Transcriptomic Analysis of Radish (Raphanus sativus L.) Roots with CLE41 Overexpression
Abstract
:1. Introduction
2. Results
2.1. RNA-Sequencing of Transgenic Radish Roots with RsCLE41 Overexpression and Those with GUS Overexpression
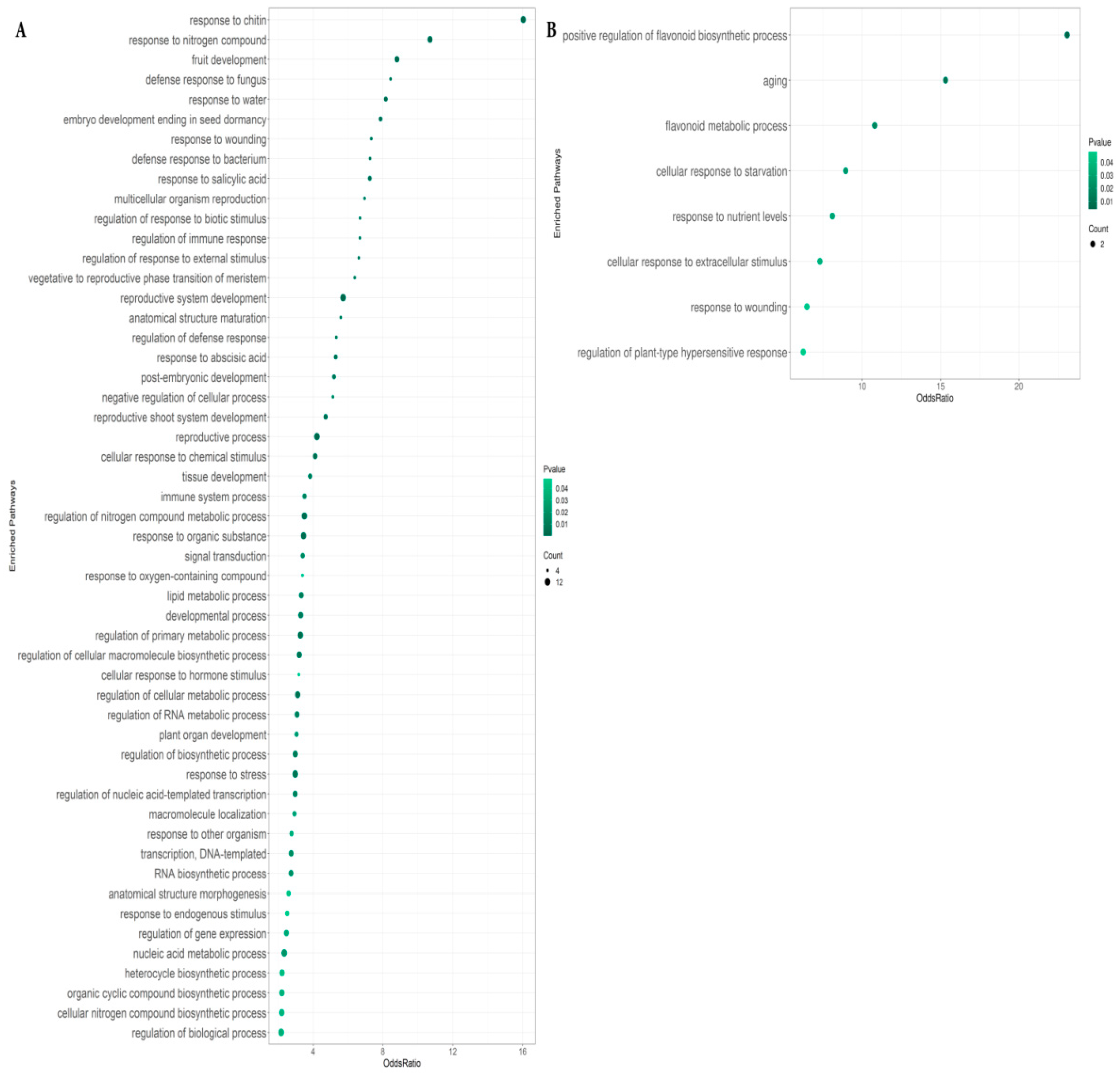
2.2. Enriched Pathways in Genes Differentially Expressed between Roots with RsCLE41 Overexpression and Those with GUS Overexpression
2.3. Individual Transcripts Differentially Expressed between Roots with RsCLE41 Overexpression and Those with GUS Overexpression
2.3.1. Genes Involved in Late Embryogenesis: Strongly Upregulated
2.3.2. Genes Governing Xylem Formation: Upregulation of Genes Involved in Early Stages of Xylem Specification and Downregulation of Genes Participating in Late Stages of Vessel Development
2.3.3. Stress-Related Genes: Upregulation of TF-Encoding Genes Involved in the Response to ABA, Ethylene and Jasmonates and Downregulation of Genes Acting in Disease Resistance
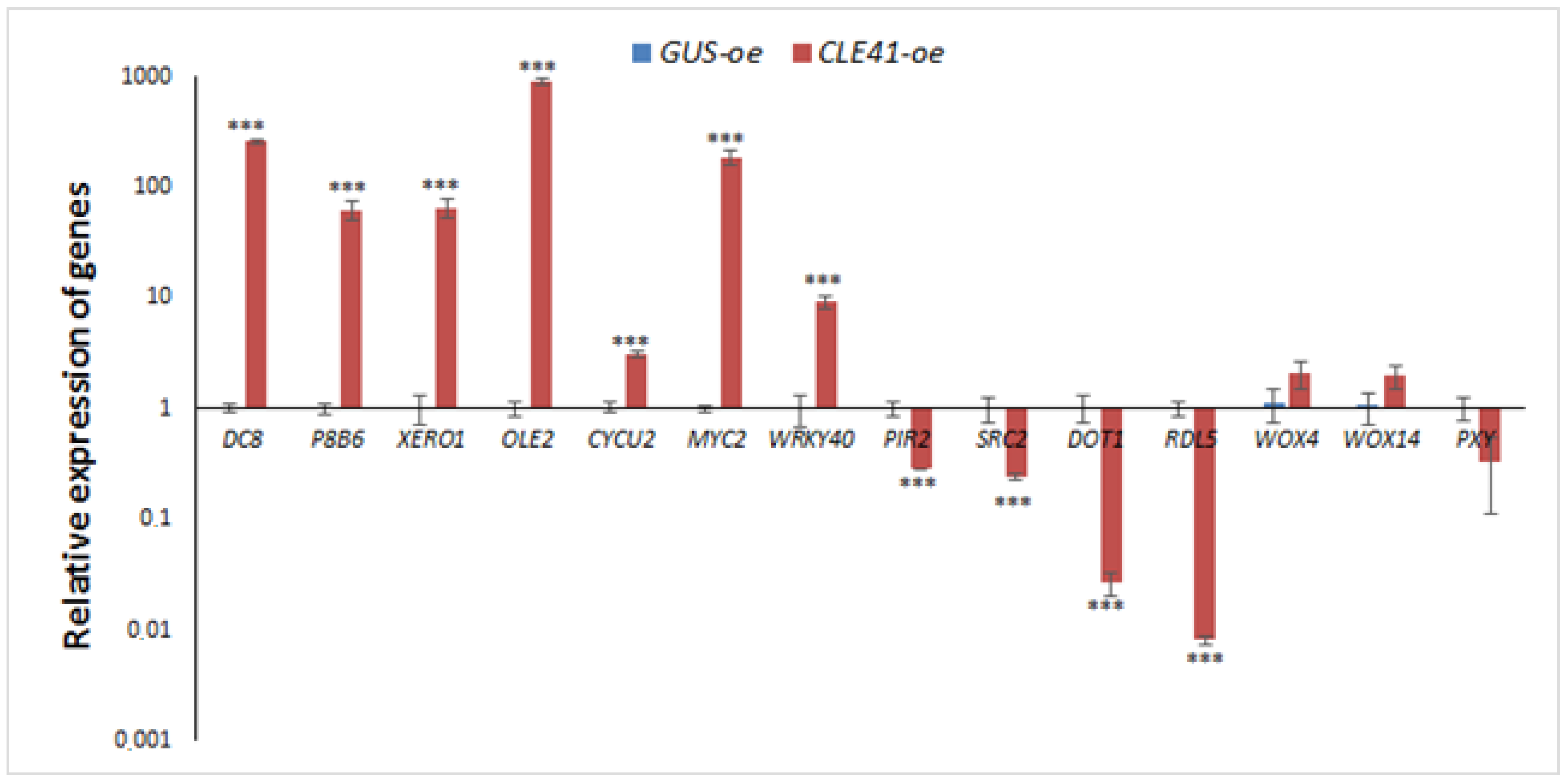
2.4. Verification of Differential Expression Analysis Data with qPCR
3. Discussion
4. Materials and Methods
4.1. Construction of Vector
4.2. Plant Growth, Transformation and Sample Collection
4.3. RNA Isolation, Library Preparation and Sequencing
4.4. Bioinformatics Processing of Sequencing Results
4.5. qPCR Expression Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Stahl, Y.; Wink, R.H.; Ingram, G.C.; Simon, R. A Signaling Module Controlling the Stem Cell Niche in Arabidopsis Root Meristems. Curr. Biol. 2009, 19, 909–914. [Google Scholar] [CrossRef] [Green Version]
- Ito, M. Conservation and Diversification of Three-Repeat Myb Transcription Factors in Plants. J. Plant Res. 2005, 118, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Etchells, J.P.; Turner, S.R. The PXY-CLE41 Receptor Ligand Pair Defines a Multifunctional Pathway That Controls the Rate and Orientation of Vascular Cell Division. Development 2010, 137, 767–774. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hirakawa, Y.; Shinohara, H.; Kondo, Y.; Inoue, A.; Nakanomyo, I.; Ogawa, M.; Sawa, S.; Ohashi-Ito, K.; Matsubayashi, Y.; Fukuda, H. Non-Cell-Autonomous Control of Vascular Stem Cell Fate by a CLE Peptide/Receptor System. Proc. Natl. Acad. Sci. USA 2008, 105, 15208–15213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fisher, K.; Turner, S. PXY, a Receptor-like Kinase Essential for Maintaining Polarity during Plant Vascular-Tissue Development. Curr. Biol. 2007, 17, 1061–1066. [Google Scholar] [CrossRef] [PubMed]
- Etchells, J.P.; Provost, C.M.; Mishra, L.; Turner, S.R. WOX4 and WOX14 Act Downstream of the PXY Receptor Kinase to Regulate Plant Vascular Proliferation Independently of Any Role in Vascular Organisation. Development 2013, 140, 2224–2234. [Google Scholar] [CrossRef] [Green Version]
- Kondo, Y.; Ito, T.; Nakagami, H.; Hirakawa, Y.; Saito, M.; Tamaki, T.; Shirasu, K.; Fukuda, H. Plant GSK3 Proteins Regulate Xylem Cell Differentiation Downstream of TDIF-TDR Signalling. Nat. Commun. 2014, 5, 3504. [Google Scholar] [CrossRef] [Green Version]
- Han, S.; Cho, H.; Noh, J.; Qi, J.; Jung, H.-J.; Nam, H.; Lee, S.; Hwang, D.; Greb, T.; Hwang, I. BIL1-Mediated MP Phosphorylation Integrates PXY and Cytokinin Signalling in Secondary Growth. Nat. Plants 2018, 4, 605–614. [Google Scholar] [CrossRef]
- Gancheva, M.S.; Dodueva, I.E.; Lebedeva, M.A.; Tvorogova, V.E.; Tkachenko, A.A.; Lutova, L.A. Identification, Expression, and Functional Analysis of CLE Genes in Radish (Raphanus sativus L.) Storage Root. BMC Plant Biol. 2016, 16, 7. [Google Scholar] [CrossRef] [Green Version]
- Jeong, Y.M.; Kim, N.; Ahn, B.O.; Oh, M.; Chung, W.H.; Chung, H.; Jeong, S.; Lim, K.B.; Hwang, Y.J.; Kim, G.B.; et al. Elucidating the triplicated ancestral genome structure of radish based on chromosome-level comparison with the Brassica genomes. Theor. Appl. Genet. 2016, 129, 1357–1372. [Google Scholar] [CrossRef]
- Gancheva, M.S.; Dodueva, I.E.; Lutova, L.A. Role of CLE41 Peptide in the Development of Root Storage Parenchyma in Species of the Genus Raphanus L. Russ. J. Plant Physiol. 2018, 65, 498–511. [Google Scholar] [CrossRef]
- Whitford, R.; Fernandez, A.; De Groodt, R.; Ortega, E.; Hilson, P. Plant CLE Peptides from Two Distinct Functional Classes Synergistically Induce Division of Vascular Cells. Proc. Natl. Acad. Sci. USA 2008, 105, 18625–18630. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smit, M.E.; McGregor, S.R.; Sun, H.; Gough, C.; Bågman, A.-M.; Soyars, C.L.; Kroon, J.T.; Gaudinier, A.; Williams, C.J.; Yang, X.; et al. A PXY-Mediated Transcriptional Network Integrates Signaling Mechanisms to Control Vascular Development in Arabidopsis. Plant Cell 2020, 32, 319–335. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, Y.; Tan, S.; Gao, Y.; Kan, C.; Wang, H.-L.; Yang, Q.; Xia, X.; Ishida, T.; Sawa, S.; Guo, H.; et al. CLE42 Delays Leaf Senescence by Antagonizing Ethylene Pathway in Arabidopsis. New Phytol. 2022, 235, 550–562. [Google Scholar] [CrossRef]
- Buzovkina, I.S.; Lutova, L.A. The Genetic Collection of Radish Inbred Lines: History and Prospects. Russ. J. Genet. 2007, 43, 1181–1192. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A Fast Spliced Aligner with Low Memory Requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [Green Version]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.-C.; Mendell, J.T.; Salzberg, S.L. StringTie Enables Improved Reconstruction of a Transcriptome from RNA-Seq Reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef] [Green Version]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the Unification of Biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [Green Version]
- GSEABase: Gene Set Enrichment Data Structures and Methods Version 1.52.1 from Bioconductor. Available online: https://rdrr.io/bioc/GSEABase/ (accessed on 29 July 2022).
- Chigri, F.; Flosdorff, S.; Pilz, S.; Kölle, E.; Dolze, E.; Gietl, C.; Vothknecht, U.C. The Arabidopsis Calmodulin-like Proteins AtCML30 and AtCML3 Are Targeted to Mitochondria and Peroxisomes, Respectively. Plant Mol. Biol. 2012, 78, 211–222. [Google Scholar] [CrossRef]
- Hromadová, D.; Soukup, A.; Tylová, E. Arabinogalactan Proteins in Plant Roots—An Update on Possible Functions. Front. Plant Sci. 2021, 12, 674010. [Google Scholar] [CrossRef] [PubMed]
- Xu, B.; Sathitsuksanoh, N.; Tang, Y.; Udvardi, M.K.; Zhang, J.-Y.; Shen, Z.; Balota, M.; Harich, K.; Zhang, P.Y.-H.; Zhao, B. Overexpression of AtLOV1 in Switchgrass Alters Plant Architecture, Lignin Content, and Flowering Time. PLoS ONE 2012, 7, e47399. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, L.; Henty-Ridilla, J.L.; Blanchoin, L.; Staiger, C.J. Profilin-Dependent Nucleation and Assembly of Actin Filaments Controls Cell Elongation in Arabidopsis. Plant Physiol. 2016, 170, 220–233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- de Lara Campos Arcuri, M.; Nunes-Laitz, A.V.; Lima, R.P.M.; Barreto, P.; Marinho, A.N.; Arruda, P.; Maia, I.G. Knockdown of Mitochondrial Uncoupling Proteins 1 and 2 (AtUCP1 and 2) in Arabidopsis Thaliana Impacts Vegetative Development and Fertility. Plant Cell Physiol. 2021, 62, 1630–1644. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Leal, D.; León-Martínez, G.; Abad-Vivero, U.; Vielle-Calzada, J.-P. Natural Variation in Epigenetic Pathways Affects the Specification of Female Gamete Precursors in Arabidopsis. Plant Cell 2015, 27, 1034–1045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hundertmark, M.; Hincha, D.K. LEA (Late Embryogenesis Abundant) Proteins and Their Encoding Genes in Arabidopsis Thaliana. BMC Genom. 2008, 9, 118. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Candat, A.; Paszkiewicz, G.; Neveu, M.; Gautier, R.; Logan, D.C.; Avelange-Macherel, M.-H.; Macherel, D. The Ubiquitous Distribution of Late Embryogenesis Abundant Proteins across Cell Compartments in Arabidopsis Offers Tailored Protection against Abiotic Stress. Plant Cell 2014, 26, 3148–3166. [Google Scholar] [CrossRef] [Green Version]
- Leprince, O.; Pellizzaro, A.; Berriri, S.; Buitink, J. Late Seed Maturation: Drying without Dying. J. Exp. Bot. 2017, 68, 827–841. [Google Scholar] [CrossRef] [Green Version]
- Finn, R.D.; Mistry, J.; Tate, J.; Coggill, P.; Heger, A.; Pollington, J.E.; Gavin, O.L.; Gunasekaran, P.; Ceric, G.; Forslund, K.; et al. The Pfam Protein Families Database. Nucleic Acids Res. 2010, 38, D211–D222. [Google Scholar] [CrossRef]
- Mao, Z.; Sun, W. Arabidopsis Seed-Specific Vacuolar Aquaporins Are Involved in Maintaining Seed Longevity under the Control of ABSCISIC ACID INSENSITIVE 3. J. Exp. Bot. 2015, 66, 4781–4794. [Google Scholar] [CrossRef] [Green Version]
- Hanano, A.; Burcklen, M.; Flenet, M.; Ivancich, A.; Louwagie, M.; Garin, J.; Blée, E. Plant Seed Peroxygenase Is an Original Heme-Oxygenase with an EF-Hand Calcium Binding Motif. J. Biol. Chem. 2006, 281, 33140–33151. [Google Scholar] [CrossRef] [Green Version]
- Dietz, K.-J. Peroxiredoxins in Plants and Cyanobacteria. Antioxid. Redox Signal. 2011, 15, 1129–1159. [Google Scholar] [CrossRef] [Green Version]
- Růžička, K.; Ursache, R.; Hejátko, J.; Helariutta, Y. Xylem Development—From the Cradle to the Grave. New Phytol. 2015, 207, 519–535. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Liu, P.; Zhao, H.; Sun, J. The HD-ZIP II Transcription Factors Regulate Plant Architecture through the Auxin Pathway. Int. J. Mol. Sci. 2020, 21, 3250. [Google Scholar] [CrossRef] [PubMed]
- Izhaki, A.; Bowman, J.L. KANADI and Class III HD-Zip Gene Families Regulate Embryo Patterning and Modulate Auxin Flow during Embryogenesis in Arabidopsis. Plant Cell 2007, 19, 495–508. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, C.J.; Valdés, A.E.; Wang, G.; Ramachandran, P.; Beste, L.; Uddenberg, D.; Carlsbecker, A. PHABULOSA Mediates an Auxin Signaling Loop to Regulate Vascular Patterning in Arabidopsis. Plant Physiol. 2016, 170, 956–970. [Google Scholar] [CrossRef] [Green Version]
- Wenkel, S.; Emery, J.; Hou, B.-H.; Evans, M.M.S.; Barton, M.K. A Feedback Regulatory Module Formed by LITTLE ZIPPER and HD-ZIPIII Genes. Plant Cell 2007, 19, 3379–3390. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.-S.; Kim, S.-G.; Lee, M.; Lee, I.; Park, H.-Y.; Seo, P.J.; Jung, J.-H.; Kwon, E.-J.; Suh, S.W.; Paek, K.-H.; et al. HD-ZIP III Activity Is Modulated by Competitive Inhibitors via a Feedback Loop in Arabidopsis Shoot Apical Meristem Development. Plant Cell 2008, 20, 920–933. [Google Scholar] [CrossRef] [Green Version]
- Husbands, A.Y.; Aggarwal, V.; Ha, T.; Timmermans, M.C.P. In Planta Single-Molecule Pull-Down Reveals Tetrameric Stoichiometry of HD-ZIPIII:LITTLE ZIPPER Complexes. Plant Cell 2016, 28, 1783–1794. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Sun, S.; Ren, H.; Grison, M.; Boutté, Y.; Bai, W.; Men, S. The SMO1 Family of Sterol 4α-Methyl Oxidases Is Essential for Auxin- and Cytokinin-Regulated Embryogenesis. Plant Physiol. 2019, 181, 578–594. [Google Scholar] [CrossRef] [Green Version]
- Zhao, H.; Lü, S.; Li, R.; Chen, T.; Zhang, H.; Cui, P.; Ding, F.; Liu, P.; Wang, G.; Xia, Y.; et al. The Arabidopsis Gene DIG6 Encodes a Large 60S Subunit Nuclear Export GTPase 1 That Is Involved in Ribosome Biogenesis and Affects Multiple Auxin-Regulated Development Processes. J. Exp. Bot. 2015, 66, 6863–6875. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weng, J.-K.; Chapple, C. The Origin and Evolution of Lignin Biosynthesis. New Phytol. 2010, 187, 273–285. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Sztojka, B.; Escamez, S.; Vanholme, R.; Hedenström, M.; Wang, Y.; Turumtay, H.; Gorzsás, A.; Boerjan, W.; Tuominen, H. PIRIN2 Suppresses S-Type Lignin Accumulation in a Noncell-Autonomous Manner in Arabidopsis Xylem Elements. New Phytol. 2020, 225, 1923–1935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, B.; Tremousaygue, D.; Denancé, N.; van Esse, H.P.; Hörger, A.C.; Dabos, P.; Goffner, D.; Thomma, B.P.H.J.; van der Hoorn, R.A.L.; Tuominen, H. PIRIN2 Stabilizes Cysteine Protease XCP2 and Increases Susceptibility to the Vascular Pathogen Ralstonia Solanacearum in Arabidopsis. Plant J. 2014, 79, 1009–1019. [Google Scholar] [CrossRef] [Green Version]
- Petricka, J.J.; Clay, N.K.; Nelson, T.M. Vein Patterning Screens and the Defectively Organized Tributaries Mutants in Arabidopsis Thaliana. Plant J. 2008, 56, 251–263. [Google Scholar] [CrossRef]
- Song, C.-P.; Galbraith, D.W. AtSAP18, an Orthologue of Human SAP18, Is Involved in the Regulation of Salt Stress and Mediates Transcriptional Repression in Arabidopsis. Plant Mol. Biol. 2006, 60, 241–257. [Google Scholar] [CrossRef]
- de Oliveira Carvalho, A.; Gomes, V.M. Plant Defensins and Defensin-like Peptides—Biological Activities and Biotechnological Applications. Curr. Pharm. Des. 2011, 17, 4270–4293. [Google Scholar] [CrossRef]
- Xu, X.; Chen, C.; Fan, B.; Chen, Z. Physical and Functional Interactions between Pathogen-Induced Arabidopsis WRKY18, WRKY40, and WRKY60 Transcription Factors. Plant Cell 2006, 18, 1310–1326. [Google Scholar] [CrossRef] [Green Version]
- Chen, H.; Lai, Z.; Shi, J.; Xiao, Y.; Chen, Z.; Xu, X. Roles of Arabidopsis WRKY18, WRKY40 and WRKY60 Transcription Factors in Plant Responses to Abscisic Acid and Abiotic Stress. BMC Plant Biol. 2010, 10, 281. [Google Scholar] [CrossRef] [Green Version]
- Shang, Y.; Yan, L.; Liu, Z.-Q.; Cao, Z.; Mei, C.; Xin, Q.; Wu, F.-Q.; Wang, X.-F.; Du, S.-Y.; Jiang, T.; et al. The Mg-Chelatase H Subunit of Arabidopsis Antagonizes a Group of WRKY Transcription Repressors to Relieve ABA-Responsive Genes of Inhibition. Plant Cell 2010, 22, 1909–1935. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Sun, J.; Xu, Y.; Jiang, H.; Wu, X.; Li, C. The BHLH-Type Transcription Factor AtAIB Positively Regulates ABA Response in Arabidopsis. Plant Mol. Biol. 2007, 65, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Nakata, M.; Mitsuda, N.; Herde, M.; Koo, A.J.K.; Moreno, J.E.; Suzuki, K.; Howe, G.A.; Ohme-Takagi, M. A BHLH-Type Transcription Factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, Acts as a Repressor to Negatively Regulate Jasmonate Signaling in Arabidopsis. Plant Cell 2013, 25, 1641–1656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.; Zhang, L.; Yu, Y.; Quan, R.; Zhang, Z.; Zhang, H.; Huang, R. The Ethylene Response Factor AtERF11 That Is Transcriptionally Modulated by the BZIP Transcription Factor HY5 Is a Crucial Repressor for Ethylene Biosynthesis in Arabidopsis. Plant J. 2011, 68, 88–99. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Xing, J.; Zhang, K.; Pang, X.; Zhao, Y.; Wang, G.; Zang, J.; Huang, R.; Dong, J. Ethylene Response Factor ERF11 Activates BT4 Transcription to Regulate Immunity to Pseudomonas Syringae. Plant Physiol. 2019, 180, 1132–1151. [Google Scholar] [CrossRef] [Green Version]
- Walley, J.W.; Kelley, D.R.; Nestorova, G.; Hirschberg, D.L.; Dehesh, K. Arabidopsis Deadenylases AtCAF1a and AtCAF1b Play Overlapping and Distinct Roles in Mediating Environmental Stress Responses. Plant Physiol. 2010, 152, 866–875. [Google Scholar] [CrossRef] [Green Version]
- Silverstein, K.A.T.; Graham, M.A.; Paape, T.D.; VandenBosch, K.A. Genome Organization of More than 300 Defensin-like Genes in Arabidopsis. Plant Physiol. 2005, 138, 600–610. [Google Scholar] [CrossRef] [Green Version]
- Hawamda, A.I.M.; Reichert, S.; Ali, M.A.; Nawaz, M.A.; Austerlitz, T.; Schekahn, P.; Abbas, A.; Tenhaken, R.; Bohlmann, H. Characterization of an Arabidopsis Defensin-like Gene Conferring Resistance against Nematodes. Plants 2022, 11, 280. [Google Scholar] [CrossRef]
- Mondragón-Palomino, M.; Stam, R.; John-Arputharaj, A.; Dresselhaus, T. Diversification of Defensins and NLRs in Arabidopsis Species by Different Evolutionary Mechanisms. BMC Evol. Biol. 2017, 17, 255. [Google Scholar] [CrossRef] [Green Version]
- Parker, M.T.; Knop, K.; Zacharaki, V.; Sherwood, A.V.; Tomé, D.; Yu, X.; Martin, P.G.; Beynon, J.; Michaels, S.D.; Barton, G.J.; et al. Widespread Premature Transcription Termination of Arabidopsis Thaliana NLR Genes by the Spen Protein FPA. Elife 2021, 10, e65537. [Google Scholar] [CrossRef]
- Alomrani, S.; Kunert, K.J.; Foyer, C.H. Papain-like Cysteine Proteases Are Required for the Regulation of Photosynthetic Gene Expression and Acclimation to High Light Stress. J. Exp. Bot. 2021, 72, 3441–3454. [Google Scholar] [CrossRef]
- McLellan, H.; Gilroy, E.M.; Yun, B.-W.; Birch, P.R.J.; Loake, G.J. Functional Redundancy in the Arabidopsis Cathepsin B Gene Family Contributes to Basal Defence, the Hypersensitive Response and Senescence. New Phytol. 2009, 183, 408–418. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Sharma, A.; Niewiara, M.J.; Singh, R.; Ming, R.; Yu, Q. Papain-like Cysteine Proteases in Carica Papaya: Lineage-Specific Gene Duplication and Expansion. BMC Genom. 2018, 19, 26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Hu, M.; Wang, Q.; Cheng, L.; Zhang, Z. Role of Papain-Like Cysteine Proteases in Plant Development. Front. Plant Sci. 2018, 9, 1717. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nadimpalli, R.; Yalpani, N.; Johal, G.S.; Simmons, C.R. Prohibitins, Stomatins, and Plant Disease Response Genes Compose a Protein Superfamily That Controls Cell Proliferation, Ion Channel Regulation, and Death. J. Biol. Chem. 2000, 275, 29579–29586. [Google Scholar] [CrossRef] [Green Version]
- Jung, H.W.; Lim, C.W.; Lee, S.C.; Choi, H.W.; Hwang, C.H.; Hwang, B.K. Distinct Roles of the Pepper Hypersensitive Induced Reaction Protein Gene CaHIR1 in Disease and Osmotic Stress, as Determined by Comparative Transcriptome and Proteome Analyses. Planta 2008, 227, 409–425. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.; Katagiri, F. Purification of Low-Abundance Arabidopsis Plasma-Membrane Protein Complexes and Identification of Candidate Components. Plant J. 2009, 57, 932–944. [Google Scholar] [CrossRef]
- Zhou, L.; Cheung, M.-Y.; Li, M.-W.; Fu, Y.; Sun, Z.; Sun, S.-M.; Lam, H.-M. Rice Hypersensitive Induced Reaction Protein 1 (OsHIR1) Associates with Plasma Membrane and Triggers Hypersensitive Cell Death. BMC Plant Biol. 2010, 10, 290. [Google Scholar] [CrossRef] [Green Version]
- Choi, D.S.; Hwang, I.S.; Hwang, B.K. Requirement of the Cytosolic Interaction between PATHOGENESIS-RELATED PROTEIN10 and LEUCINE-RICH REPEAT PROTEIN1 for Cell Death and Defense Signaling in Pepper. Plant Cell 2012, 24, 1675–1690. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.-X.; Li, J.; Liu, Z.; Yin, J.; Chang, Z.-Y.; Rong, C.; Wu, J.-L.; Bi, F.-C.; Yao, N. The Arabidopsis Ceramidase AtACER Functions in Disease Resistance and Salt Tolerance. Plant J. 2015, 81, 767–780. [Google Scholar] [CrossRef]
- Nakano, Y.; Yamaguchi, M.; Endo, H.; Rejab, N.A.; Ohtani, M. NAC-MYB-Based Transcriptional Regulation of Secondary Cell Wall Biosynthesis in Land Plants. Front. Plant Sci. 2015, 6, 288. [Google Scholar] [CrossRef] [Green Version]
- Abdul Aziz, M.; Sabeem, M.; Mullath, S.K.; Brini, F.; Masmoudi, K. Plant Group II LEA Proteins: Intrinsically Disordered Structure for Multiple Functions in Response to Environmental Stresses. Biomolecules 2021, 11, 1662. [Google Scholar] [CrossRef] [PubMed]
- Kagaya, Y.; Okuda, R.; Ban, A.; Toyoshima, R.; Tsutsumida, K.; Usui, H.; Yamamoto, A.; Hattori, T. Indirect ABA-Dependent Regulation of Seed Storage Protein Genes by FUSCA3 Transcription Factor in Arabidopsis. Plant Cell Physiol. 2005, 46, 300–311. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Ruan, J.; Chu, P.; Fu, W.; Liang, Z.; Li, Y.; Tong, J.; Xiao, L.; Liu, J.; Li, C.; et al. AtPER1 Enhances Primary Seed Dormancy and Reduces Seed Germination by Suppressing the ABA Catabolism and GA Biosynthesis in Arabidopsis Seeds. Plant J. 2020, 101, 310–323. [Google Scholar] [CrossRef] [PubMed]
- Murashige, T.; Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Siebert, P.D.; Chenchik, A. Modified Acid Guanidinium Thiocyanate-Phenol-Chloroform RNA Extraction Method Which Greatly Reduces DNA Contamination. Nucleic Acids Res. 1993, 21, 2019–2020. [Google Scholar] [CrossRef] [Green Version]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- Ewels, P.; Magnusson, M.; Lundin, S.; Käller, M. MultiQC: Summarize Analysis Results for Multiple Tools and Samples in a Single Report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef] [Green Version]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New Capabilities and Interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Dodueva, I.E.; Kiryushkin, A.S.; Yurlova, E.V.; Osipova, M.A.; Buzovkina, I.S.; Lutova, L.A. Effect of Cytokinins on Expression of Radish CLE Genes. Russ. J. Plant Physiol. 2013, 60, 388–395. [Google Scholar] [CrossRef]




Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kuznetsova, K.; Dodueva, I.; Gancheva, M.; Lutova, L. Transcriptomic Analysis of Radish (Raphanus sativus L.) Roots with CLE41 Overexpression. Plants 2022, 11, 2163. https://doi.org/10.3390/plants11162163
Kuznetsova K, Dodueva I, Gancheva M, Lutova L. Transcriptomic Analysis of Radish (Raphanus sativus L.) Roots with CLE41 Overexpression. Plants. 2022; 11(16):2163. https://doi.org/10.3390/plants11162163
Chicago/Turabian StyleKuznetsova, Ksenia, Irina Dodueva, Maria Gancheva, and Lyudmila Lutova. 2022. "Transcriptomic Analysis of Radish (Raphanus sativus L.) Roots with CLE41 Overexpression" Plants 11, no. 16: 2163. https://doi.org/10.3390/plants11162163
APA StyleKuznetsova, K., Dodueva, I., Gancheva, M., & Lutova, L. (2022). Transcriptomic Analysis of Radish (Raphanus sativus L.) Roots with CLE41 Overexpression. Plants, 11(16), 2163. https://doi.org/10.3390/plants11162163




