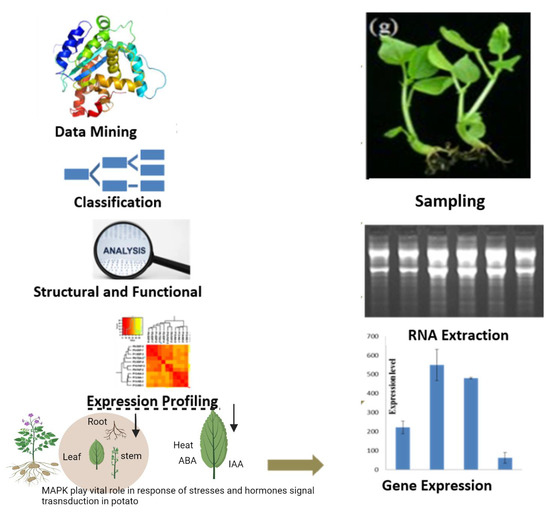
Mitogen-Activated Protein Kinase Expression Profiling Revealed Its Role in Regulating Stress Responses in Potato (Solanum tuberosum)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification of the StMAPKs
2.2. Multiple Sequence Alignment and Phylogenetic Analysis
2.3. Chromosomal Locations, Synteny, and Selective Pressure Analysis
2.4. Gene Structure Analysis and Conserved Motif Identification
2.5. Cis-Elements Analysis
2.6. Expression Analysis of MAPK Genes
2.7. Plant Material and Sample Collection
2.8. RNA Extraction and Quantitative Real-Time PCR Analysis
3. Results
3.1. Identification and Physiological Properties of MAPK Genes
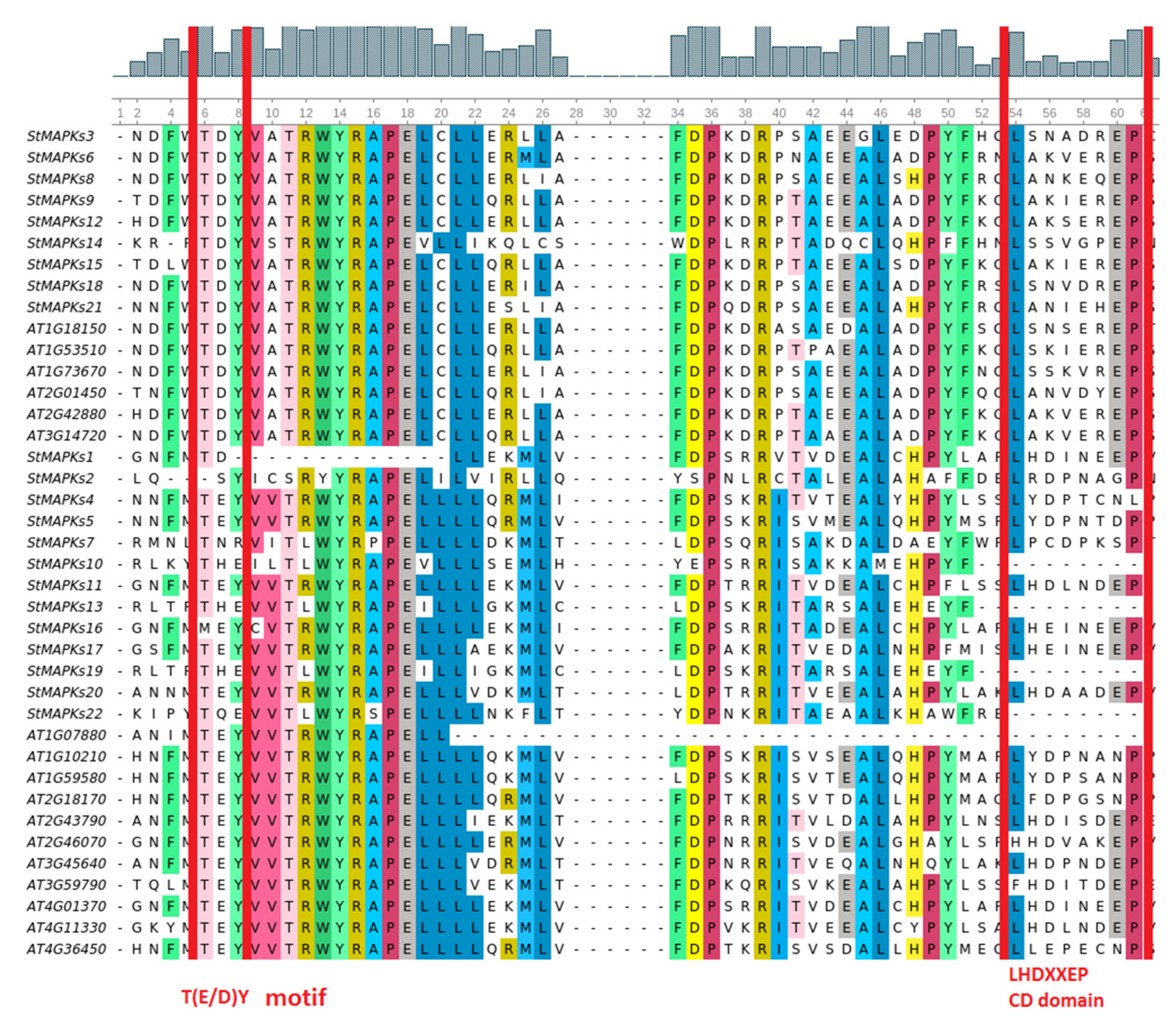
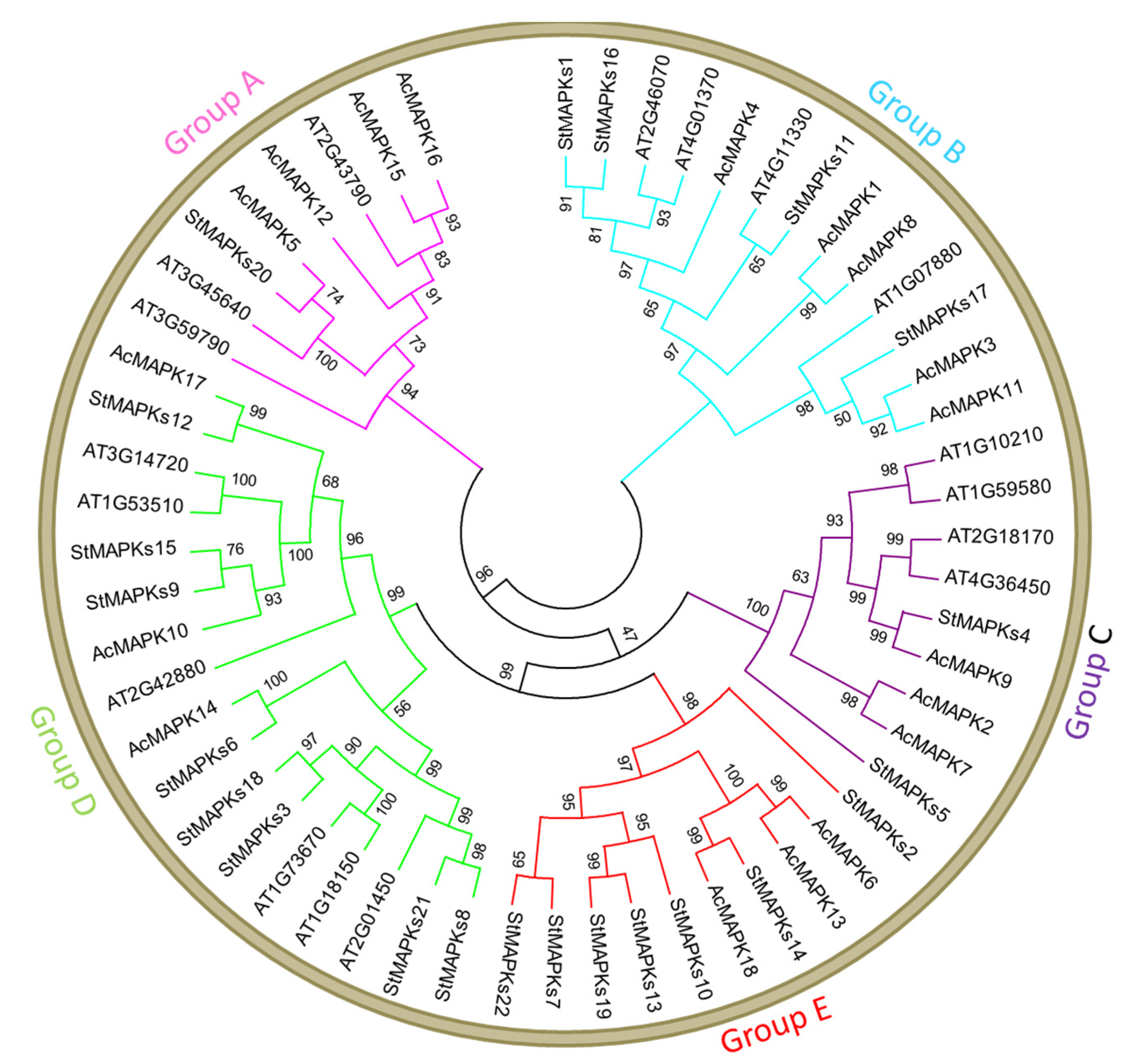
3.2. Multiple Sequence Alignment and Phylogenetic Analysis of MAPK Genes
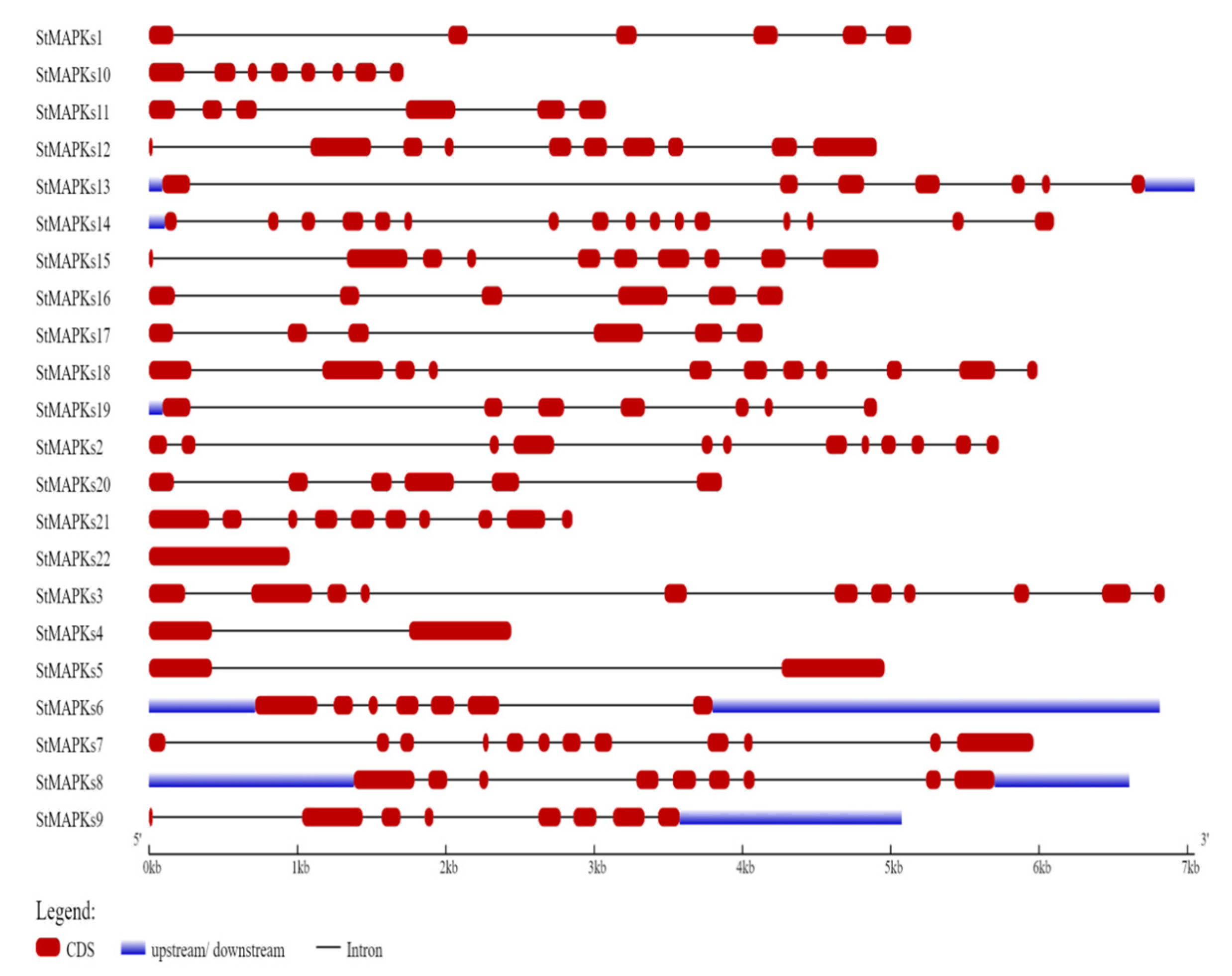
3.3. Gene Structure and Conserved Motif Analysis
3.4. Chromosomal Distribution of MAPK Genes and Cis-Element Analysis
3.5. Gene Duplications of MAPK Genes
3.6. Comparative Synteny Analysis of Identified MAPK Protein Sequences
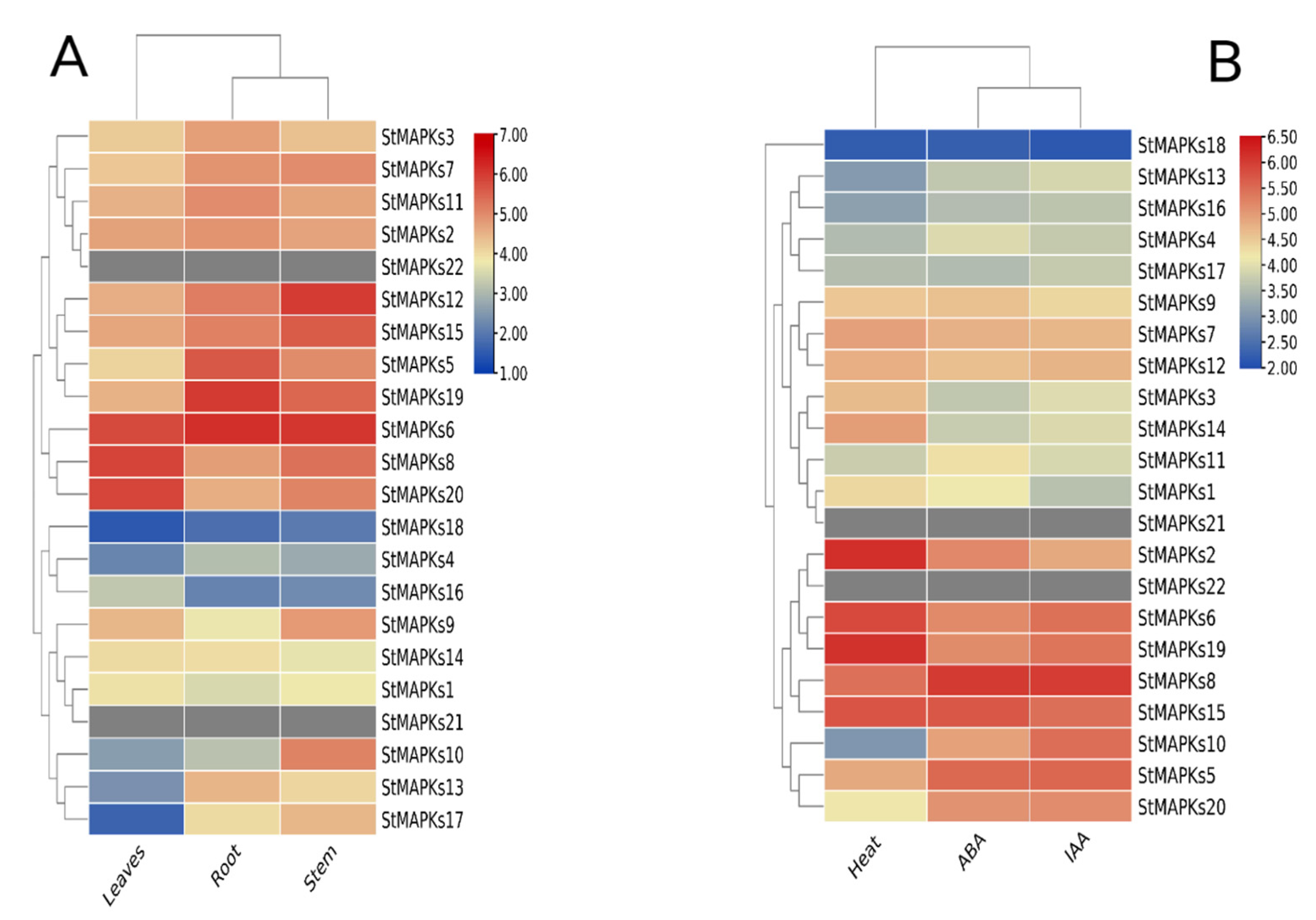
3.7. Tissue-Specific Expression Profiling
3.8. Expression Patterns of StMAPK Genes in Response to Heat Stress
3.9. Expression Patterns of StMAPK Genes in Response to Phytohormones
3.10. RT-qPCR Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Zaynab, M.; Sharif, Y.; Abbas, S.; Afzal, M.Z.; Qasim, M.; Khalofah, A.; Ansari, M.J.; Khan, K.A.; Tao, L.; Li, S. Saponin toxicity as key player in plant defence against pathogens. Toxicon 2021, 193, 21–27. [Google Scholar] [CrossRef]
- Zaynab, M.; Fatima, M.; Abbas, S.; Sharif, Y.; Umair, M.; Zafar, M.H.; Bahadar, K. Role of secondary metabolites in plant defense against pathogens. Microb. Pathog. 2018, 124, 198–202. [Google Scholar] [CrossRef]
- Chen, T.; Zhu, H.; Ke, D.; Cai, K.; Wang, C.; Gou, H.; Hong, Z.; Zhang, Z. A MAP Kinase Kinase Interacts with SymRK and Regulates Nodule Organogenesis in Lotus japonicus. Plant Cell 2012, 24, 823–838. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zaynab, M.; Sharif, Y.; Fatima, M.; Afzal, M.Z.; Aslam, M.M.; Raza, M.F.; Anwar, M.; Sajjad, N.; Yang, X.; Li, S. CRISPR/Cas9 to generate plant immunity against pathogen. Microb. Pathog. 2020, 141, 103996. [Google Scholar] [CrossRef] [PubMed]
- Ijaz, B.; Formentin, E.; Ronci, B.; Locato, V.; Barizza, E.; Hyder, M.Z.; Schiavo, F.L.; Yasmin, T. Salt tolerance in indica rice cell cultures depends on a fine tuning of ROS signalling and homeostasis. PLoS ONE 2019, 14, e0213986. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhou, Y.; Zhang, Q.; Liu, Q.; Li, L.; Sun, C.; Wang, K.; Wang, Y.; Zhao, M.; Li, H.; et al. Structural variation, functional differentiation and expression characteristics of the AP2/ERF gene family and its response to cold stress and methyl jasmonate in Panax ginseng C.A. Meyer. PLoS ONE 2020, 15, e0226055. [Google Scholar] [CrossRef]
- Takahashi, Y.; Soyano, T.; Kosetsu, K.; Sasabe, M.; Machida, Y. HINKEL kinesin, ANP MAPKKKs and MKK6/ANQ MAPKK, which phosphorylates and activates MPK4 MAPK, constitute a pathway that is required for cytokinesis in Arabidopsis thaliana. Plant Cell Physiol. 2010, 51, 1766–1776. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, F.Y.; Hu, F.; Zhang, S.Y.; Wang, K.; Zhang, C.R.; Liu, T. MAPKs regulate root growth by influencing auxin signaling and cell cycle-related gene expression in cadmium-stressed rice. Environ. Sci. Pollut. Res. 2013, 20, 5449–5460. [Google Scholar] [CrossRef]
- Shang, X.; Dong, G.; Zhang, H.; Zhang, L.; Yu, X.; Li, J.; Wang, X.; Yue, B.; Zhao, Y.; Wu, Y. Polybrominated diphenyl ethers (PBDEs) and indicator polychlorinated biphenyls (PCBs) in various marine fish from Zhoushan fishery, China. Food Control. 2016, 67, 240–246. [Google Scholar] [CrossRef]
- Dóczi, R.; Hatzimasoura, E.; Bilooei, S.F.; Ahmad, Z.; Ditengou, F.A.; López-Juez, E.; Palme, K.; Bögre, L. The MKK7-MPK6 MAP Kinase Module Is a Regulator of Meristem Quiescence or Active Growth in Arabidopsis. Front. Plant Sci. 2019, 10, 202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asai, T.; Tena, G.; Plotnikova, J.; Willmann, M.R.; Chiu, W.-L.; Gomez-Gomez, L.; Boller, T.; Ausubel, F.M.; Sheen, J. MAP kinase signalling cascade in Arabidopsis innate immunity. Nat. Cell Biol. 2002, 415, 977–983. [Google Scholar] [CrossRef]
- Virk, N.; Li, D.; Tian, L.; Huang, L.; Hong, Y.; Li, X.; Zhang, Y.; Liu, B.; Zhang, H.; Song, F. Arabidopsis Raf-Like Mitogen-Activated Protein Kinase Kinase Kinase Gene Raf43 Is Required for Tolerance to Multiple Abiotic Stresses. PLoS ONE 2015, 10, e0133975. [Google Scholar] [CrossRef]
- Meng, X.; Zhang, S. MAPK Cascades in Plant Disease Resistance Signaling. Annu. Rev. Phytopathol. 2013, 51, 245–266. [Google Scholar] [CrossRef]
- Cristina, M.; Petersen, M.; Mundy, J. Mitogen-Activated Protein Kinase Signaling in Plants. Annu. Rev. Plant Biol. 2010, 61, 621–649. [Google Scholar] [CrossRef] [PubMed]
- Fiil, B.K.; Petersen, K.; Petersen, M.; Mundy, J. Gene regulation by MAP kinase cascades. Curr. Opin. Plant Biol. 2009, 12, 615–621. [Google Scholar] [CrossRef] [PubMed]
- Colcombet, J.; Hirt, H. Arabidopsis MAPKs: A complex signalling network involved in multiple biological processes. Biochem. J. 2008, 413, 217–226. [Google Scholar] [CrossRef]
- Chen, L.; Hu, W.; Tan, S.; Wang, M.; Ma, Z.; Zhou, S.; Deng, X.; Zhang, Y.; Huang, C.; Yang, G.; et al. Genome-Wide Identification and Analysis of MAPK and MAPKK Gene Families in Brachypodium distachyon. PLoS ONE 2012, 7, e46744. [Google Scholar] [CrossRef] [Green Version]
- Agrawal, G.K.; Iwahashi, H.; Rakwal, R. Rice MAPKs. Biochem. Biophys. Res. Commun. 2003, 302, 171–180. [Google Scholar] [CrossRef]
- Rohila, J.S.; Yang, Y. Rice Mitogen-activated Protein Kinase Gene Family and Its Role in Biotic and Abiotic Stress Response. J. Integr. Plant Biol. 2007, 49, 751–759. [Google Scholar] [CrossRef]
- Wei, C.; Liu, X.; Long, D.; Guo, Q.; Fang, Y.; Bian, C.; Zhang, D.; Zeng, Q.; Xiang, Z.; Zhao, A. Molecular cloning and expression analysis of mulberry MAPK gene family. Plant Physiol. Biochem. 2014, 77, 108–116. [Google Scholar] [CrossRef]
- Lian, W.-W.; Tang, Y.-M.; Gao, S.-Q.; Zhang, Z.; Zhao, X.; Zhao, C.-P. Phylogenetic Analysis and Expression Patterns of the MAPK Gene Family in Wheat (Triticum aestivum L.). J. Integr. Agric. 2012, 11, 1227–1235. [Google Scholar] [CrossRef]
- Hyun, T.K.; Kim, J.-S.; Kwon, S.-Y.; Kim, S.-H. Comparative genomic analysis of mitogen activated protein kinase gene family in grapevine. Genes Genom. 2010, 32, 275–281. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, D.; Wang, L.; Li, D. Genome-Wide Analysis of Mitogen-Activated Protein Kinase Gene Family in Maize. Plant Mol. Biol. Rep. 2013, 31, 1446–1460. [Google Scholar] [CrossRef]
- Kong, F.; Wang, J.; Cheng, L.; Liu, S.; Wu, J.; Peng, Z.; Lu, G. Genome-wide analysis of the mitogen-activated protein kinase gene family in Solanum lycopersicum. Gene 2012, 499, 108–120. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Cheng, T.; Wang, G.; Yan, Y.; Xia, Q. Cloning and evolutionary analysis of tobacco MAPK gene family. Mol. Biol. Rep. 2012, 40, 1407–1415. [Google Scholar] [CrossRef]
- Asif, M.H.; Lakhwani, D.; Pathak, S.; Bhambhani, S.; Bag, S.K.; Trivedi, P.K. Genome-wide identification and expression analysis of the mitogen-activated protein kinase gene family from banana suggest involvement of specific members in different stages of fruit ripening. Funct. Integr. Genom. 2013, 14, 161–175. [Google Scholar] [CrossRef]
- Hamel, L.-P.; Nicole, M.-C.; Sritubtim, S.; Morency, M.-J.; Ellis, M.; Ehlting, J.; Beaudoin, N.; Barbazuk, B.; Klessig, D.; Lee, J.; et al. Ancient signals: Comparative genomics of plant MAPK and MAPKK gene families. Trends Plant Sci. 2006, 11, 192–198. [Google Scholar] [CrossRef]
- Zhang, S.; Xu, R.; Luo, X.; Jiang, Z.; Shu, H. Genome-wide identification and expression analysis of MAPK and MAPKK gene family in Malus domestica. Gene 2013, 531, 377–387. [Google Scholar] [CrossRef]
- Sattar, F.; Hamooh, B.; Wellman, G.; Ali, A.; Shah, S.; Anwar, Y.; Mousa, M. Growth and Biochemical Responses of Potato Cultivars under In Vitro Lithium Chloride and Mannitol Simulated Salinity and Drought Stress. Plants 2021, 10, 924. [Google Scholar] [CrossRef]
- Dahal, K.; Li, X.-Q.; Tai, H.; Creelman, A.; Bizimungu, B. Improving potato stress tolerance and tuber yield under a climate change scenario—A current overview. Front. Plant Sci. 2019, 10, 563. [Google Scholar] [CrossRef]
- Li, G.; Hou, M.; Liu, Y.; Pei, Y.; Ye, M.; Zhou, Y.; Huang, C.; Zhao, Y.; Ma, H. Genome-wide identification, characterization and expression analysis of the non-specific lipid transfer proteins in potato. BMC Genom. 2019, 20, 375. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Zaynab, M.; Chen, H.; Chen, Y.; Ouyang, L.; Yang, X.; Hu, Z.; Li, S. Signs of biofilm formation in the genome of Labrenzia sp. PO1. Saudi J. Biol. Sci. 2021, 28, 1900–1912. [Google Scholar] [CrossRef]
- Zaynab, M.; Chen, H.; Chen, Y.; Ouyang, L.; Yang, X.; Xu, W.; Zeng, Q.; Khan, K.A.; Hassan, M.M.; Hassan, S.; et al. Transcriptome analysis revealed transporter proteins role in the growth of Labrenzia sp. PO1 and SY1. J. King Saud Univ. Sci. 2021, 33, 101433. [Google Scholar] [CrossRef]
- Cao, X.; Zhou, P.; Zhang, X.; Zhu, S.; Zhong, X.; Xiao, Q.; Ding, B.; Li, Y. Identification of an RNA Silencing Suppressor from a Plant Double-Stranded RNA Virus. J. Virol. 2005, 79, 13018–13027. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van De Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Bielawski, J.P. Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 2000, 15, 496–503. [Google Scholar] [CrossRef]
- Zaynab, M.; Kanwal, S.; Hussain, I.; Qasim, M.; Noman, A.; Ali, G.; Jamil, A.; Rehman, N.; Buriro, M.; Abbas, S.; et al. Rice chitinase gene expression in genetically engineered potato confers resistance against Fusarium solani and Rhizictonia solani. PSM Microbiol. 2017, 2, 63–73. [Google Scholar]
- Quandahor, P.; Gou, Y.; Lin, C.; Coulter, J.A.; Liu, C. Comparison of root tolerance to drought and aphid (Myzus persicae Sulzer) resistance among different potato (Solanum tuberosum L.) cultivars. Sci. Rep. 2021, 11, 628. [Google Scholar] [CrossRef]
- Rehman, N.; Khan, M.R.; Abbas, Z.; Rafique, R.S.; Zaynab, M.; Qasim, M.; Noor, S.; Inam, S.; Ali, G.M. Functional characterization of Mitogen-Activated Protein Kinase Kinase (MAPKK) gene in Halophytic Salicornia europaea against salt stress. Environ. Exp. Bot. 2020, 171, 103934. [Google Scholar] [CrossRef]
- Reyna, N.S.; Yang, Y. Molecular Analysis of the Rice MAP Kinase Gene Family in Relation to Magnaporthe grisea Infection. Mol. Plant Microbe Interact. 2006, 19, 530–540. [Google Scholar] [CrossRef] [Green Version]
- Nicole, M.-C.; Hamel, L.-P.; Morency, M.-J.; Beaudoin, N.; E Ellis, B.; Séguin, A. MAP-ping genomic organization and organ-specific expression profiles of poplar MAP kinases and MAP kinase kinases. BMC Genom. 2006, 7, 223. [Google Scholar] [CrossRef] [Green Version]
- MAPK Group; Ichimura, K.; Shinozaki, K.; Tena, G.; Sheen, J.; Henry, Y.; Champion, A.; Kreis, M.; Zhang, S.; Hirt, H.; et al. Mitogen-activated protein kinase cascades in plants: A new nomenclature. Trends Plant Sci. 2002, 7, 301–308. [Google Scholar] [CrossRef]
- Moore, R.C.; Purugganan, M.D. The evolutionary dynamics of plant duplicate genes. Curr. Opin. Plant Biol. 2005, 8, 122–128. [Google Scholar] [CrossRef]
- Flagel, L.E.; Wendel, J.F. Gene duplication and evolutionary novelty in plants. New Phytol. 2009, 183, 557–564. [Google Scholar] [CrossRef]
- Mattick, J.; Gagen, M. The Evolution of Controlled Multitasked Gene Networks: The Role of Introns and Other Noncoding RNAs in the Development of Complex Organisms. Mol. Biol. Evol. 2001, 18, 1611–1630. [Google Scholar] [CrossRef] [Green Version]
- Jeffares, D.C.; Penkett, C.J.; Bahler, J. Rapidly regulated genes are intron poor. Trends Genet. 2008, 24, 375–378. [Google Scholar] [CrossRef] [PubMed]
- Waqas, M.; Azhar, M.T.; Rana, I.A.; Azeem, F.; Ali, M.A.; Nawaz, M.A.; Chung, G.; Atif, R.M. Genome-wide identification and expression analyses of WRKY transcription factor family members from chickpea (Cicer arietinum L.) reveal their role in abiotic stress-responses. Genes Genom. 2019, 41, 467–481. [Google Scholar] [CrossRef] [PubMed]
- Bari, A.; Farooq, M.; Hussain, A.; Qamar, M.T.U.; Abbas, M.W.; Mustafa, G.; Karim, A.; Ahmed, I.; Hussain, T. Genome-Wide Bioinformatics Analysis of Aquaporin Gene Family in Maize (Zea mays L.). J. Phylogenet. Evol. Biol. 2018, 6, 1–15. [Google Scholar] [CrossRef]
- Wu, J.; Wang, J.; Pan, C.; Guan, X.; Wang, Y.; Liu, S.; He, Y.; Chen, J.; Chen, L.; Lu, G. Genome-Wide Identification of MAPKK and MAPKKK Gene Families in Tomato and Transcriptional Profiling Analysis during Development and Stress Response. PLoS ONE 2014, 9, e103032. [Google Scholar] [CrossRef]
- Lavin, M.; Herendeen, P.; Wojciechowski, M.F. Evolutionary Rates Analysis of Leguminosae Implicates a Rapid Diversification of Lineages during the Tertiary. Syst. Biol. 2005, 54, 575–594. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goyal, R.K.; Tulpan, D.; Chomistek, N.; Fundora, D.G.-P.; West, C.; Ellis, B.E.; Frick, M.; Laroche, A.; Foroud, N.A. Analysis of MAPK and MAPKK gene families in wheat and related Triticeae species. BMC Genom. 2018, 19, 178. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Wan, Y.; Meng, X.; Zhang, X.; Yao, M.; Miu, W.; Zhu, D.; Yuan, D.; Lu, K.; Li, J.; et al. Genome-Wide Identification and Analysis of MKK and MAPK Gene Families in Brassica Species and Response to Stress in Brassica napus. Int. J. Mol. Sci. 2021, 22, 544. [Google Scholar] [CrossRef]
- Fatima, M.; Ma, X.; Zhou, P.; Zaynab, M.; Ming, R. Auxin regulated metabolic changes underlying sepal retention and development after pollination in spinach. BMC Plant Biol. 2021, 21, 166. [Google Scholar] [CrossRef]
- Zaynab, M.; Kanwal, S.; Furqan, M.; Islam, W.; Noman, A.; Ali, G.M.; Rehman, N.; Zafar, S.; Sughra, K.; Jahanzab, M. Proteomic approach to address low seed germination in Cyclobalnopsis gilva. Biotechnol. Lett. 2017, 39, 1441–1451. [Google Scholar] [CrossRef]
- Kumar, K.; Raina, S.K.; Sultan, S.M. Arabidopsis MAPK signaling pathways and their cross talks in abiotic stress response. J. Plant Biochem. Biotechnol. 2020, 29, 700–714. [Google Scholar] [CrossRef]
- Zhou, H.; Ren, S.; Han, Y.; Zhang, Q.; Qin, L.; Xing, Y. Identification and Analysis of Mitogen-Activated Protein Kinase (MAPK) Cascades in Fragaria vesca. Int. J. Mol. Sci. 2017, 18, 1766. [Google Scholar] [CrossRef] [PubMed] [Green Version]





| Gene | CDS | Genomic | Transcript Name | Location | Star-End | No. of Amino Acids | Molecular Weight | pI | Sub-Cellular Localization |
|---|---|---|---|---|---|---|---|---|---|
| StMAPKs1 | 927 | 5139 | PGSC0003DMT400000192 | ST4.03ch01 | 72545240..72550378 | 308 | 35404.2 | 6.13 | Cytoplasmic |
| StMAPKs2 | 1233 | 5729 | PGSC0003DMT400001392 | ST4.03ch02 | 47182708..47188436 | 410 | 46208.71 | 8.43 | Cytoplasmic |
| StMAPKs3 | 1728 | 6847 | PGSC0003DMT400006281 | ST4.03ch12 | 39794625..39801471 | 575 | 65699.51 | 6.47 | Nuclear |
| StMAPKs4 | 1113 | 2442 | PGSC0003DMT400009066 | ST4.03ch02 | 40117154..40119595 | 370 | 42708.53 | 8.02 | Nuclear |
| StMAPKs5 | 1119 | 4960 | PGSC0003DMT400009491 | ST4.03ch04 | 70130371..70135330 | 372 | 42825.56 | 6.32 | Nuclear |
| StMAPKs6 | 1260 | 3087 | PGSC0003DMT400011136 | ST4.03ch01 | 61223835..61226921 | 419 | 48384.75 | 8.37 | Cytoplasmic |
| StMAPKs7 | 1521 | 5963 | PGSC0003DMT400015241 | ST4.03ch06 | 54220156..54226118 | 506 | 56582.89 | 9.31 | Nuclear |
| StMAPKs8 | 1488 | 5700 | PGSC0003DMT400015263 | ST4.03ch04 | 2187114..2192813 | 495 | 56445.57 | 6.61 | Cytoplasmic |
| StMAPKs9 | 1284 | 3577 | PGSC0003DMT400018187 | ST4.03ch07 | 52242577..52246153 | 427 | 49320.14 | 9.03 | Cytoplasmic |
| StMAPKs10 | 948 | 1716 | PGSC0003DMT400025910 | ST4.03ch04 | 72007479..72009194 | 315 | 36055 | 8.88 | Cytoplasmic |
| StMAPKs11 | 1140 | 3080 | PGSC0003DMT400031773 | ST4.03ch08 | 55187632..55190711 | 379 | 43454.69 | 5.91 | Cytoplasmic |
| StMAPKs12 | 1842 | 4908 | PGSC0003DMT400044709 | ST4.03ch07 | 51103717..51108624 | 613 | 69709.84 | 9.25 | Nuclear |
| StMAPKs13 | 885 | 6714 | PGSC0003DMT400047739 | ST4.03ch11 | 42450013..42456726 | 294 | 33480.78 | 7.09 | Cytoplasmic |
| StMAPKs14 | 1314 | 6102 | PGSC0003DMT400051979 | ST4.03ch06 | 58205983..58212084 | 437 | 49665 | 5.44 | Cytoplasmic |
| StMAPKs15 | 1782 | 4917 | PGSC0003DMT400054750 | ST4.03ch10 | 2533056..2537972 | 593 | 67184.24 | 9.22 | Nuclear |
| StMAPKs16 | 1131 | 4243 | PGSC0003DMT400055751 | ST4.03ch05 | 44001222..44005494 | 376 | 43056.15 | 6.76 | Nuclear |
| StMAPKs17 | 1119 | 4136 | PGSC0003DMT400065261 | ST4.03ch11 | 44614589..44618724 | 372 | 42685.76 | 4.97 | Cytoplasmic |
| StMAPKs18 | 1818 | 5991 | PGSC0003DMT400074096 | ST4.03ch06 | 50702397..50708387 | 605 | 69042.25 | 7.07 | Nuclear |
| StMAPKs19 | 885 | 5909 | PGSC0003DMT400075349 | ST4.03ch12 | 57689402..57694310 | 294 | 33688.02 | 7.12 | Cytoplasmic |
| StMAPKs20 | 1112 | 3862 | PGSC0003DMT400077272 | ST4.03ch06 | 7463661..7467522 | 373 | 42891.19 | 5.53 | Cytoplasmic |
| StMAPKs21 | 1536 | 2855 | PGSC0003DMT400078323 | ST4.03ch05 | 4324632..4327486 | 511 | 58650.61 | 6.01 | Nuclear |
| StMAPKs22 | 948 | 948 | PGSC0003DMT400086876 | ST4.03ch08 | 4905365..4906312 | 315 | 36394.48 | 9.4 | Cytoplasmic |
| Seq_1 | Seq_2 | Ka | Ks | Ka_Ks | Time (MYA) |
|---|---|---|---|---|---|
| StMAPKs3 | StMAPKs18 | 0.1025318 | 0.5577344 | 0.1838362 | 19.717651 |
| StMAPKs9 | StMAPKs15 | 0.0786731 | 0.5823474 | 0.1350966 | 15.129448 |
| StMAPKs8 | StMAPKs21 | 0.1247762 | 0.5100702 | 0.2446256 | 23.995425 |
| StMAPKs4 | StMAPKs5 | 0.1439549 | NaN | NaN | 27.683632 |
| StMAPKs1 | StMAPKs16 | 0.08686 | 0.4515929 | 0.1923414 | 16.703851 |
| StMAPKs2 | StMAPKs12 | 0.623371 | 3.7728284 | 0.1652264 | 119.87904 |
| StMAPKs13 | StMAPKs19 | 0.0515455 | 0.5004247 | 0.1030036 | 9.9126019 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zaynab, M.; Hussain, A.; Sharif, Y.; Fatima, M.; Sajid, M.; Rehman, N.; Yang, X.; Khan, K.A.; Ghramh, H.A.; Li, S. Mitogen-Activated Protein Kinase Expression Profiling Revealed Its Role in Regulating Stress Responses in Potato (Solanum tuberosum). Plants 2021, 10, 1371. https://doi.org/10.3390/plants10071371
Zaynab M, Hussain A, Sharif Y, Fatima M, Sajid M, Rehman N, Yang X, Khan KA, Ghramh HA, Li S. Mitogen-Activated Protein Kinase Expression Profiling Revealed Its Role in Regulating Stress Responses in Potato (Solanum tuberosum). Plants. 2021; 10(7):1371. https://doi.org/10.3390/plants10071371
Chicago/Turabian StyleZaynab, Madiha, Athar Hussain, Yasir Sharif, Mahpara Fatima, Mateen Sajid, Nazia Rehman, Xuewei Yang, Khalid Ali Khan, Hamed A. Ghramh, and Shuangfei Li. 2021. "Mitogen-Activated Protein Kinase Expression Profiling Revealed Its Role in Regulating Stress Responses in Potato (Solanum tuberosum)" Plants 10, no. 7: 1371. https://doi.org/10.3390/plants10071371