A Systemic View of Carbohydrate Metabolism in Rice to Facilitate Productivity
Abstract
:1. Introduction
2. Materials and Methods
2.1. Integration of Carbohydrate Metabolism Annotation Data
2.2. Collection and Clustering of Microarray Data
2.3. Functional Classification via Literature Search
2.4. Gene Ontology (GO) Enrichment Analysis
2.5. KEGG Enrichment Analysis
2.6. RNA Extraction and qRT-PCR
2.7. Construction of the Conceptual Carbohydrate Metabolism Model
3. Results
3.1. Identification of Genome-Wide Candidate Genes Related to Carbohydrate Metabolism
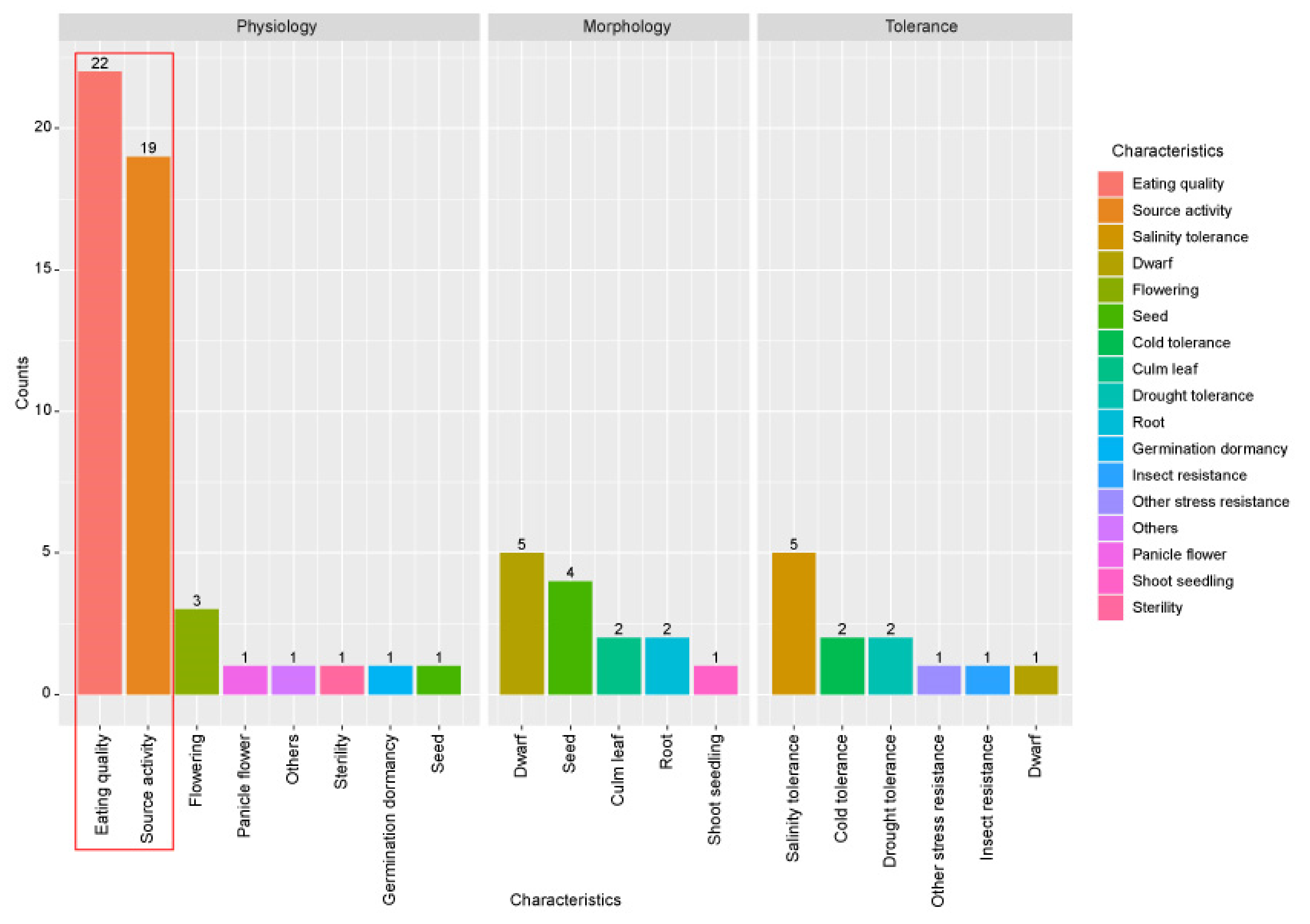
3.2. Functional Analysis of the Characterized Carbohydrate Metabolism-Related Genes
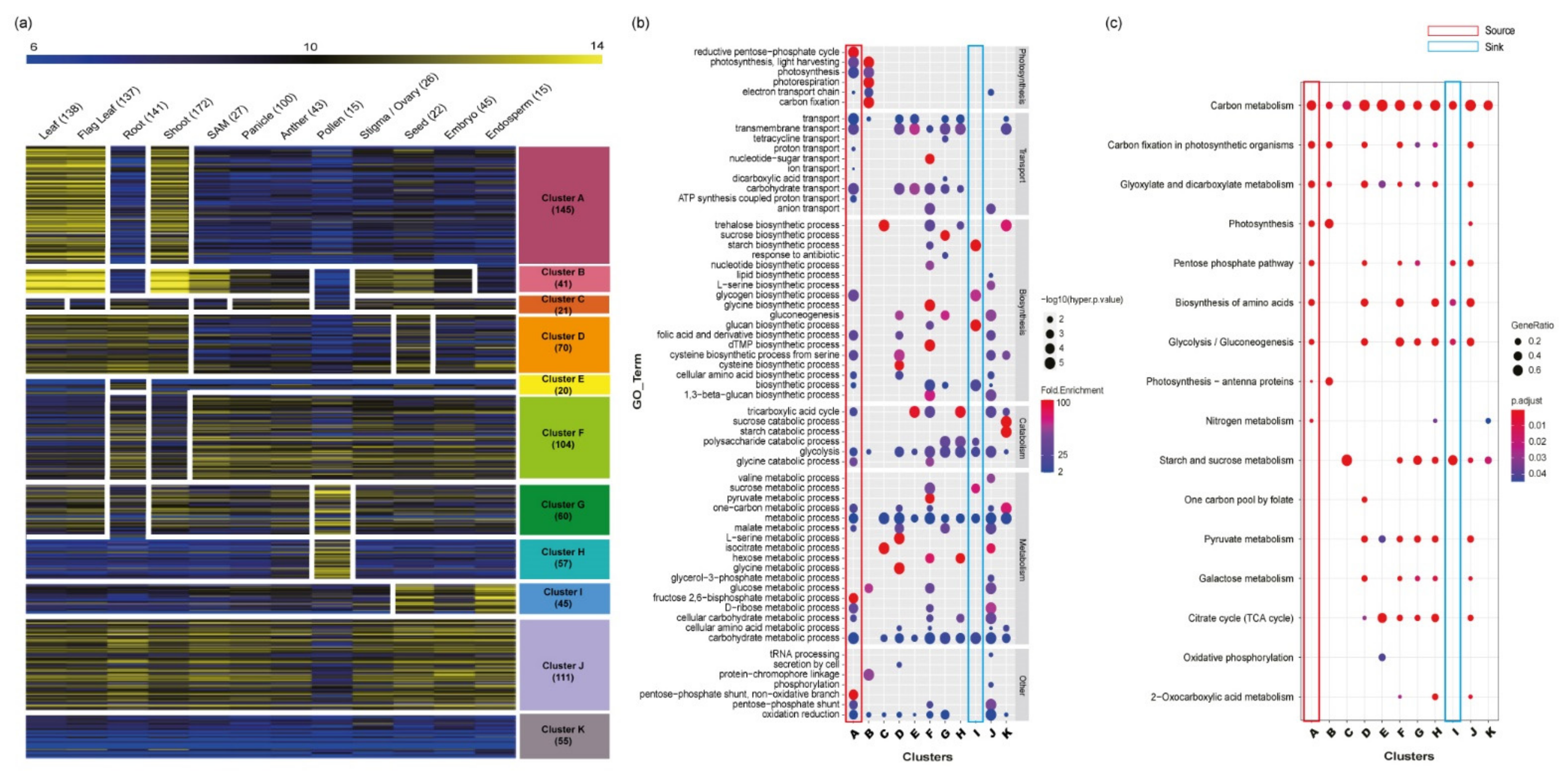
3.3. Anatomical Dissection of Carbohydrate Metabolism-Related Genes via Meta-Expression Analysis
3.4. Functional Comparison and Enrichment Analysis of 11 Anatomical Clusters
3.5. In Silico and In Vitro Expression Verification of Tissue-Preferred Genes
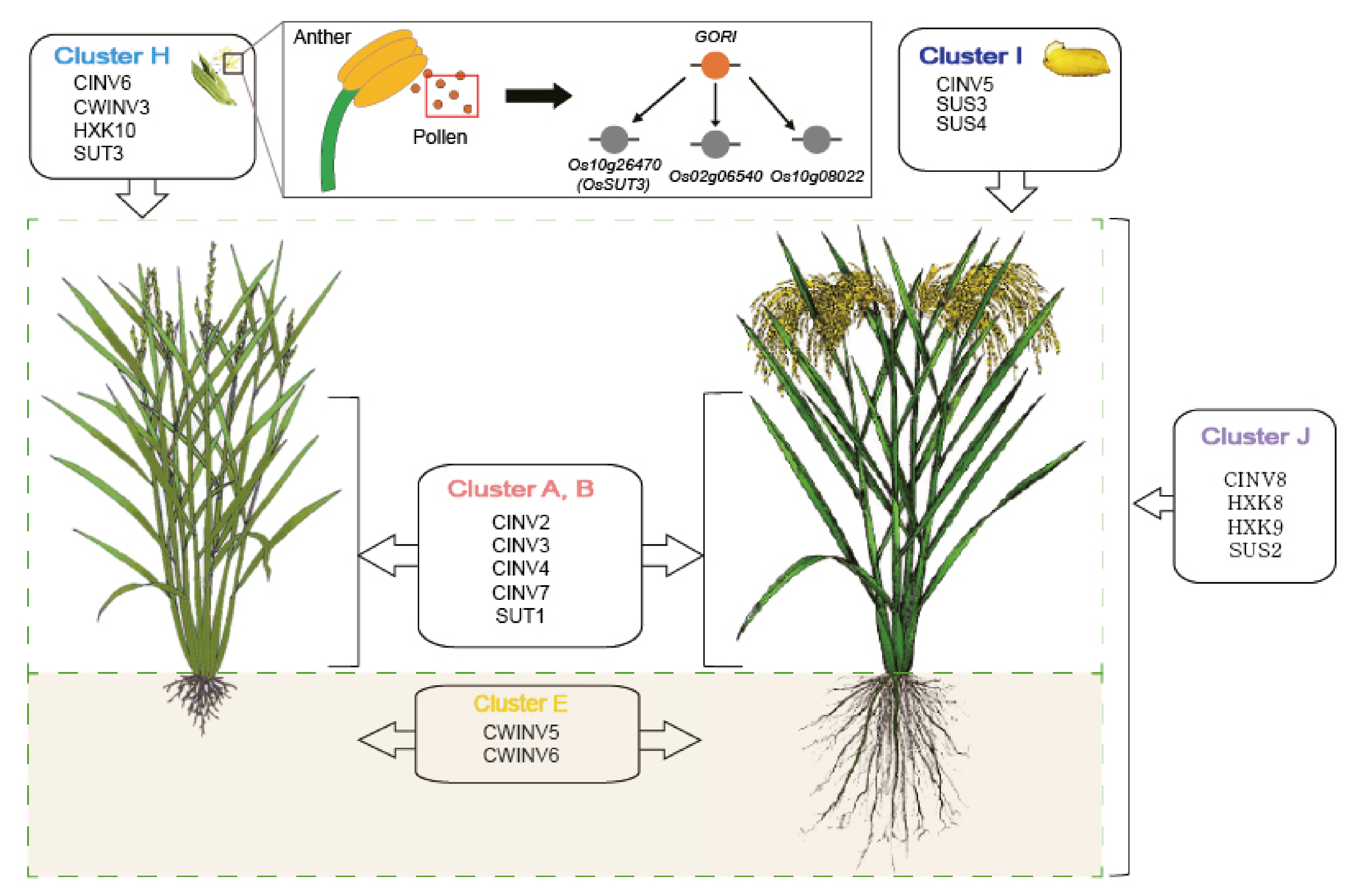
3.6. Construction of a Conceptual Carbohydrate Metabolism Model
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Godfray, H.C.J.; Beddington, J.R.; Crute, I.R.; Haddad, L.; Lawrence, D.; Muir, J.F.; Pretty, J.; Robinson, S.; Thomas, S.M.; Toulmin, C. Food Security: The Challenge of Feeding 9 Billion People. Science 2010, 327, 812. [Google Scholar] [CrossRef] [Green Version]
- Zhao, C.; Liu, B.; Piao, S.; Wang, X.; Lobell, D.B.; Huang, Y.; Huang, M.; Yao, Y.; Bassu, S.; Ciais, P.; et al. Temperature Increase Reduces Global Yields of Major Crops in Four Independent Estimates. Proc. Natl. Acad. Sci. USA 2017, 114, 9326. [Google Scholar] [CrossRef] [Green Version]
- Yu, S.-M.; Lo, S.-F.; Ho, T.-H.D. Source–Sink Communication: Regulated by Hormone, Nutrient, and Stress Cross-Signaling. Trends Plant Sci. 2015, 20, 844–857. [Google Scholar] [CrossRef]
- Sinclair, T.R.; Sheehy, J.E. Erect Leaves and Photosynthesis in Rice. Science 1999, 283, 1455. [Google Scholar] [CrossRef]
- Sakamoto, T.; Morinaka, Y.; Ohnishi, T.; Sunohara, H.; Fujioka, S.; Ueguchi-Tanaka, M.; Mizutani, M.; Sakata, K.; Takatsuto, S.; Yoshida, S.; et al. Erect Leaves Caused by Brassinosteroid Deficiency Increase Biomass Production and Grain Yield in Rice. Nat. Biotechnol. 2006, 24, 105–109. [Google Scholar] [CrossRef]
- Yoo, S.-C.; Cho, S.-H.; Zhang, H.; Paik, H.-C.; Lee, C.-H.; Li, J.; Yoo, J.-H.; Lee, B.-W.; Koh, H.-J.; Seo, H.S.; et al. Quantitative Trait Loci Associated with Functional Stay-Green SNU-SG1 in Rice. Mol. Cells 2007, 24, 83–94. [Google Scholar] [PubMed]
- Lalonde, S.; Tegeder, M.; Throne-Holst, M.; Frommer, W.B.; Patrick, J.W. Phloem Loading and Unloading of Sugars and Amino Acids. Plant Cell Environ. 2003, 26, 37–56. [Google Scholar] [CrossRef] [Green Version]
- Zhang, C.; Turgeon, R. Mechanisms of Phloem Loading. Curr. Opin. Plant Biol. 2018, 43, 71–75. [Google Scholar] [CrossRef]
- Chen, L.-Q.; Qu, X.-Q.; Hou, B.-H.; Sosso, D.; Osorio, S.; Fernie, A.R.; Frommer, W.B. Sucrose Efflux Mediated by SWEET Proteins as a Key Step for Phloem Transport. Science 2012, 335, 207. [Google Scholar] [CrossRef] [PubMed]
- Eom, J.-S.; Chen, L.-Q.; Sosso, D.; Julius, B.T.; Lin, I.; Qu, X.-Q.; Braun, D.M.; Frommer, W.B. SWEETs, Transporters for Intracellular and Intercellular Sugar Translocation. Curr. Opin. Plant Biol. 2015, 25, 53–62. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cho, J.-I.; Ryoo, N.; Ko, S.; Lee, S.-K.; Lee, J.; Jung, K.-H.; Lee, Y.-H.; Bhoo, S.H.; Winderickx, J.; An, G.; et al. Structure, Expression, and Functional Analysis of the Hexokinase Gene Family in Rice (Oryza Sativa L.). Planta 2006, 224, 598–611. [Google Scholar] [CrossRef] [PubMed]
- Wang, E.; Wang, J.; Zhu, X.; Hao, W.; Wang, L.; Li, Q.; Zhang, L.; He, W.; Lu, B.; Lin, H.; et al. Control of Rice Grain-Filling and Yield by a Gene with a Potential Signature of Domestication. Nat. Genet. 2008, 40, 1370–1374. [Google Scholar] [CrossRef]
- Li, B.; Liu, H.; Zhang, Y.; Kang, T.; Zhang, L.; Tong, J.; Xiao, L.; Zhang, H. Constitutive Expression of Cell Wall Invertase Genes Increases Grain Yield and Starch Content in Maize. Plant Biotechnol. J. 2013, 11, 1080–1091. [Google Scholar] [CrossRef]
- Jeon, J.-S.; Jung, K.-H.; Kim, H.-B.; Suh, J.-P.; Khush, G.S. Genetic and Molecular Insights into the Enhancement of Rice Yield Potential. J. Plant Biol. 2011, 54, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Zhang, J. Grain-Filling Problem in ‘Super’ Rice. J. Exp. Bot. 2010, 61, 1–5. [Google Scholar] [CrossRef] [Green Version]
- Slewinski, T.L.; Zhang, C.; Turgeon, R. Structural and Functional Heterogeneity in Phloem Loading and Transport. Front. Plant Sci. 2013, 4, 244. [Google Scholar] [CrossRef] [Green Version]
- Ruan, Y.-L.; Jin, Y.; Huang, J. Capping Invertase Activity by Its Inhibitor: Roles and Implications in Sugar Signaling, Carbon Allocation, Senescence and Evolution. Plant Signal. Behav. 2009, 4, 983–985. [Google Scholar] [CrossRef]
- Zhu, T.; Budworth, P.; Chen, W.; Provart, N.; Chang, H.-S.; Guimil, S.; Su, W.; Estes, B.; Zou, G.; Wang, X. Transcriptional Control of Nutrient Partitioning during Rice Grain Filling. Plant Biotechnol. J. 2003, 1, 59–70. [Google Scholar] [CrossRef] [Green Version]
- Yamakawa, H.; Hirose, T.; Kuroda, M.; Yamaguchi, T. Comprehensive Expression Profiling of Rice Grain Filling-Related Genes under High Temperature Using DNA Microarray. Plant Physiol. 2007, 144, 258–277. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: A User-Driven Tool to Display Genomics Data Sets onto Diagrams of Metabolic Pathways and Other Biological Processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- Ouyang, S.; Zhu, W.; Hamilton, J.; Lin, H.; Campbell, M.; Childs, K.; Thibaud-Nissen, F.; Malek, R.L.; Lee, Y.; Zheng, L.; et al. The TIGR Rice Genome Annotation Resource: Improvements and New Features. Nucleic Acids Res. 2007, 35, D883–D887. [Google Scholar] [CrossRef] [Green Version]
- Chandran, A.K.N.; Yoo, Y.-H.; Cao, P.; Sharma, R.; Sharma, M.; Dardick, C.; Ronald, P.C.; Jung, K.-H. Updated Rice Kinase Database RKD 2.0: Enabling Transcriptome and Functional Analysis of Rice Kinase Genes. Rice 2016, 9, 40. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, P.; Jung, K.-H.; Choi, D.; Hwang, D.; Zhu, J.; Ronald, P.C. The Rice Oligonucleotide Array Database: An Atlas of Rice Gene Expression. Rice 2012, 5, 17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chandran, A.K.N.; Hong, W.-J.; Abhijith, B.; Lee, J.; Kim, Y.-J.; Park, S.K.; Jung, K.-H. Rice Male Gamete Expression Database (RMEDB): A Web Resource for Functional Genomic Studies of Rice Male Organ Development. J. Plant Biol. 2020, 63, 421–430. [Google Scholar] [CrossRef]
- Saeed, A.I.; Sharov, V.; White, J.; Li, J.; Liang, W.; Bhagabati, N.; Braisted, J.; Klapa, M.; Currier, T.; Thiagarajan, M.; et al. TM4: A Free, Open-Source System for Microarray Data Management and Analysis. Biotechniques 2003, 34, 374–378. [Google Scholar] [CrossRef] [Green Version]
- Moon, S.; Oo, M.M.; Kim, B.; Koh, H.-J.; Oh, S.A.; Yi, G.; An, G.; Park, S.K.; Jung, K.-H. Genome-Wide Analyses of Late Pollen-Preferred Genes Conserved in Various Rice Cultivars and Functional Identification of a Gene Involved in the Key Processes of Late Pollen Development. Rice 2018, 11, 28. [Google Scholar] [CrossRef] [Green Version]
- Yamamoto, E.; Yonemaru, J.-I.; Yamamoto, T.; Yano, M. OGRO: The Overview of Functionally Characterized Genes in Rice Online Database. Rice 2012, 5, 26. [Google Scholar] [CrossRef] [Green Version]
- Hong, W.-J.; Jung, K.-H. Comparative Analysis of Flanking Sequence Tags of T-DNA/Transposon Insertional Mutants and Genetic Variations of Fast-Neutron Treated Mutants in Rice. J. Plant Biol. 2018, 61, 80–84. [Google Scholar] [CrossRef]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-3-319-24277-4. [Google Scholar]
- Khatri, P.; Drăghici, S. Ontological Analysis of Gene Expression Data: Current Tools, Limitations, and Open Problems. Bioinformatics 2005, 21, 3587–3595. [Google Scholar] [CrossRef] [Green Version]
- Jung, K.-H.; Dardick, C.; Bartley, L.E.; Cao, P.; Phetsom, J.; Canlas, P.; Seo, Y.-S.; Shultz, M.; Ouyang, S.; Yuan, Q.; et al. Refinement of Light-Responsive Transcript Lists Using Rice Oligonucleotide Arrays: Evaluation of Gene-Redundancy. PLoS ONE 2008, 3, e3337. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.-G.; Han, Y.; He, Q.-Y. ClusterProfiler: An R Package for Comparing Biological Themes among Gene Clusters. OMICS 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Sakai, H.; Lee, S.S.; Tanaka, T.; Numa, H.; Kim, J.; Kawahara, Y.; Wakimoto, H.; Yang, C.; Iwamoto, M.; Abe, T.; et al. Rice Annotation Project Database (RAP-DB): An Integrative and interactive Database for Rice Genomics. Plant Cell Physiol. 2013, 54, e6. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-J.; Kim, M.-H.; Hong, W.-J.; Moon, S.; Kim, E.-J.; Silva, J.; Lee, J.; Lee, S.; Kim, S.T.; Park, S.K.; et al. GORI, Encoding the WD40 Domain Protein, Is Required for Pollen Tube Germination and Elongation in Rice. Plant J. 2021, 105, 1645–1664. [Google Scholar] [CrossRef] [PubMed]
- Hong, W.-J.; Jiang, X.; Ahn, H.R.; Choi, J.; Kim, S.-R.; Jung, K.-H. Systematic Analysis of Cold Stress Response and Diurnal Rhythm Using Transcriptome Data in Rice Reveals the Molecular Networks Related to Various Biological Processes. Int. J. Mol. Sci. 2020, 21, 6872. [Google Scholar] [CrossRef]
- Ge, L.-F.; Chao, D.-Y.; Shi, M.; Zhu, M.-Z.; Gao, J.-P.; Lin, H.-X. Overexpression of the Trehalose-6-Phosphate Phosphatase Gene OsTPP1 Confers Stress Tolerance in Rice and Results in the Activation of Stress Responsive Genes. Planta 2008, 228, 191–201. [Google Scholar] [CrossRef]
- Li, H.-W.; Zang, B.-S.; Deng, X.-W.; Wang, X.-P. Overexpression of the Trehalose-6-Phosphate Synthase Gene OsTPS1 Enhances Abiotic Stress Tolerance in Rice. Planta 2011, 234, 1007–1018. [Google Scholar] [CrossRef]
- Koumoto, T.; Shimada, H.; Kusano, H.; She, K.-C.; Iwamoto, M.; Takano, M. Rice Monoculm Mutation Moc2, Which Inhibits Outgrowth of the Second Tillers, Is Ascribed to Lack of a Fructose-1,6-Bisphosphatase. Plant Biotechnol. 2013, 30, 47–56. [Google Scholar] [CrossRef] [Green Version]
- Eom, J.-S.; Cho, J.-I.; Reinders, A.; Lee, S.-W.; Yoo, Y.; Tuan, P.Q.; Choi, S.-B.; Bang, G.; Park, Y.-I.; Cho, M.-H.; et al. Impaired Function of the Tonoplast-Localized Sucrose Transporter in Rice, OsSUT2, Limits the Transport of Vacuolar Reserve Sucrose and Affects Plant Growth. Plant Physiol. 2011, 157, 109–119. [Google Scholar] [CrossRef] [Green Version]
- Duan, J.; Zhang, M.; Zhang, H.; Xiong, H.; Liu, P.; Ali, J.; Li, J.; Li, Z. OsMIOX, a Myo-Inositol Oxygenase Gene, Improves Drought Tolerance through Scavenging of Reactive Oxygen Species in Rice (Oryza Sativa L.). Plant Sci. 2012, 196, 143–151. [Google Scholar] [CrossRef]
- Ishimaru, K.; Hirose, T.; Aoki, N.; Takahashi, S.; Ono, K.; Yamamoto, S.; Wu, J.; Saji, S.; Baba, T.; Ugaki, M.; et al. Antisense Expression of a Rice Sucrose Transporter OsSUT1 in Rice (Oryza Sativa L.). Plant Cell Physiol. 2001, 42, 1181–1185. [Google Scholar] [CrossRef] [Green Version]
- Sharma, A.; Komatsu, S. Involvement of a Ca(2+)-Dependent Protein Kinase Component Downstream to the Gibberellin-Binding Phosphoprotein, RuBisCO Activase, in Rice. Biochem. Biophys. Res. Commun. 2002, 290, 690–695. [Google Scholar] [CrossRef]
- Lu, Y.; Li, Y.; Yang, Q.; Zhang, Z.; Chen, Y.; Zhang, S.; Peng, X.-X. Suppression of Glycolate Oxidase Causes Glyoxylate Accumulation That Inhibits Photosynthesis through Deactivating Rubisco in Rice. Physiol. Plant 2014, 150, 463–476. [Google Scholar] [CrossRef]
- Zhang, Y.; Xiao, W.; Luo, L.; Pang, J.; Rong, W.; He, C. Downregulation of OsPK1, a Cytosolic Pyruvate Kinase, by T-DNA Insertion Causes Dwarfism and Panicle Enclosure in Rice. Planta 2012, 235, 25–38. [Google Scholar] [CrossRef]
- Zhang, Y.; Feng, F.; He, C. Downregulation of OsPK1 Contributes to Oxidative Stress and the Variations in ABA/GA Balance in Rice. Plant Mol. Biol. Rep. 2012, 30, 1006–1013. [Google Scholar] [CrossRef]
- Lee, S.-K.; Hwang, S.-K.; Han, M.; Eom, J.-S.; Kang, H.-G.; Han, Y.; Choi, S.-B.; Cho, M.-H.; Bhoo, S.H.; An, G.; et al. Identification of the ADP-Glucose Pyrophosphorylase Isoforms Essential for Starch Synthesis in the Leaf and Seed Endosperm of Rice (Oryza Sativa L.). Plant Mol. Biol. 2007, 65, 531–546. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Cai, H.; Xiao, J.; Li, X.; Zhang, Q.; Lian, X. Over-Expression of Aspartate Aminotransferase Genes in Rice Resulted in Altered Nitrogen Metabolism and Increased Amino Acid Content in Seeds. Theor. Appl. Genet. 2009, 118, 1381–1390. [Google Scholar] [CrossRef] [PubMed]
- Hakata, M.; Kuroda, M.; Miyashita, T.; Yamaguchi, T.; Kojima, M.; Sakakibara, H.; Mitsui, T.; Yamakawa, H. Suppression of α-Amylase Genes Improves Quality of Rice Grain Ripened under High Temperature. Plant Biotechnol. J. 2012, 10, 1110–1117. [Google Scholar] [CrossRef] [Green Version]
- Lin, D.-G.; Chou, S.-Y.; Wang, A.Z.; Wang, Y.-W.; Kuo, S.-M.; Lai, C.-C.; Chen, L.-J.; Wang, C.-S. A Proteomic Study of Rice Cultivar TNG67 and Its High Aroma Mutant SA0420. Plant Sci. 2014, 214, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Kuwano, M.; Takaiwa, F.; Yoshida, K.T. Differential Effects of a Transgene to Confer Low Phytic Acid in Caryopses Located at Different Positions in Rice Panicles. Plant Cell Physiol. 2009, 50, 1387–1392. [Google Scholar] [CrossRef] [Green Version]
- Kuwano, M.; Mimura, T.; Takaiwa, F.; Yoshida, K.T. Generation of Stable ‘Low Phytic Acid’ Transgenic Rice through Antisense Repression of the 1d-Myo-Inositol 3-Phosphate Synthase Gene (RINO1) Using the 18-KDa Oleosin Promoter. Plant Biotechnol. J. 2009, 7, 96–105. [Google Scholar] [CrossRef]
- Kim, S.I.; Andaya, C.B.; Newman, J.W.; Goyal, S.S.; Tai, T.H. Isolation and Characterization of a Low Phytic Acid Rice Mutant Reveals a Mutation in the Rice Orthologue of Maize MIK. Theor. Appl. Genet. 2008, 117, 1291. [Google Scholar] [CrossRef]
- Satoh, H.; Shibahara, K.; Tokunaga, T.; Nishi, A.; Tasaki, M.; Hwang, S.-K.; Okita, T.W.; Kaneko, N.; Fujita, N.; Yoshida, M.; et al. Mutation of the Plastidial α-Glucan Phosphorylase Gene in Rice Affects the Synthesis and Structure of Starch in the Endosperm. Plant Cell 2008, 20, 1833–1849. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Utsumi, Y.; Utsumi, C.; Sawada, T.; Fujita, N.; Nakamura, Y. Functional Diversity of Isoamylase Oligomers: The ISA1 Homo-Oligomer Is Essential for Amylopectin Biosynthesis in Rice Endosperm. Plant Physiol. 2011, 156, 61–77. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kang, H.-G.; Park, S.; Matsuoka, M.; An, G. White-Core Endosperm Floury Endosperm-4 in Rice Is Generated by Knockout Mutations in the C4-Type Pyruvate Orthophosphate Dikinase Gene (OsPPDKB). Plant J. 2005, 42, 901–911. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.-Y.; Zheng, F.-Q.; Shen, G.-Z.; Gao, J.-P.; Snustad, D.P.; Li, M.-G.; Zhang, J.-L.; Hong, M.-M. The Amylose Content in Rice Endosperm Is Related to the Post-Transcriptional Regulation of the Waxy Gene. Plant J. 1995, 7, 613–622. [Google Scholar] [CrossRef]
- Fujita, N.; Yoshida, M.; Asakura, N.; Ohdan, T.; Miyao, A.; Hirochika, H.; Nakamura, Y. Function and Characterization of Starch Synthase I Using Mutants in Rice. Plant Physiol. 2006, 140, 1070–1084. [Google Scholar] [CrossRef] [Green Version]
- Zhang, G.; Cheng, Z.; Zhang, X.; Guo, X.; Su, N.; Jiang, L.; Mao, L.; Wan, J. Double Repression of Soluble Starch Synthase Genes SSIIa and SSIIIa in Rice (Oryza Sativa L.) Uncovers Interactive Effects on the Physicochemical Properties of Starch. Genome 2011, 54, 448–459. [Google Scholar] [CrossRef]
- Gao, Z.; Zeng, D.; Cheng, F.; Tian, Z.; Guo, L.; Su, Y.; Yan, M.; Jiang, H.; Dong, G.; Huang, Y.; et al. ALK, the Key Gene for Gelatinization Temperature, Is a Modifier Gene for Gel Consistency in RiceF. J. Integr. Plant Biol. 2011, 53, 756–765. [Google Scholar] [PubMed]
- Ryoo, N.; Yu, C.; Park, C.-S.; Baik, M.-Y.; Park, I.M.; Cho, M.-H.; Bhoo, S.H.; An, G.; Hahn, T.-R.; Jeon, J.-S. Knockout of a Starch Synthase Gene OsSSIIIa/Flo5 Causes White-Core Floury Endosperm in Rice (Oryza Sativa L.). Plant Cell Rep. 2007, 26, 1083–1095. [Google Scholar] [CrossRef] [PubMed]
- Fujita, N.; Satoh, R.; Hayashi, A.; Kodama, M.; Itoh, R.; Aihara, S.; Nakamura, Y. Starch Biosynthesis in Rice Endosperm Requires the Presence of Either Starch Synthase I or IIIa. J. Exp. Bot. 2011, 62, 4819–4831. [Google Scholar] [CrossRef] [Green Version]
- Yun, M.-S.; Umemoto, T.; Kawagoe, Y. Rice Debranching Enzyme Isoamylase3 Facilitates Starch Metabolism and Affects Plastid Morphogenesis. Plant Cell Physiol. 2011, 52, 1068–1082. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saito, H.; Okumoto, Y.; Yoshitake, Y.; Inoue, H.; Yuan, Q.; Teraishi, M.; Tsukiyama, T.; Nishida, H.; Tanisaka, T. Complete Loss of Photoperiodic Response in the Rice Mutant Line X61 Is Caused by Deficiency of Phytochrome Chromophore Biosynthesis Gene. Theor. Appl. Genet. 2011, 122, 109–118. [Google Scholar] [CrossRef]
- Jia, L.; Zhang, B.; Mao, C.; Li, J.; Wu, Y.; Wu, P.; Wu, Z. OsCYT-INV1 for Alkaline/Neutral Invertase Is Involved in Root Cell Development and Reproductivity in Rice (Oryza Sativa L.). Planta 2008, 228, 51–59. [Google Scholar] [CrossRef] [PubMed]
- Asatsuma, S.; Sawada, C.; Itoh, K.; Okito, M.; Kitajima, A.; Mitsui, T. Involvement of α-Amylase I-1 in Starch Degradation in Rice Chloroplasts. Plant Cell Physiol. 2005, 46, 858–869. [Google Scholar] [CrossRef]
- Ibraheem, O.; Botha, C.E.J.; Bradley, G.; Dealtry, G.; Roux, S. Rice Sucrose Transporter1 (OsSUT1) up-Regulation in Xylem Parenchyma Is Caused by Aphid Feeding on Rice Leaf Blade Vascular Bundles. Plant Biol. 2014, 16, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Wang, K.; Li, Y.; Tan, Y.; Kong, J.; Li, H.; Li, Y.; Zhu, Y. Overexpression of SBPase Enhances Photosynthesis against High Temperature Stress in Transgenic Rice Plants. Plant Cell Rep. 2007, 26, 1635–1646. [Google Scholar] [CrossRef]
- Ye, N.; Yang, G.; Chen, Y.; Zhang, C.; Zhang, J.; Peng, X. Two Hydroxypyruvate Reductases Encoded by OsHPR1 and OsHPR2 Are Involved in Photorespiratory Metabolism in Rice. J. Integr. Plant Biol. 2014, 56, 170–180. [Google Scholar] [CrossRef]
- Cook, F.R.; Fahy, B.; Trafford, K. A Rice Mutant Lacking a Large Subunit of ADP-Glucose Pyrophosphorylase Has Drastically Reduced Starch Content in the Culm but Normal Plant Morphology and Yield. Funct. Plant Biol. 2012, 39, 1068–1078. [Google Scholar] [CrossRef]
- Hirose, T.; Hashida, Y.; Aoki, N.; Okamura, M.; Yonekura, M.; Ohto, C.; Terao, T.; Ohsugi, R. Analysis of Gene-Disruption Mutants of a Sucrose Phosphate Synthase Gene in Rice, OsSPS1, Shows the Importance of Sucrose Synthesis in Pollen Germination. Plant Sci. 2014, 225, 102–106. [Google Scholar] [CrossRef]
- Cao, H.; Guo, S.; Xu, Y.; Jiang, K.; Jones, A.M.; Chong, K. Reduced Expression of a Gene Encoding a Golgi Localized Monosaccharide Transporter (OsGMST1) Confers Hypersensitivity to Salt in Rice (Oryza sativa). J. Exp. Bot. 2011, 62, 4595–4604. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siahpoosh, M.R.; Sanchez, D.H.; Schlereth, A.; Scofield, G.N.; Furbank, R.T.; van Dongen, J.T.; Kopka, J. Modification of OsSUT1 Gene Expression Modulates the Salt Response of Rice Oryza Sativa Cv. Taipei 309. Plant Sci. 2012, 182, 101–111. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-H.; Rao, X.-L.; Shi, H.-T.; Li, R.-J.; Lu, Y.-T. Overexpression of a Cytosolic Glyceraldehyde-3-Phosphate Dehydrogenase Gene OsGAPC3 Confers Salt Tolerance in Rice. Plant Cell Tissue Organ Cult. 2011, 107, 1. [Google Scholar] [CrossRef]
- Tuncel, A.; Kawaguchi, J.; Ihara, Y.; Matsusaka, H.; Nishi, A.; Nakamura, T.; Kuhara, S.; Hirakawa, H.; Nakamura, Y.; Cakir, B.; et al. The Rice Endosperm ADP-Glucose Pyrophosphorylase Large Subunit Is Essential for Optimal Catalysis and Allosteric Regulation of the Heterotetrameric Enzyme. Plant Cell Physiol. 2014, 55, 1169–1183. [Google Scholar] [CrossRef] [Green Version]
- Masumoto, C.; Miyazawa, S.-I.; Ohkawa, H.; Fukuda, T.; Taniguchi, Y.; Murayama, S.; Kusano, M.; Saito, K.; Fukayama, H.; Miyao, M. Phosphoenolpyruvate Carboxylase Intrinsically Located in the Chloroplast of Rice Plays a Crucial Role in Ammonium Assimilation. Proc. Natl. Acad. Sci. USA 2010, 107, 5226–5231. [Google Scholar] [CrossRef] [Green Version]
- Lee, S.-K.; Jeon, J.-S.; Bornke, F.; Voll, L.; Cho, J.-I.; Goh, C.-H.; Jeong, S.-W.; Park, Y.-I.; Kim, S.J.; Choi, S.-B.; et al. Loss of Cytosolic Fructose-1,6-Bisphosphatase Limits Photosynthetic Sucrose Synthesis and Causes Severe Growth Retardations in Rice (Oryza sativa). Plant Cell Environ. 2008, 31, 1851–1863. [Google Scholar] [CrossRef]
- Kasajima, I.; Ebana, K.; Yamamoto, T.; Takahara, K.; Yano, M.; Kawai-Yamada, M.; Uchimiya, H. Molecular Distinction in Genetic Regulation of Nonphotochemical Quenching in Rice. Proc. Natl. Acad. Sci. USA 2011, 108, 13835–13840. [Google Scholar] [CrossRef] [Green Version]
- Hubbart, S.; Ajigboye, O.O.; Horton, P.; Murchie, E.H. The Photoprotective Protein PsbS Exerts Control over CO2 Assimilation Rate in Fluctuating Light in Rice. Plant J. 2012, 71, 402–412. [Google Scholar] [CrossRef]
- Abe, N.; Asai, H.; Yago, H.; Oitome, N.F.; Itoh, R.; Crofts, N.; Nakamura, Y.; Fujita, N. Relationships between Starch Synthase I and Branching Enzyme Isozymes Determined Using Double Mutant Rice Lines. BMC Plant Biol. 2014, 14, 80. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, H.; Zhang, J.; Zeng, J.; Jiang, L.; Liu, E.; Peng, C.; He, Z.; Peng, X. Inducible Antisense Suppression of Glycolate Oxidase Reveals Its Strong Regulation over Photosynthesis in Rice. J. Exp. Bot. 2009, 60, 1799–1809. [Google Scholar] [CrossRef] [Green Version]
- Jin, S.-H.; Hong, J.; Li, X.-Q.; Jiang, D.-A. Antisense Inhibition of Rubisco Activase Increases Rubisco Content and Alters the Proportion of Rubisco Activase in Stroma and Thylakoids in Chloroplasts of Rice Leaves. Ann. Bot. 2006, 97, 739–744. [Google Scholar] [CrossRef] [Green Version]
- Nath, K.; Poudyal, R.S.; Eom, J.-S.; Park, Y.S.; Zulfugarov, I.S.; Mishra, S.R.; Tovuu, A.; Ryoo, N.; Yoon, H.-S.; Nam, H.G.; et al. Loss-of-Function of OsSTN8 Suppresses the Photosystem II Core Protein Phosphorylation and Interferes with the Photosystem II Repair Mechanism in Rice (Oryza sativa). Plant J. 2013, 76, 675–686. [Google Scholar] [CrossRef] [PubMed]
- Nishikawa, Y.; Yamamoto, H.; Okegawa, Y.; Wada, S.; Sato, N.; Taira, Y.; Sugimoto, K.; Makino, A.; Shikanai, T. PGR5-Dependent Cyclic Electron Transport Around PSI Contributes to the Redox Homeostasis in Chloroplasts Rather Than CO2 Fixation and Biomass Production in Rice. Plant Cell Physiol. 2012, 53, 2117–2126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, Q.; Yu, Q.; Wang, Z.; Pan, Y.; Lv, W.; Zhu, L.; Chen, R.; He, G. Knockdown of GDCH Gene Reveals Reactive Oxygen Species-Induced Leaf Senescence in Rice. Plant Cell Environ. 2013, 36, 1476–1489. [Google Scholar] [CrossRef]
- Makino, A.; Shimada, T.; Takumi, S.; Kaneko, K.; Matsuoka, M.; Shimamoto, K.; Nakano, H.; Miyao-Tokutomi, M.; Mae, T.; Yamamoto, N. Does Decrease in Ribulose-1,5-Bisphosphate Carboxylase by Antisense RbcS Lead to a Higher N-Use Efficiency of Photosynthesis under Conditions of Saturating CO2 and Light in Rice Plants? Plant Physiol. 1997, 114, 483–491. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogawa, S.; Suzuki, Y.; Yoshizawa, R.; Kanno, K.; Makino, A. Effect of Individual Suppression of RBCS Multigene Family on Rubisco Contents in Rice Leaves. Plant Cell Environ. 2012, 35, 546–553. [Google Scholar] [CrossRef]
- Makino, A.; Sage, R.F. Temperature Response of Photosynthesis in Transgenic Rice Transformed with ‘Sense’ or ‘Antisense’ RbcS. Plant Cell Physiol. 2007, 48, 1472–1483. [Google Scholar] [CrossRef] [Green Version]
- Hong, W.-J.; Yoo, Y.-H.; Park, S.-A.; Moon, S.; Kim, S.-R.; An, G.; Jung, K.-H. Genome-Wide Identification and Extensive Analysis of Rice-Endosperm Preferred Genes Using Reference Expression Database. J. Plant Biol. 2017, 60, 249–258. [Google Scholar] [CrossRef]
- Xu, F.-Q.; Li, X.-R.; Ruan, Y.-L. RNAi-Mediated Suppression of Hexokinase Gene OsHXK10 in Rice Leads to Non-Dehiscent Anther and Reduction of Pollen Germination. Plant Sci. 2008, 175, 674–684. [Google Scholar] [CrossRef]
- Oliver, S.N.; Van Dongen, J.T.; Alfred, S.C.; Mamun, E.A.; Zhao, X.; Saini, H.S.; Fernandes, S.F.; Blanchard, C.L.; Sutton, B.G.; Geigenberger, P.; et al. Cold-Induced Repression of the Rice Anther-Specific Cell Wall Invertase Gene OSINV4 Is Correlated with Sucrose Accumulation and Pollen Sterility. Plant Cell Environ. 2005, 28, 1534–1551. [Google Scholar] [CrossRef]
- Fan, C.; Wang, G.; Wu, L.; Liu, P.; Huang, J.; Jin, X.; Zhang, G.; He, Y.; Peng, L.; Luo, K.; et al. Distinct Cellulose and Callose Accumulation for Enhanced Bioethanol Production and Biotic Stress Resistance in OsSUS3 Transgenic Rice. Carbohydr. Polym. 2020, 232, 115448. [Google Scholar] [CrossRef]
- Chen, W.; Gong, P.; Guo, J.; Li, H.; Li, R.; Xing, W.; Yang, Z.; Guan, Y. Glycolysis Regulates Pollen Tube Polarity via Rho GTPase Signaling. PLoS Genet. 2018, 14, e1007373. [Google Scholar] [CrossRef] [Green Version]
- You, C.; Zhu, H.; Xu, B.; Huang, W.; Wang, S.; Ding, Y.; Liu, Z.; Li, G.; Chen, L.; Ding, C.; et al. Effect of Removing Superior Spikelets on Grain Filling of Inferior Spikelets in Rice. Front. Plant Sci. 2016, 7, 1161. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Zhang, D.; Miao, Q.; Yang, J.; Xuan, Y.; Hu, Y. Essential Role of Sugar Transporter OsSWEET11 During the Early Stage of Rice Grain Filling. Plant Cell Physiol. 2017, 58, 863–873. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.; Luo, D.; Yang, B.; Frommer, W.B.; Eom, J.-S. SWEET11 and 15 as Key Players in Seed Filling in Rice. New Phytol. 2018, 218, 604–615. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tabuchi, M.; Sugiyama, K.; Ishiyama, K.; Inoue, E.; Sato, T.; Takahashi, H.; Yamaya, T. Severe Reduction in Growth Rate and Grain Filling of Rice Mutants Lacking OsGS1;1, a Cytosolic Glutamine Synthetase1;1. Plant J. 2005, 42, 641–651. [Google Scholar] [CrossRef]
- Samuel, A.D.; Bungau, S.; Tit, D.M.; Melinte, C.E.; Purza, L.; Badea, G.E. Effects of Long Term Application of Organic and Mineral Fertilizers on Soil Enzymes. Rev. Chim. 2018, 69, 2608–2612. [Google Scholar] [CrossRef]
- Ramakrishna, W.; Yadav, R.; Li, K. Plant Growth Promoting Bacteria in Agriculture: Two Sides of a Coin. Appl. Soil Ecol. 2019, 138, 10–18. [Google Scholar] [CrossRef]
- Preece, C.; Peñuelas, J. A Return to the Wild: Root Exudates and Food Security. Trends Plant Sci. 2020, 25, 14–21. [Google Scholar] [CrossRef] [PubMed]


| LOC_id | Symbol | Character_major 1 | Character_minor | Method 2 | Detailed Functions | Reference |
|---|---|---|---|---|---|---|
| Os02g44230 | OsTPP1 | R or T | Cold tolerance | OX | Cold and salinity tolerance | [37] |
| Os05g44210 | OsTPS1 | R or T | Cold tolerance | OX | Cold, drought, and salinity tolerance | [38] |
| Os01g64660 | moc2 | MT | Culm leaf | KO | Tiller bud outgrowth; tillering | [39] |
| Os12g44380 | ossut2 | MT | Culm leaf | KO | Sugar export; growth retardation | [40] |
| Os05g44210 | OsTPS1 | R or T | Drought tolerance | OX | Cold, drought, and salinity tolerance | [38] |
| Os06g36560 | OsMIOX | R or T | Drought tolerance | OX | Drought tolerance | [41] |
| Os03g07480 | OsSUT1 | MT | Dwarf | KD | Dwarfism; flowering time | [42] |
| Os04g56320 | OsrcaA2 | MT | Dwarf | KD/OX | Dwarfism | [43] |
| Os07g05820 | OsGLO4 | R or T | Dwarf | KD | Growth inhibition | [44] |
| Os11g05110 | OsPK1 | MT | Dwarf | KO | Elongation of uppermost internode; dwarfism | [45] |
| Os11g05110 | ospk1 | MT | Dwarf | KO | Dwarfism; seed color; internode color; regulation of gibberellin and ABA biosynthesis | [46] |
| Os12g44380 | ossut2 | MT | Dwarf | KO | Sugar export; growth retardation | [40] |
| Os01g44220 | osagpl2 | PT | Eating quality | KO | Starch biosynthesis | [47] |
| Os01g55540 | OsAAT2 | PT | Eating quality | OX | Seed amino acid and protein content | [48] |
| Os02g52710 | Amy1A | PT | Eating quality | KD | Seed starch content; high temperature-triggered grain chalkiness | [49] |
| Os03g03720 | OsGAPDHB | PT | Eating quality | KD | Fragrance rice | [50] |
| Os03g09250 | RINO1 | PT | Eating quality | KD | Seed phytic acid content | [51] |
| Os03g09250 | RINO1 | PT | Eating quality | KD | Seed phytic acid content | [52] |
| Os03g52760 | lpaN15-186 | PT | Eating quality | KO | Seed phytic acid content | [53] |
| Os03g55090 | pho1 | PT | Eating quality | KO | Seed starch content; grain maturation | [54] |
| Os05g32710 | OsISA2 | PT | Eating quality | OX | Seed starch content | [55] |
| Os05g33570 | flo4 | PT | Eating quality | KO | Seed protein and lipid content | [56] |
| Os06g04200 | wx | PT | Eating quality | NV | Seed amylose content | [57] |
| Os06g06560 | OsSSI | PT | Eating quality | KO | Seed amylopectin content | [58] |
| Os06g12450 | SSIIa | PT | Eating quality | KD | Chalky kernel; amylose content | [59] |
| Os06g12450 | ALK | PT | Eating quality | NV | Gelatinization temperature; gel consistency | [60] |
| Os08g09230 | SSIIIa | PT | Eating quality | KD | Chalky kernel; amylose content | [59] |
| Os08g09230 | flo5 | PT | Eating quality | KO | Seed starch content | [61] |
| Os08g09230 | ss3a | PT | Eating quality | KO | Seed starch content | [62] |
| Os08g25734 | osagps2 | PT | Eating quality | KO | Seed starch content | [47] |
| Os08g40930 | OsISA1 | PT | Eating quality | OX | Seed starch content | [55] |
| Os09g28400 | Amy3A | PT | Eating quality | KD | Seed starch content; high temperature-triggered grain chalkiness | [49] |
| Os09g28420 | Amy3B | PT | Eating quality | KD | Seed starch content; high temperature-triggered grain chalkiness | [49] |
| Os09g29404 | isa3 | PT | Eating quality | KO | Seed starch content | [63] |
| Os01g72090 | se13 | PT | Flowering | KO | Photoperiodic response | [64] |
| Os02g34560 | Oscyt-inv1 | PT | Flowering | KO | Root cell development; flowering time; fertility | [65] |
| Os03g07480 | OsSUT1 | PT | Flowering | KD | Dwarfism; flowering time | [42] |
| Os02g52710 | AmyI-1 | PT | Germination dormancy | KD/OX | Starch degradation; seed germination; seedling growth | [66] |
| Os03g07480 | OsSUT1 | R or T | Insect resistance | Others | Upregulation by aphid feeding; transfer sucrose | [67] |
| Os02g47020 | SBPase | R or T | Other stress resistance | OX | Photosynthetic ability under high-temperature condition | [68] |
| Os02g01150 | OsHPR1 | PT | Others | KD/OX | Photorespiratory pathway | [69] |
| Os05g50380 | LSU3 | Others | Others | KO | Culm starch content | [70] |
| Os01g69030 | OsSPS1 | PT | Panicle flower | KO | Pollen germination through sucrose synthesis | [71] |
| Os02g34560 | Oscyt-inv1 | MT | Root | KO | Root cell development; flowering time; fertility | [65] |
| Os12g44380 | ossut2 | MT | Root | KO | Sugar export; growth retardation | [40] |
| Os02g17500 | OsGMST1 | R or T | Salinity tolerance | KD | Salinity tolerance | [72] |
| Os02g44230 | OsTPP1 | R or T | Salinity tolerance | OX | Cold and salinity tolerance | [37] |
| Os03g07480 | OsSUT1 | R or T | Salinity tolerance | KD | Salinity tolerance | [73] |
| Os05g44210 | OsTPS1 | R or T | Salinity tolerance | OX | Cold, drought, and salinity tolerance | [38] |
| Os08g03290 | OsGAPC3 | R or T | Salinity tolerance | OX | Salinity tolerance | [74] |
| Os01g44220 | OsAGPL2 | MT | Seed | KO | Glassy/vitreous, shrunken grain | [75] |
| Os03g55090 | pho1 | MT | Seed | KO | Seed starch content; grain maturation | [54] |
| Os04g33740 | GIF1 | MT | Seed | NV | Grain filling; grain size | [12] |
| Os08g25734 | OsAGPS2b | PT | Seed | KO | Seed weight; starch content; AGPase activities from developing endosperms of the seed | [75] |
| Os11g05110 | ospk1 | MT | Seed | KO | Dwarfism; seed color; internode color; Regulation of gibberellin and ABA biosynthesis | [46] |
| Os02g52710 | AmyI-1 | MT | Shoot seedling | KD/OX | Starch degradation; seed germination; seedling growth | [66] |
| Os01g11054 | Osppc4 | PT | Source activity | KD | Ammonium assimilation in leaves | [76] |
| Os01g64660 | oscfbp1 | PT | Source activity | KO | Photosynthetic sucrose biosynthesis; growth retardation | [77] |
| Os01g64960 | qNPQ1-2 | PT | Source activity | NV | Nonphotochemical quenching capacity; protection from photoinhibition | [78] |
| Os01g64960 | PsbS | PT | Source activity | KD/OX | Nonphotochemical quenching capacity; photosynthetic rate in fluctuating light conditions | [79] |
| Os02g32660 | BE2b | PT | Source activity | Others | Starch biosynthesis in endosperm; amylopectin biosynthesis; branch formation | [80] |
| Os03g57220 | OsGLO1 | PT | Source activity | KD | Photorespiration | [81] |
| Os04g56320 | rca | PT | Source activity | KD | Rubisco activity | [82] |
| Os05g40180 | OsSTN8 | PT | Source activity | KO | Photosystem II repair during high light illumination | [83] |
| Os06g06560 | SS1 | PT | Source activity | Others | Starch biosynthesis in endosperm; chain elongation | [80] |
| Os06g51084 | BE1 | PT | Source activity | Others | Starch granule binding; amylopectin structure | [80] |
| Os07g05820 | OsGLO4 | PT | Source activity | KD | Rubisco activation; photosynthesis rate | [44] |
| Os08g45190 | PGR5 | PT | Source activity | KD | Photosynthetic capacity | [84] |
| Os10g37180 | OsGDCH | PT | Source activity | KD | Leaf senescence induced by reactive oxygen species | [85] |
| Os12g17600 | rbcS | PT | Source activity | KD | Rubisco content; photosynthetic capacity | [86] |
| Os12g17600 | OsRBCS2 | PT | Source activity | KD | Rubisco content | [87] |
| Os12g17600 | rbcS | PT | Source activity | KD/OX | Rubisco content; photosynthetic capacity | [88] |
| Os12g19381 | OsRBCS5 | PT | Source activity | KD | Rubisco content | [87] |
| Os12g19470 | OsRBCS4 | PT | Source activity | KD | Rubisco content | [87] |
| Os12g44380 | ossut2 | PT | Source activity | KO | Sugar export; growth retardation | [40] |
| Os02g34560 | Oscyt-inv1 | PT | Sterility | KO | Root cell development; flowering time; fertility | [65] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hong, W.-J.; Jiang, X.; Choi, S.-H.; Kim, Y.-J.; Kim, S.-T.; Jeon, J.-S.; Jung, K.-H. A Systemic View of Carbohydrate Metabolism in Rice to Facilitate Productivity. Plants 2021, 10, 1690. https://doi.org/10.3390/plants10081690
Hong W-J, Jiang X, Choi S-H, Kim Y-J, Kim S-T, Jeon J-S, Jung K-H. A Systemic View of Carbohydrate Metabolism in Rice to Facilitate Productivity. Plants. 2021; 10(8):1690. https://doi.org/10.3390/plants10081690
Chicago/Turabian StyleHong, Woo-Jong, Xu Jiang, Seok-Hyun Choi, Yu-Jin Kim, Sun-Tae Kim, Jong-Seong Jeon, and Ki-Hong Jung. 2021. "A Systemic View of Carbohydrate Metabolism in Rice to Facilitate Productivity" Plants 10, no. 8: 1690. https://doi.org/10.3390/plants10081690






