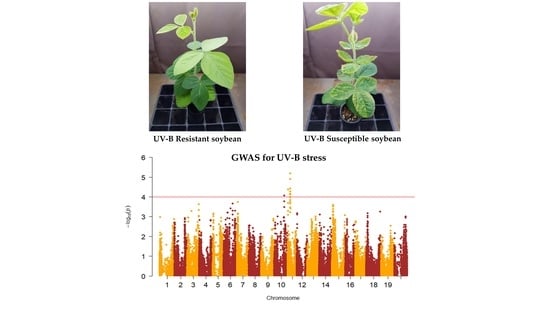
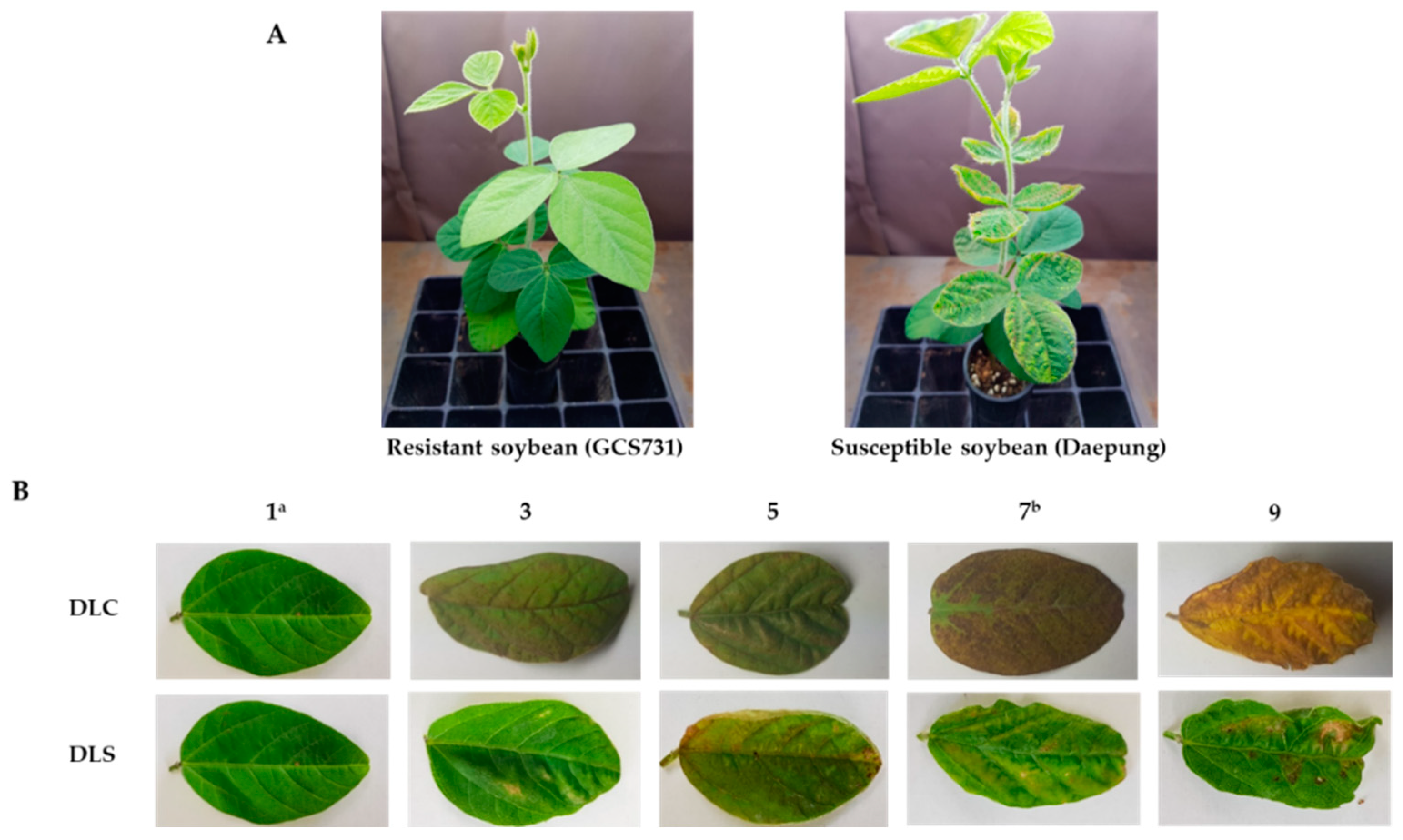
2.1. Phenotypic Evaluation of Soybean Germplasm Collection under Enhanced UV-B Conditions
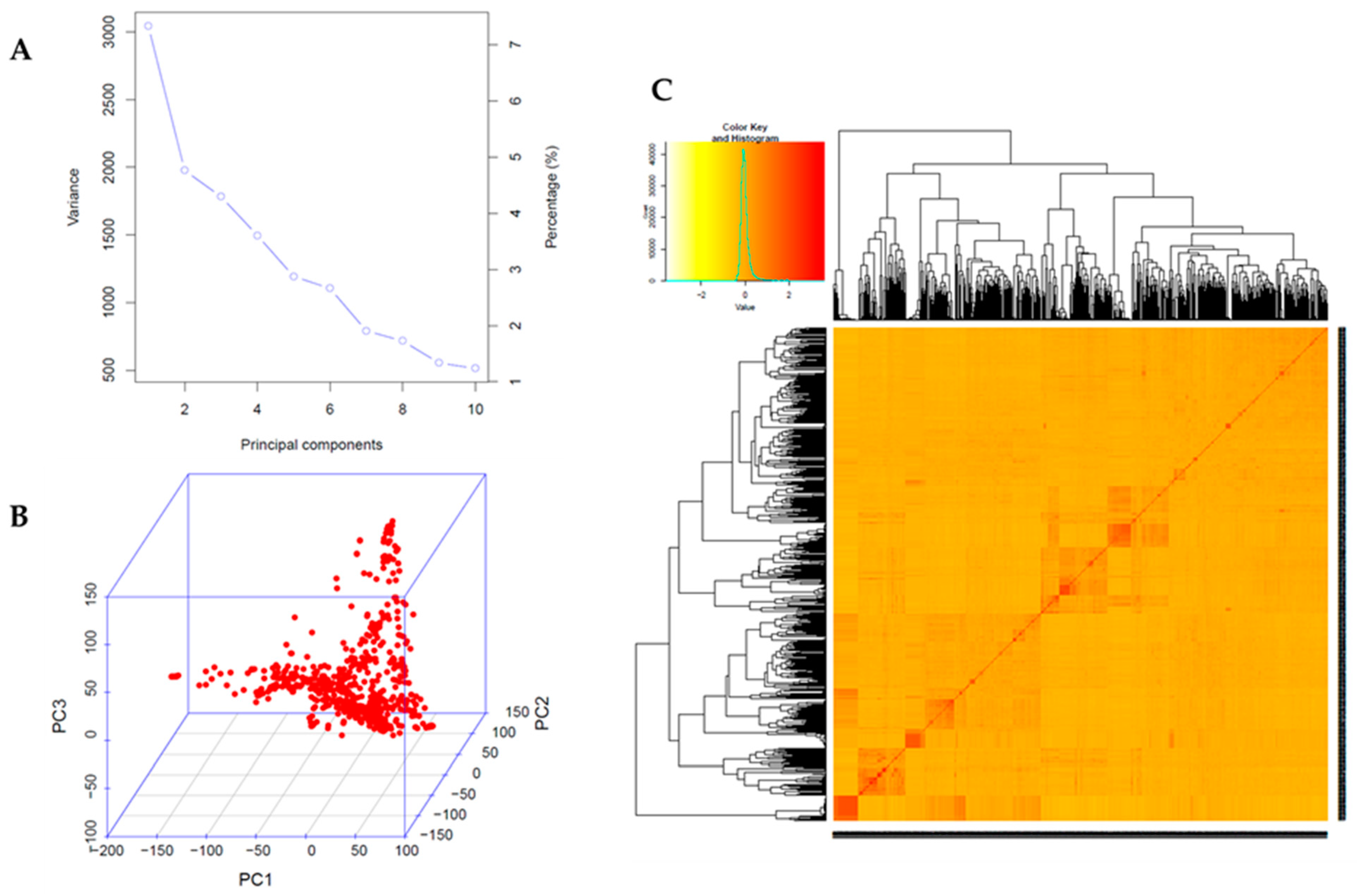
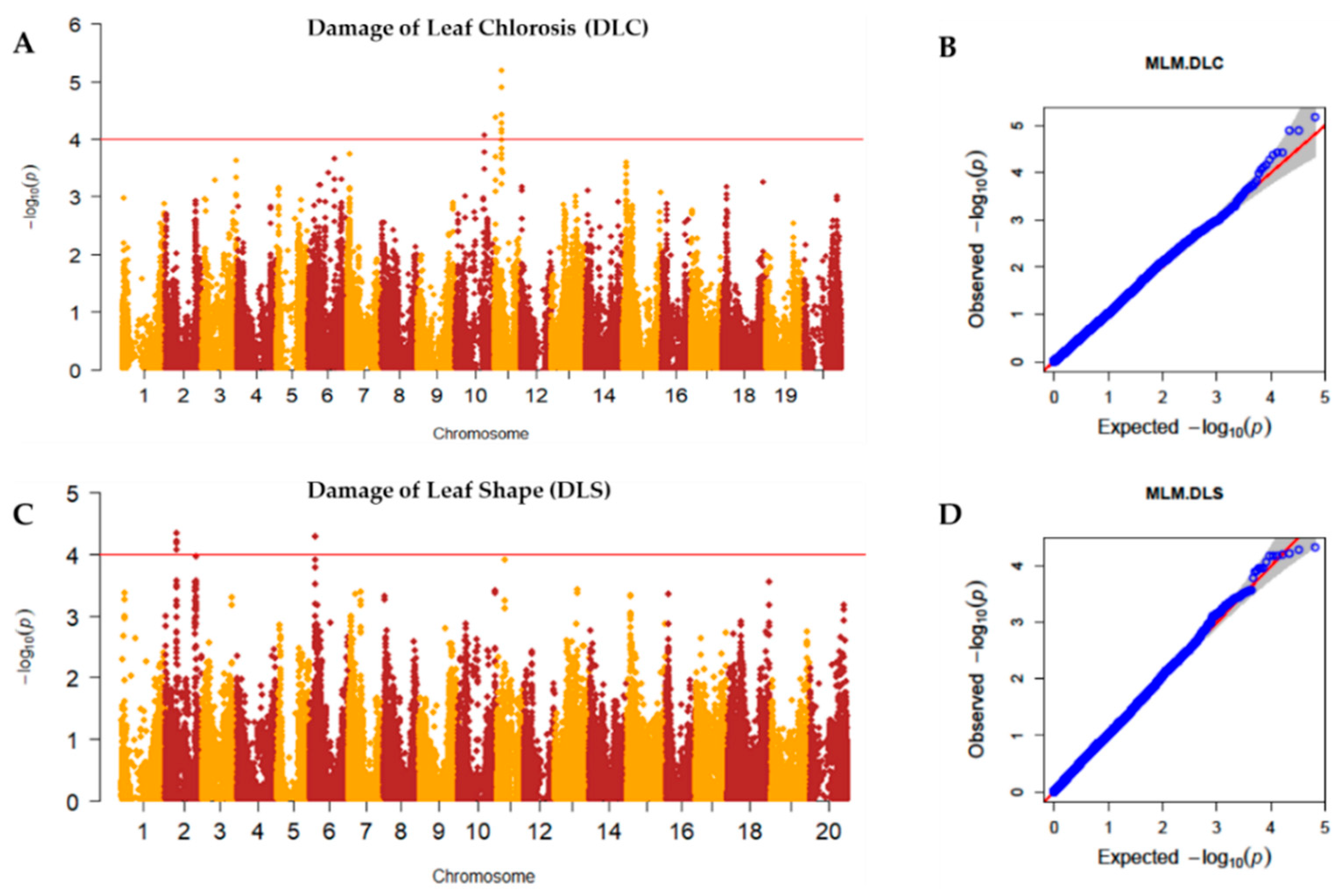
2.2. Genome-Wide Association Study (GWAS)
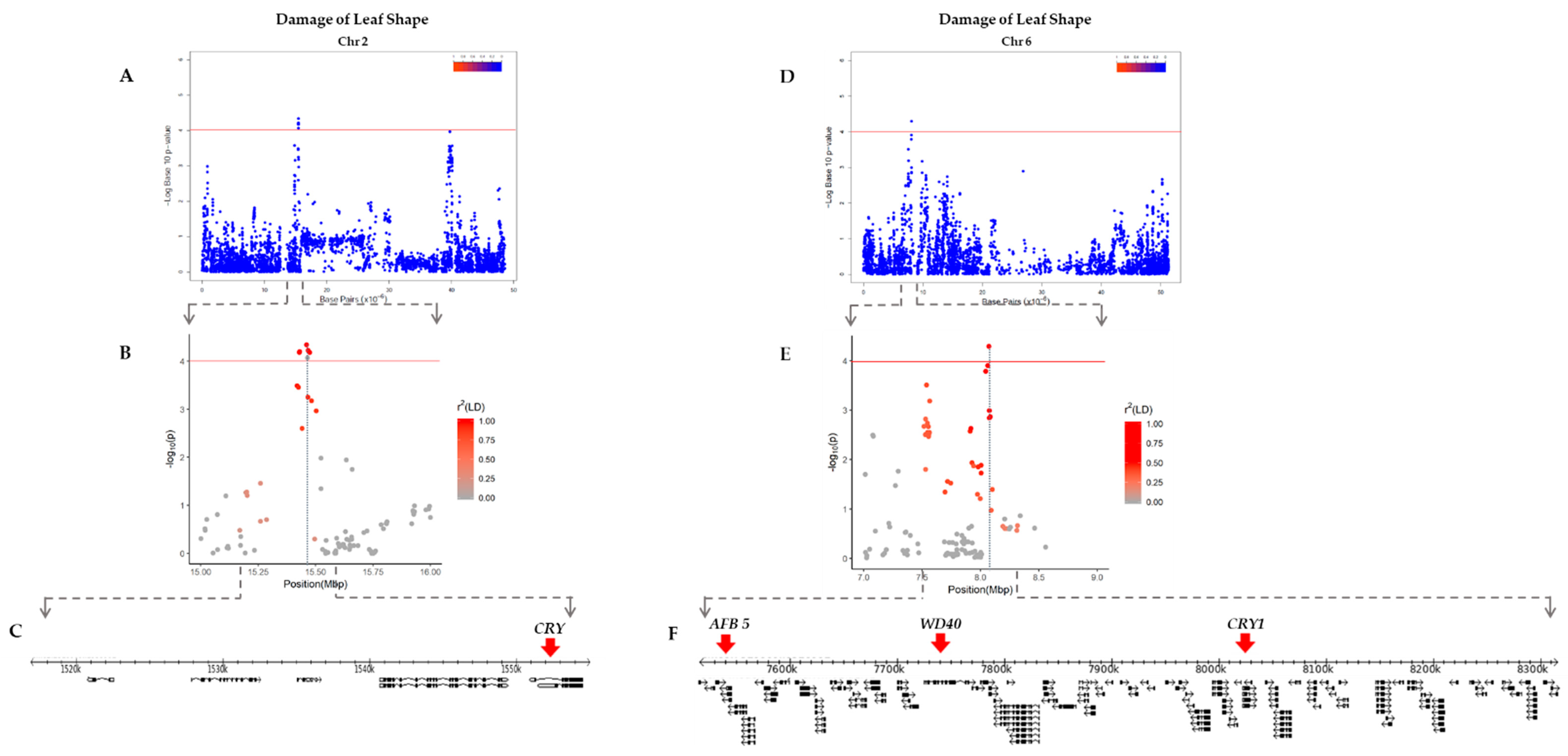
2.3. Linkage Disequilibrium (LD) Analysis
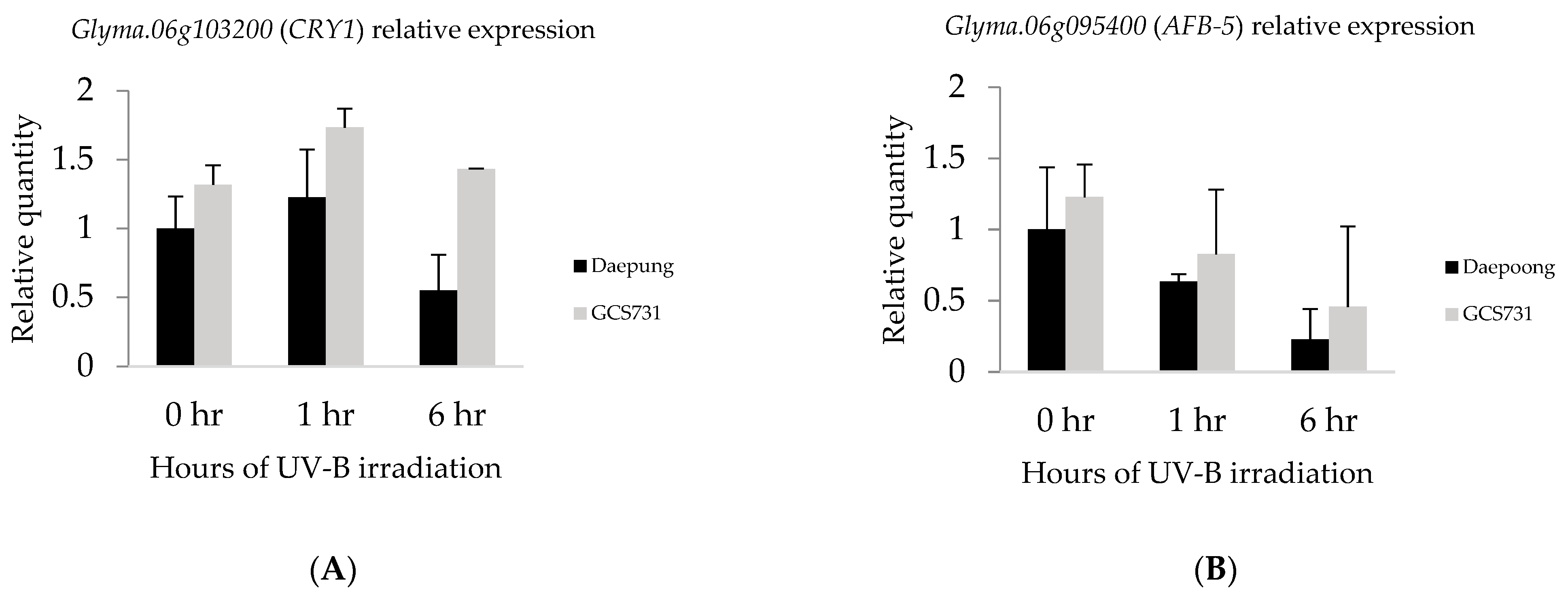
2.4. Gene Expression Analysis by qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Evaluation of UV-B Resistance
4.3. Genome-Wide Association Study (GWAS)
4.4. Linkage Disequilibrium (LD) Analysis and Candidate Gene Identification
4.5. Real-Time Quantitative Reverse Transcription PCR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Montzka, S.A.; Dutton, G.; Yu, P.; Ray, E.; Portmann, R.; Daniel, J.S.; Kuijpers, L.; Hall, B.D.; Mondeel, D.; Siso, C.; et al. An unexpected and persistent increase in global emissions of ozone-depleting CFC-11. Nat. Cell Biol. 2018, 557, 413–417. [Google Scholar] [CrossRef]
- Hossaini, R.; Chipperfield, M.; Montzka, S.A.; Leeson, A.A.; Dhomse, S.S.; Pyle, J.A. The increasing threat to stratospheric ozone from dichloromethane. Nat. Commun. 2017, 8, 15962. [Google Scholar] [CrossRef]
- Hollósy, F. Effects of ultraviolet radiation on plant cells. Micron 2002, 33, 179–197. [Google Scholar] [CrossRef]
- Deckmyn, G.; Impens, I. Combined effects of enhanced UV-B radiation and nitrogen deficiency on the growth, composition and photosynthesis of rye (Secale cereale). Plant Ecol. 1997, 128, 235–240. [Google Scholar] [CrossRef]
- Pang, Q.; Hays, J.B. UV-B-Inducible and Temperature-Sensitive Photoreactivation of Cyclobutane Pyrimidine Dimers in Arabidopsis thaliana. Plant Physiol. 1991, 95, 536–543. [Google Scholar] [CrossRef]
- Mitchell, D.L.; Nairn, R.S. The Biology of the (6–4) Photoproduct. Photochem. Photobiol. 1989, 49, 805–819. [Google Scholar] [CrossRef]
- Teramura, A.H. Effects of Ultraviolet-B Irradiances on Soybean: II. Interaction between Ultraviolet-B and Photosynthetically Active Radiation on Net Photosynthesis, Dark Respiration, and Transpiration. Plant Physiol. 1980, 65, 483–488. [Google Scholar] [CrossRef] [PubMed]
- Hectors, K.; Van Oevelen, S.; Guisez, Y.; Prinsen, E.; Jansen, M.A.K. The phytohormone auxin is a component of the regulatory system that controls UV-mediated accumulation of flavonoids and UV-induced morphogenesis. Physiol. Plant 2012, 145, 594–603. [Google Scholar] [CrossRef]
- Chow, W.S.; Anderson, J.M. UV-B damage and protection at the molecular level in plants. Photosynth. Res. 1994, 39, 475–489. [Google Scholar] [CrossRef]
- Kalbin, G.; Ohlsson, A.B.; Berglund, T.; Rydstrom, J.; Strid, A. Ultraviolet-B-radiation-induced changes in nicotinamide and glutathione metabolism and gene expression in plants. Eur. J. Biochem. 1997, 249, 465–472. [Google Scholar] [CrossRef]
- Hidema, J.; Taguchi, T.; Ono, T.; Teranishi, M.; Yamamoto, K.; Kumagai, T. Increase in CPD photolyase activity functions effectively to prevent growth inhibition caused by UVB radiation. Plant J. 2007, 50, 70–79. [Google Scholar] [CrossRef]
- Takahashi, M.; Teranishi, M.; Ishida, H.; Kawasaki, J.; Takeuchi, A.; Yamaya, T.; Watanabe, M.; Makino, A.; Hidema, J. Cyclobutane pyrimidine dimer (CPD) photolyase repairs ultraviolet-B-induced CPDs in rice chloroplast and mitochondrial DNA. Plant J. 2011, 66, 433–442. [Google Scholar] [CrossRef]
- Sancar, A. Structure and function of DNA photolyase. Biochemistry 1994, 33, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Biever, J.J.; Gardner, G. The relationship between multiple UV-B perception mechanisms and DNA repair pathways in plants. Environ. Exp. Bot. 2016, 124, 89–99. [Google Scholar] [CrossRef]
- Biedermann, S.S.M.; Hellmann, H. Recognition and Repair Pathways of Damaged DNA in Higher Plants; IntechOpen: Rijeka, Croatia, 2011; pp. 201–236. [Google Scholar]
- Favory, J.-J.; Stec, A.; Gruber, H.; Rizzini, L.; Oravecz, A.; Funk, M.; Albert, A.; Cloix, C.; Jenkins, I.G.; Oakeley, E.J.; et al. Interaction of COP1 and UVR8 regulates UV-B-induced photomorphogenesis and stress acclimation in Arabidopsis. EMBO J. 2009, 28, 591–601. [Google Scholar] [CrossRef] [PubMed]
- Hayes, S.; Velanis, C.; Jenkins, G.I.; Franklin, K.A. UV-B detected by the UVR8 photoreceptor antagonizes auxin signaling and plant shade avoidance. Proc. Natl. Acad. Sci. USA 2014, 111, 11894–11899. [Google Scholar] [CrossRef]
- Hofmann, N.R. The Molecular Mechanism of the UVR8 UV-B Photoreceptor. Plant Cell 2012, 24, 3485. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Barnes, P.W.; Flint, S.D.; Caldwell, M.M. Morphological Responses of Crop and Weed Species of Different Growth Forms to Ultraviolet-B Radiation. Am. J. Bot. 1990, 77, 1354–1360. [Google Scholar] [CrossRef]
- Feng, H.; An, L.; Chen, T.; Qiang, W.; Xu, S.; Zhang, M.; Wang, X.; Cheng, G. The effect of enhanced ultraviolet-B radiation on growth, photosynthesis and stable carbon isotope composition (δ13C) of two soybean cultivars (Glycine max) under field conditions. Environ. Exp. Bot. 2003, 49, 1–8. [Google Scholar] [CrossRef]
- Liu, B.; Liu, X.-B.; Li, Y.-S.; Herbert, S. Effects of enhanced UV-B radiation on seed growth characteristics and yield components in soybean. Field Crops Res. 2013, 154, 158–163. [Google Scholar] [CrossRef]
- Kim, E.H.; Seguin, P.; Lee, J.E.; Yoon, C.G.; Song, H.-K.; Ahn, J.-K.; Chung, I.-M. Elevated Ultraviolet-B Radiation Reduces Concentrations of Isoflavones and Phenolic Compounds in Soybean Seeds. J. Agron. Crop Sci. 2011, 197, 75–80. [Google Scholar] [CrossRef]
- Shim, H.-C.; Ha, B.-K.; Yoo, M.; Kang, S.-T. Detection of quantitative trait loci controlling UV-B resistance in soybean. Euphytica 2014, 202, 109–118. [Google Scholar] [CrossRef]
- Lee, J.S.; Kim, S.; Ha, B.-K.; Kang, S. Positional mapping and identification of novel quantitative trait locus responsible for UV-B radiation tolerance in soybean [Glycine max (L.) Merr.]. Mol. Breed. 2016, 36. [Google Scholar] [CrossRef]
- Kim, K.D.; Yun, M.Y.; Shin, J.H.; Kang, Y.J.; Kim, M.Y.; Lee, S.-H. Underlying genetic variation in the response of cultivated and wild soybean to enhanced ultraviolet-B radiation. Euphytica 2014, 202, 207–217. [Google Scholar] [CrossRef]
- Yoon, M.Y.; Kim, M.Y.; Ha, J.; Lee, T.; Kim, K.D.; Lee, S.-H. QTL Analysis of Resistance to High-Intensity UV-B Irradiation in Soybean (Glycine max [L.] Merr.). Int. J. Mol. Sci. 2019, 20, 3287. [Google Scholar] [CrossRef]
- Ersoz, E.S.; Yu, J.; Buckler, E.S. Applications of Linkage Disequilibrium and Association Mapping in Crop Plants. In Genomics-Assisted Crop Improvement; Springer: Dordrecht, The Netherlands, 2007; Volume 1, pp. 97–119. [Google Scholar]
- Shim, S.; Ha, J.; Kim, M.Y.; Choi, M.S.; Kang, S.-T.; Jeong, S.-C.; Moon, J.-K.; Lee, S.-H. GmBRC1 is a Candidate Gene for Branching in Soybean (Glycine max (L.) Merrill). Int. J. Mol. Sci. 2019, 20, 135. [Google Scholar] [CrossRef]
- Jing, Y.; Zhao, X.; Wang, J.; Lian, M.; Teng, W.; Qiu, L.; Han, Y.; Li, W. Identification of loci and candidate genes for plant height in soybean (Glycine max) via genome-wide association study. Plant Breed. 2019, 138, 721–732. [Google Scholar] [CrossRef]
- Wang, L.; Yang, Y.; Zhang, S.; Che, Z.; Yuan, W.; Yu, D. GWAS reveals two novel loci for photosynthesis-related traits in soybean. Mol. Genet. Genom. 2020, 295, 705–716. [Google Scholar] [CrossRef]
- Chang, H.-X.; Hartman, G.L. Characterization of Insect Resistance Loci in the USDA Soybean Germplasm Collection Using Genome-Wide Association Studies. Front. Plant Sci. 2017, 8, 670. [Google Scholar] [CrossRef] [PubMed]
- Che, Z.; Yan, H.; Liu, H.; Yang, H.; Du, H.; Yang, Y.; Liu, B.; Yu, D. Genome-wide association study for soybean mosaic virus SC3 resistance in soybean. Mol. Breed. 2020, 40, 1–14. [Google Scholar] [CrossRef]
- Hyten, D.; Song, Q.; Choi, I.-Y.; Yoon, M.-S.; Specht, J.E.; Matukumalli, L.K.; Nelson, R.L.; Shoemaker, R.C.; Young, N.D.; Cregan, P.B. High-throughput genotyping with the GoldenGate assay in the complex genome of soybean. Theor. Appl. Genet. 2008, 116, 945–952. [Google Scholar] [CrossRef] [PubMed]
- Song, Q.; Hyten, D.L.; Jia, G.; Quigley, C.V.; Fickus, E.W.; Nelson, R.L.; Cregan, P.B. Development and Evaluation of SoySNP50K, a High-Density Genotyping Array for Soybean. PLoS ONE 2013, 8, e54985. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-G.; Jeong, N.; Kim, J.H.; Lee, K.; Kim, K.H.; Pirani, A.; Ha, B.-K.; Kang, S.-T.; Park, B.-S.; Moon, J.-K.; et al. Development, validation and genetic analysis of a large soybean SNP genotyping array. Plant J. 2015, 81, 625–636. [Google Scholar] [CrossRef]
- Tang, Y.; Liu, X.; Wang, J.; Li, M.; Wang, Q.; Tian, F.; Su, Z.; Pan, Y.; Liu, D.; Lipka, A.E.; et al. GAPIT Version 2: An Enhanced Integrated Tool for Genomic Association and Prediction. Plant Genome 2016, 9. [Google Scholar] [CrossRef]
- Müller, B.U.; Stich, B.; Piepho, H.-P. A general method for controlling the genome-wide type I error rate in linkage and association mapping experiments in plants. Heredity 2010, 106, 825–831. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Carell, T. Structural biology of DNA photolyases and cryptochromes. Curr. Opin. Struct. Biol. 2009, 19, 277–285. [Google Scholar] [CrossRef]
- Jansen, M.A.; Gaba, V.; Greenberg, B.M. Higher plants and UV-B radiation: Balancing damage, repair and acclimation. Trends Plant Sci. 1998, 3, 131–135. [Google Scholar] [CrossRef]
- Yang, X.; Yan, J.; Shah, T.; Warburton, M.L.; Li, Q.; Li, L.; Gao, Y.; Chai, Y.; Fu, Z.; Zhou, Y.; et al. Genetic analysis and characterization of a new maize association mapping panel for quantitative trait loci dissection. Theor. Appl. Genet. 2010, 121, 417–431. [Google Scholar] [CrossRef]
- Tossi, V.E.; Regalado, J.J.; Iannicelli, J.; Laino, L.E.; Burrieza, H.P.; Escandón, A.S.; Pitta-Álvarez, S.I. Beyond Arabidopsis: Differential UV-B Response Mediated by UVR8 in Diverse Species. Front. Plant Sci. 2019, 10, 780. [Google Scholar] [CrossRef]
- Zhang, C.; Guo, H.; Zhang, J.; Guo, G.; Schumaker, K.S.; Guo, Y. Arabidopsis Cockayne Syndrome A-Like Proteins 1A and 1B Form a Complex with CULLIN4 and Damage DNA Binding Protein 1A and Regulate the Response to UV Irradiation. Plant Cell 2010, 22, 2353–2369. [Google Scholar] [CrossRef]
- Al Khateeb, W.M.; Schroeder, D.F. Overexpression of Arabidopsis damaged DNA binding protein 1A (DDB1A) enhances UV tolerance. Plant Mol. Biol. 2009, 70, 371–383. [Google Scholar] [CrossRef]
- Rassoolzadeh, H.; Böhm, S.; Hedström, E.; Gad, H.; Helleday, T.; Henriksson, S.; Farnebo, M. Overexpression of the scaffold WD40 protein WRAP53beta enhances the repair of and cell survival from DNA double-strand breaks. Cell Death Dis. 2016, 7, e2267. [Google Scholar] [CrossRef]
- Vandenbussche, F.; Tilbrook, K.; Fierro, A.C.; Marchal, K.; Poelman, D.; Van Der Straeten, D.; Ulm, R. Photoreceptor-Mediated Bending towards UV-B in Arabidopsis. Mol. Plant 2014, 7, 1041–1052. [Google Scholar] [CrossRef]
- Vanhaelewyn, L.; Prinsen, E.; Van Der Straeten, D.; Vandenbussche, F. Hormone-controlled UV-B responses in plants. J. Exp. Bot. 2016, 67, 4469–4482. [Google Scholar] [CrossRef]
- Hayes, S.; Sharma, A.; Fraser, D.P.; Trevisan, M.; Cragg-Barber, C.K.; Tavridou, E.; Fankhauser, C.; Jenkins, G.I.; Franklin, K.A. UV-B Perceived by the UVR8 Photoreceptor Inhibits Plant Thermomorphogenesis. Curr. Biol. 2017, 27, 120–127. [Google Scholar] [CrossRef] [PubMed]
- Dharmasiri, N.; Dharmasiri, S.; Weijers, D.; Lechner, E.; Yamada, M.; Hobbie, L.; Ehrismann, J.S.; Jürgens, G.; Estelle, M. Plant Development Is Regulated by a Family of Auxin Receptor F Box Proteins. Dev. Cell 2005, 9, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Villalobos, L.I.A.C.; Lee, S.; De Oliveira, C.; Ivetac, A.; Brandt, W.; Armitage, L.; Sheard, L.B.; Tan, X.; Parry, G.; Mao, H.; et al. A combinatorial TIR1/AFB-Aux/IAA co-receptor system for differential sensing of auxin. Nat. Chem. Biol. 2012, 8, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Yadav, A.; Singh, D.; Lingwan, M.; Yadukrishnan, P.; Masakapalli, S.K.; Datta, S. Light signaling and UV-B-mediated plant growth regulation. J. Integr. Plant Biol. 2020, 62, 1270–1292. [Google Scholar] [CrossRef]
- Galvão, V.C.; Fankhauser, C. Sensing the light environment in plants: Photoreceptors and early signaling steps. Curr. Opin. Neurobiol. 2015, 34, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Chaves, I.; Pokorny, R.; Byrdin, M.; Hoang, N.; Ritz, T.; Brettel, K.; Essen, L.-O.; van der Horst, G.; Batschauer, A.; Ahmad, M. The Cryptochromes: Blue Light Photoreceptors in Plants and Animals. Annu. Rev. Plant Biol. 2011, 62, 335–364. [Google Scholar] [CrossRef]
- Wulff, D.L.; Rupert, C.S. Disappearance of thymine photodimer in ultraviolet irradiated DNA upon treatment with a photoreactivating enzyme from Baker’s yeast. Biochem. Biophys. Res. Commun. 1962, 7, 237–240. [Google Scholar] [CrossRef]
- Tagua, V.G.; Pausch, M.; Eckel, M.; Gutiérrez, G.; Miralles-Durán, A.; Sanz, C.; Eslava, A.P.; Pokorny, R.; Corrochano, L.M.; Batschauer, A. Fungal cryptochrome with DNA repair activity reveals an early stage in cryptochrome evolution. Proc. Natl. Acad. Sci. USA 2015, 112, 15130–15135. [Google Scholar] [CrossRef]
- Lipka, A.E.; Tian, F.; Wang, Q.; Peiffer, J.; Li, M.; Bradbury, P.J.; Gore, M.; Buckler, E.S.; Zhang, Z. GAPIT: Genome association and prediction integrated tool. Bioinformatics 2012, 28, 2397–2399. [Google Scholar] [CrossRef]
- VanRaden, P.M. Efficient Methods to Compute Genomic Predictions. J. Dairy Sci. 2008, 91, 4414–4423. [Google Scholar] [CrossRef] [PubMed]
- Astorkia, M.; Hernandez, M.; Bocs, S.; De Armentia, E.L.; Herran, A.; Ponce, K.; León, O.; Morales, S.; Quezada, N.; Orellana, F.; et al. Association Mapping Between Candidate Gene SNP and Production and Oil Quality Traits in Interspecific Oil Palm Hybrids. Plants 2019, 8, 377. [Google Scholar] [CrossRef]
- Gao, L.; Turner, M.K.; Chao, S.; Kolmer, J.; Anderson, J.A. Genome Wide Association Study of Seedling and Adult Plant Leaf Rust Resistance in Elite Spring Wheat Breeding Lines. PLoS ONE 2016, 11, e0148671. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Grant, D.; Nelson, R.T.; Cannon, S.B.; Shoemaker, R.C. SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res. 2009, 38, 843–846. [Google Scholar] [CrossRef]
- Yoon, M.Y.; Kim, M.Y.; Shim, S.; Kim, K.D.; Ha, J.; Shin, J.H.; Kang, S.; Lee, S.-H. Transcriptomic Profiling of Soybean in Response to High-Intensity UV-B Irradiation Reveals Stress Defense Signaling. Front. Plant Sci. 2016, 7, 1917. [Google Scholar] [CrossRef]
- Guénin, S.; Mauriat, M.; Pelloux, J.; Van Wuytswinkel, O.; Bellini, C.; Gutierrez, L. Normalization of qRT-PCR data: The necessity of adopting a systematic, experimental conditions-specific, validation of references. J. Exp. Bot. 2009, 60, 487–493. [Google Scholar] [CrossRef]
- Rao, X.; Huang, X.; Zhou, Z.; Lin, X. An improvement of the 2ˆ(–delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinform. Biomath. 2013, 3, 71–85. [Google Scholar]


| Trait | Chr. | SNP | Position | FDR Unadjusted p-Values | −log(p) | maf | R2 without SNP | R2 with SNP | FDR Adjusted p-Values | Effect | Linkage Disequilibrium Block | Distance | No. of Genes |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DLC | 10 | AX-90454793 | 38322497 | 8.65 × 10−5 | 4.06 | 0.16 | 0.02 | 0.04 | 0.57 | −0.46 | 382,934~383,225 | 292 bp | 1 |
| 11 | AX-90522955 | 1168027 | 4.22 × 10−5 | 4.38 | 0.40 | 0.02 | 0.05 | 0.46 | 0.37 | 9,939,957~1,013,188 | 19,194 bp | 18 | |
| AX-90521132 | 9939957 | 6.50 × 10−6 | 5.19 | 0.08 | 0.02 | 0.05 | 0.28 | −0.69 | 1,144,379-1,168,553 | 24,174 bp | 8 | ||
| AX-90466824 | 9955814 | 7.83 × 10−5 | 4.11 | 0.16 | 0.02 | 0.05 | 0.57 | −0.45 | - | - | - | ||
| AX-90325683 | 10099676 | 1.27 × 10−5 | 4.90 | 0.08 | 0.02 | 0.05 | 0.28 | −0.65 | - | - | - | ||
| AX-90318887 | 10104602 | 1.27 × 10−5 | 4.90 | 0.08 | 0.02 | 0.05 | 0.28 | −0.65 | - | - | - | ||
| AX-90379400 | 10109488 | 3.77 × 10−5 | 4.42 | 0.10 | 0.02 | 0.05 | 0.46 | −0.57 | - | - | - | ||
| AX-90353742 | 10119213 | 3.73 × 10−5 | 4.43 | 0.10 | 0.02 | 0.05 | 0.46 | −0.58 | - | - | - | ||
| AX-90477047 | 10121857 | 6.84 × 10−5 | 4.16 | 0.10 | 0.02 | 0.05 | 0.56 | −0.56 | - | - | - | ||
| AX-90427669 | 10126101 | 5.30 × 10−5 | 4.28 | 0.10 | 0.02 | 0.05 | 0.50 | −0.58 | - | - | - | ||
| DLS | 2 | AX-90319706 | 15429030 | 6.67 × 10−5 | 4.18 | 0.18 | 0.05 | 0.07 | 0.63 | −0.45 | - | - | - |
| AX-90457128 | 15430402 | 6.26 × 10−5 | 4.20 | 0.18 | 0.05 | 0.08 | 0.63 | −0.46 | - | - | - | ||
| AX-90334094 | 15460775 | 4.56 × 10−5 | 4.34 | 0.18 | 0.05 | 0.08 | 0.63 | −0.47 | 1,516,984~1,550,224 | 29,095 bp | 9 | ||
| AX-90344232 | 15464333 | 0.0000847 | 4.07 | 0.19 | 0.05 | 0.07 | 0.64 | −0.45 | - | - | - | ||
| AX-90436068 | 15467717 | 6.09 × 10−5 | 4.22 | 0.18 | 0.05 | 0.08 | 0.63 | −0.46 | - | - | - | ||
| AX-90474824 | 15473795 | 6.60 × 10−5 | 4.18 | 0.18 | 0.05 | 0.07 | 0.63 | −0.45 | - | - | - | ||
| AX-90329251 | 15474547 | 6.60 × 10−5 | 4.18 | 0.18 | 0.05 | 0.07 | 0.63 | −0.45 | - | - | - | ||
| 6 | AX-90333167 | 8071856 | 5.13 × 10−5 | 4.29 | 0.23 | 0.05 | 0.08 | 0.63 | −0.43 | 7,516,555~8,315,464 | 798,909 bp | 94 |
| Trait | Chr. | Linkage Disequilibrium Block | Candidate Gene | Gene Position | Description |
|---|---|---|---|---|---|
| DLC 1 | 11 | 9,939,957–10,131,882 | Glyma.11g130800 | 9,983,698–9,993,001 | Transducin/WD40 repeat-like superfamily protein, WD40 repeat family |
| DLS 2 | 2 | 1,516,984–1,550,224 | Glyma.02g017500 | 1,550,874–1,554,518 | CRYPTOCHROME, photolyase/blue-light receptor 2, DNA repair, DNA photolyase activity |
| 6 | 7,516,555–8,315,464 | Glyma.06g095400 | 7,530,783–7,533,737 | Auxin F-box protein 5, protein binding | |
| Glyma.06g097800 | 7,725,465–7,766,431 | Protein binding, related BEACH and WD40 repeat proteins, WD domain | |||
| Glyma.06g103200 | 8,205,073–8,210,842 | CRYPTOCHROME 1, DNA repair, blue/ultraviolet sensing protein C terminal, DNA photolyase activity |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, T.; Kim, K.D.; Kim, J.-M.; Shin, I.; Heo, J.; Jung, J.; Lee, J.; Moon, J.-K.; Kang, S. Genome-Wide Association Study for Ultraviolet-B Resistance in Soybean (Glycine max L.). Plants 2021, 10, 1335. https://doi.org/10.3390/plants10071335
Lee T, Kim KD, Kim J-M, Shin I, Heo J, Jung J, Lee J, Moon J-K, Kang S. Genome-Wide Association Study for Ultraviolet-B Resistance in Soybean (Glycine max L.). Plants. 2021; 10(7):1335. https://doi.org/10.3390/plants10071335
Chicago/Turabian StyleLee, Taeklim, Kyung Do Kim, Ji-Min Kim, Ilseob Shin, Jinho Heo, Jiyeong Jung, Juseok Lee, Jung-Kyung Moon, and Sungteag Kang. 2021. "Genome-Wide Association Study for Ultraviolet-B Resistance in Soybean (Glycine max L.)" Plants 10, no. 7: 1335. https://doi.org/10.3390/plants10071335
APA StyleLee, T., Kim, K. D., Kim, J.-M., Shin, I., Heo, J., Jung, J., Lee, J., Moon, J.-K., & Kang, S. (2021). Genome-Wide Association Study for Ultraviolet-B Resistance in Soybean (Glycine max L.). Plants, 10(7), 1335. https://doi.org/10.3390/plants10071335