Genome-Wide Association Study of Arabinoxylan Content from a 562 Hexaploid Wheat Collection
Abstract
:1. Introduction
2. Results
2.1. Population Structure Analysis
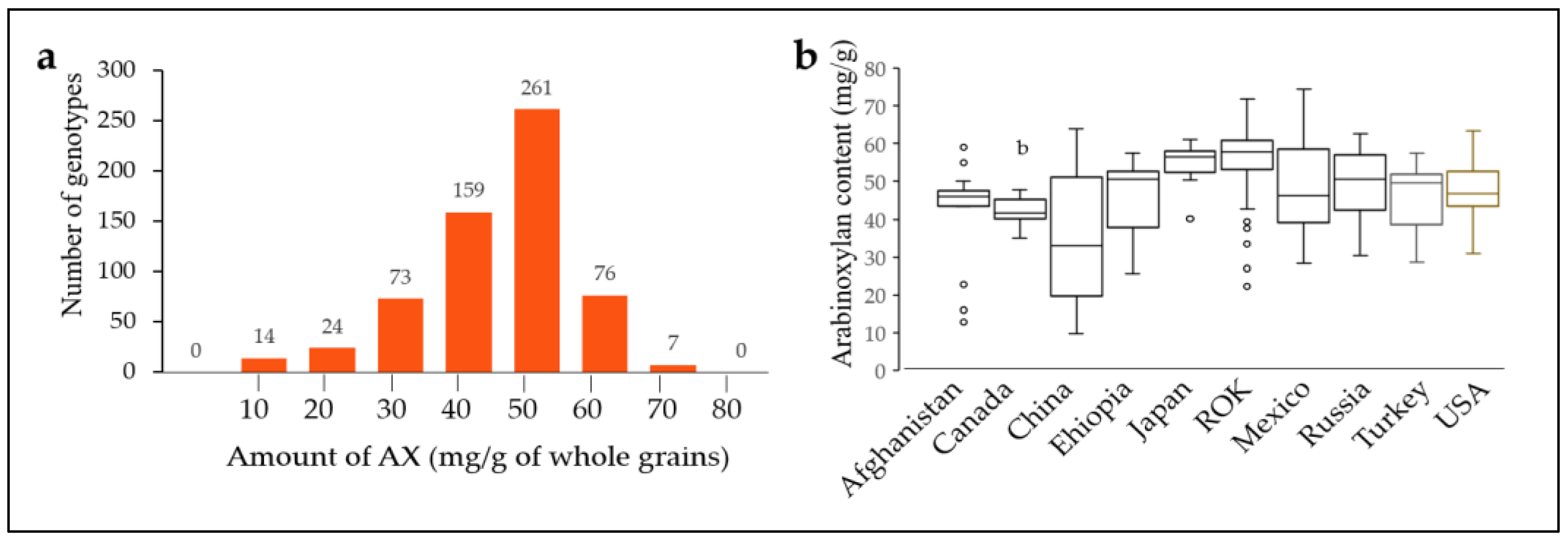
2.2. Arabinoxylan Content of 562 Bread Wheat Varieties
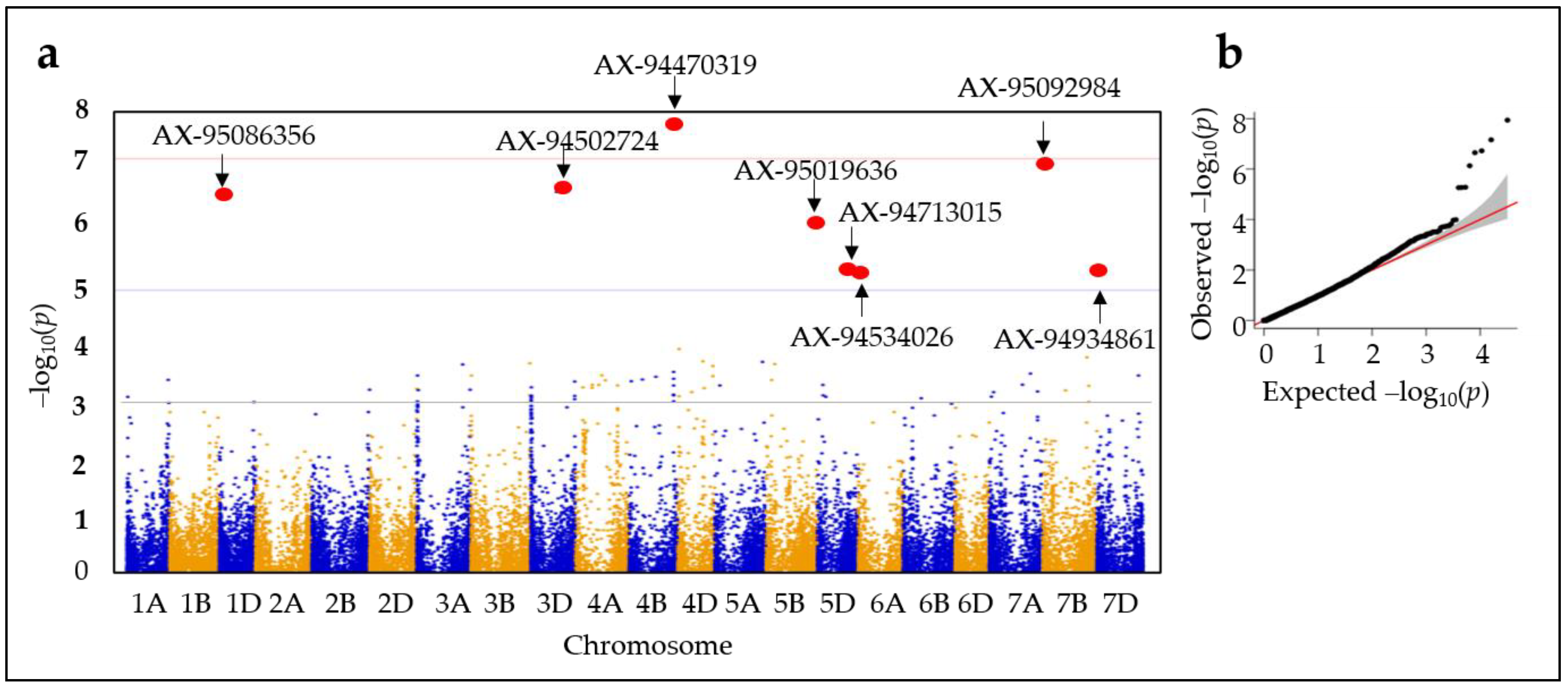
2.3. Genome-Wide Association Studies (GWASs)
2.4. Candidate Gene Identification
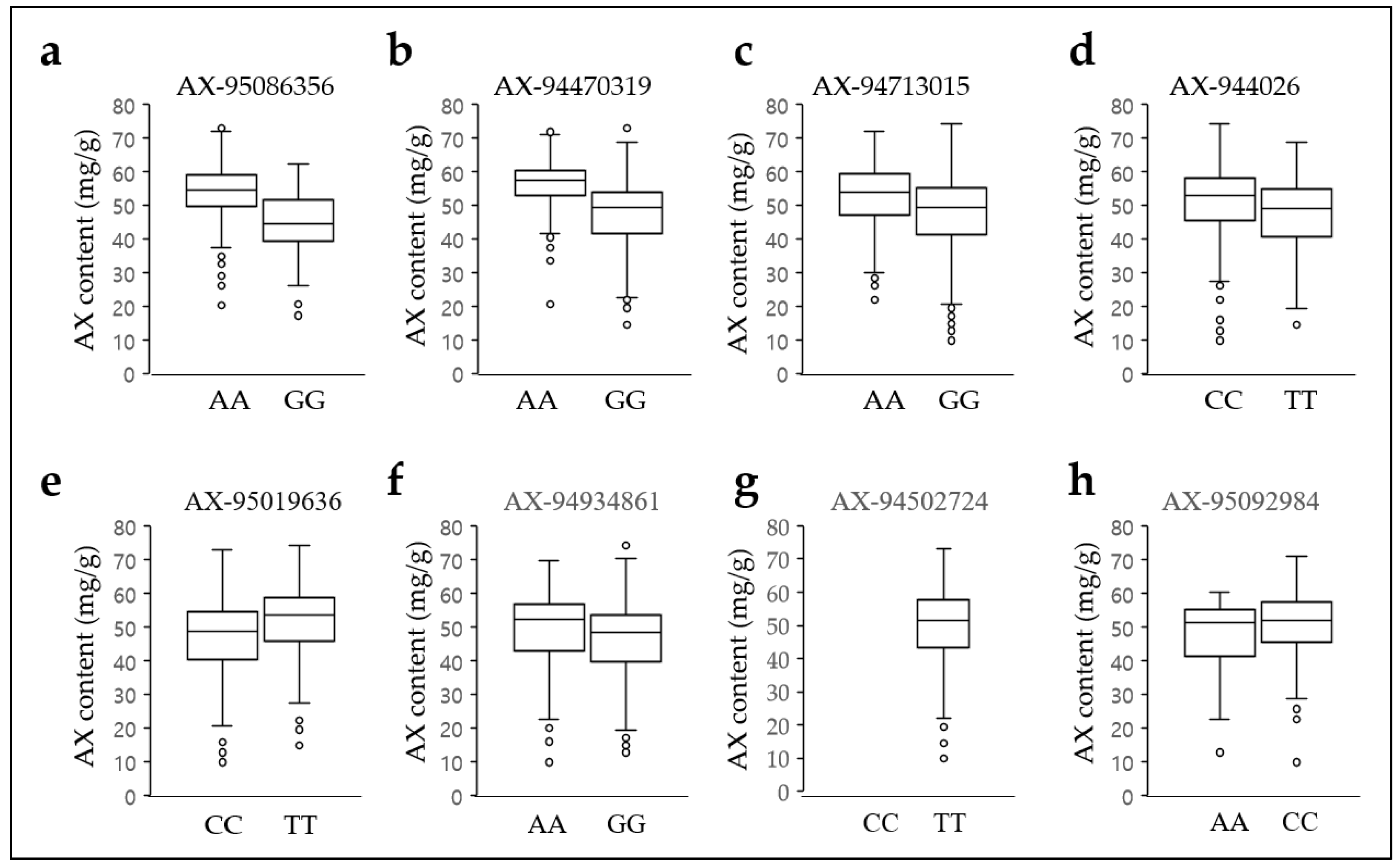
2.5. Arabinoxylan Distribution of the Genotypes for the Three SNPs
2.6. Arabinoxylan Content by Country According to the Three SNP Genotypes
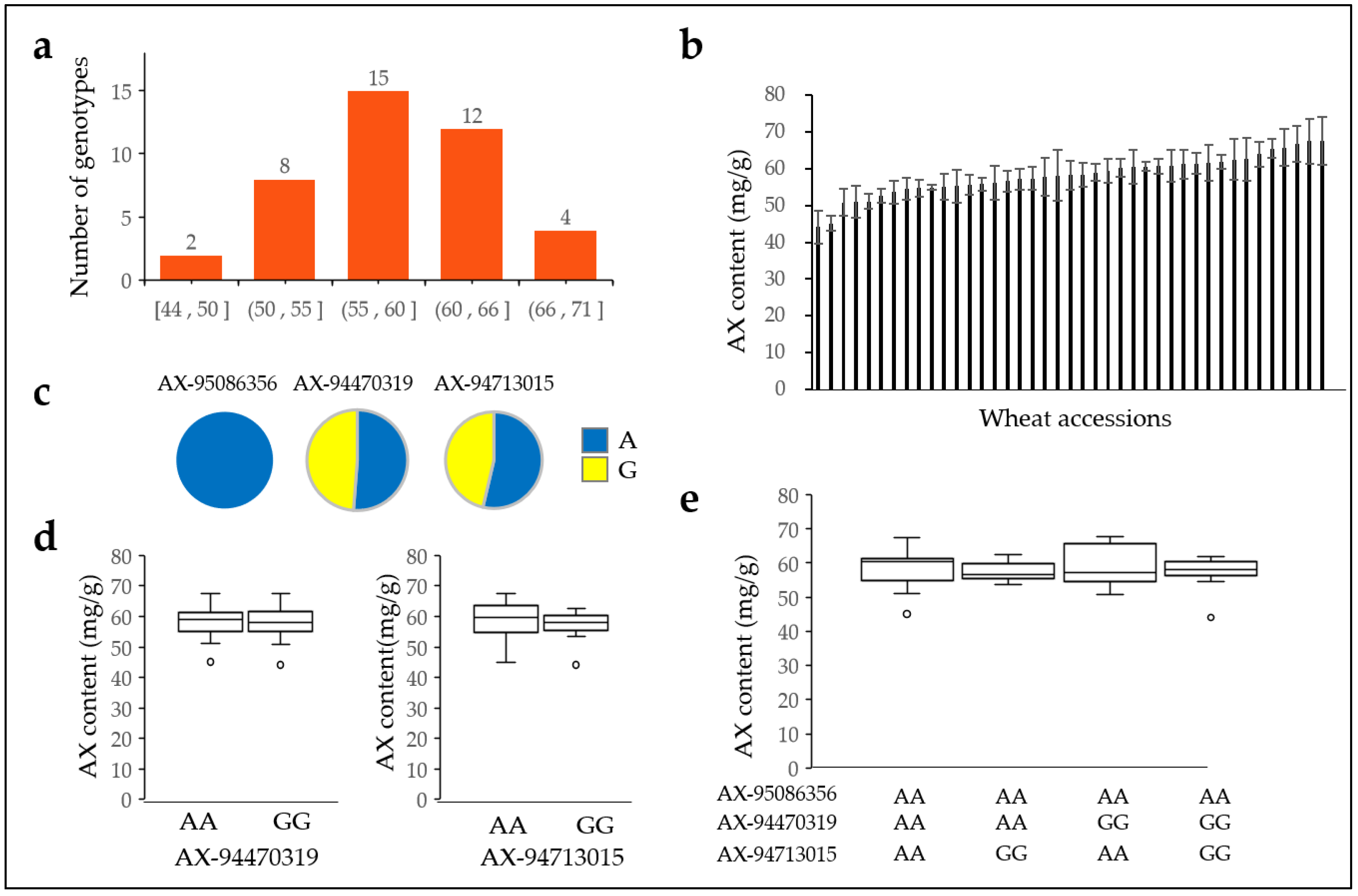
2.7. Arabinoxylan Content in Three Allelic Combinations
2.8. Arabinoxylan Content of 41 Domestic Wheat Varieties
3. Discussion
4. Materials and Methods
4.1. Materials and Arabinoxylan Extraction
4.2. DNA Extraction and Genotyping
4.3. Genome-Wide Association Analysis and Population Structure
4.4. Identification of Putative Candidate Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Mendis, M.; Simsek, S. Arabinoxylans and human health. Food Hydrocoll. 2014, 42, 239–243. [Google Scholar] [CrossRef]
- Yadav, M.P.; Kale, M.S.; Hicks, K.B.; Hanah, K. Isolation, characterization and the functional properties of cellulosic arabinoxylan fiber isolated from agricultural processing by-products, agricultural residues and energy crops. Food Hydrocoll. 2017, 63, 545–551. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Li, S.; Fu, Y.; Li, C.; Chen, D.; Chen, H. Arabinoxylan structural characteristics, interaction with gut microbiota and potential health functions. J. Funct. Foods 2019, 54, 536–551. [Google Scholar] [CrossRef]
- Casas, G.A.; Lærke, H.N.; Knudsen, K.E.B.; Stein, H.H. Arabinoxylan is the main polysaccharide in fiber from rice coproducts, and increased concentration of fiber decreases in vitro digestibility of dry matter. Anim. Feed. Sci. Technol. 2018, 247, 255–261. [Google Scholar] [CrossRef]
- He, H.J.; Qiao, J.; Liu, Y.; Guo, Q.; Ou, X.; Wang, X. Isolation, Structural, Functional, and Bioactive Properties of Cereal Arabinoxylan—A Critical Review. J. Agric. Food Chem. 2021, 69, 15437–15457. [Google Scholar] [CrossRef]
- Correia, M.; Mazumder, K.; Brás, J.; Firbank, S.J.; Zhu, Y.; Lewis, R.; York, W.S.; Fontes, C.M.; Gilbert, H.J. Structure and function of an arabinoxylan-specific xylanase. J. Biol. Chem. 2011, 286, 22510–22520. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gopalan, N.; Rodríguez-Duran, L.; Saucedo-Castaneda, G.; Nampoothiri, K.M. Review on technological and scientific aspects of feruloyl esterases: A versatile enzyme for biorefining of biomass. Bioresour. Technol. 2015, 193, 534–544. [Google Scholar] [CrossRef]
- El-Seedi, H.R.; El-Said, A.M.A.; Khalifa, S.A.M.; Göransson, U.; Bohlin, L.; Borg-Karlson, A.-K.; Verpoorte, R. Biosynthesis, natural sources, dietary intake, pharmacokinetic properties, and biological activities of hydroxycinnamic acids. J. Agric. Food Chem. 2012, 60, 10877–10895. [Google Scholar] [CrossRef]
- De Santis, M.A.; Kosik, O.; Passmore, D.; Flagella, Z.; Shewry, P.R.; Lovegrove, A. Comparison of the dietary fibre composition of old and modern durum wheat (Triticum turgidum spp. durum) genotypes. Food Chem. 2017, 244, 304–310. [Google Scholar] [CrossRef]
- Dodevska, M.S.; Djordjevic, B.I.; Sobajic, S.S.; Miletic, I.D.; Djordjevic, P.B.; Dimitrijevic-Sreckovic, V.S. Characterisation of dietary fibre components in cereals and legumes used in Serbian diet. Food Chem. 2013, 141, 1624–1629. [Google Scholar] [CrossRef]
- Rainakari, A.-I.; Rita, H.; Putkonen, T.; Pastell, H. New dietary fibre content results for cereals in the Nordic countries using AOAC 2011.25 method. J. Food Compos. Anal. 2016, 51, 1–8. [Google Scholar] [CrossRef]
- Onipe, O.O.; Jideani, A.I.O.; Beswa, D. Composition and functionality of wheat bran and its application in some cereal food products. Int. J. Food Sci. Technol. 2015, 50, 2509–2518. [Google Scholar] [CrossRef]
- Marcotuli, I.; Hsieh, Y.S.-Y.; Lahnstein, J.; Yap, K.; Burton, R.A.; Blanco, A.; Fincher, G.B.; Gadaleta, A. Structural Variation and Content of Arabinoxylans in Endosperm and Bran of Durum Wheat (Triticum turgidum L.). J. Agric. Food Chem. 2016, 64, 2883–2892. [Google Scholar] [CrossRef] [PubMed]
- Michelini, E.; Guardigli, M.; Magliulo, M.; Mirasoli, M.; Roda, A.; Simoni, P.; Baraldini, M. Bioluminescent biosensors based on genetically engineered living cells in environmental and food analysis. Anal. Lett. 2006, 39, 1503–1515. [Google Scholar] [CrossRef]
- Saulnier, L.; Sado, P.-E.; Branlard, G.; Charmet, G.; Guillon, F. Wheat arabinoxylans: Exploiting variation in amount and composition to develop enhanced varieties. J. Cereal Sci. 2007, 46, 261–281. [Google Scholar] [CrossRef]
- Barron, C.; Bar-L’Helgouac’h, C.; Champ, M.; Saulnier, L. Arabinoxylan content and grain tissue distribution are good predictors of the dietary fibre content and their nutritional properties in wheat products. Food Chem. 2020, 328, 127111. [Google Scholar] [CrossRef]
- Ebringerová, A.; Hromádková, Z.; Heinze, T.; Hemicellulose, T.H. Polysaccharides I. Adv. Polym. Sci. 2005, 186, 67. [Google Scholar]
- Courtin, C.M.; Delcour, J.A. Arabinoxylans and endoxylanases in wheat flour bread-making. J. Cereal Sci. 2002, 35, 225–243. [Google Scholar] [CrossRef]
- Finnie, S.M.; Bettge, A.D.; Morris, C.F. Influence of cultivar and environment on water-soluble and water-insoluble arabinoxylans in soft wheat. Cereal Chem. 2006, 83, 617–623. [Google Scholar] [CrossRef]
- Tremmel-Bede, K.; Láng, L.; Török, K.; Tömösközi, S.; Vida, G.; Shewry, P.R.; Bedő, Z.; Rakszegi, M. Development and characterization of wheat lines with increased levels of arabinoxylan. Euphytica 2017, 213, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Wang, M.; Hamer, R.; van Vliet, T.; Gruppen, H.; Marseille, H.; Weegels, P. Effect of water unextractable solids on gluten formation and properties: Mechanistic considerations. J. Cereal Sci. 2003, 37, 55–64. [Google Scholar] [CrossRef]
- Li, S.; Morris, C.F.; Bettge, A.D. Genotype and environment variation for arabinoxylans in hard winter and spring wheats of the u.s. pacific northwest. Cereal Chem. 2009, 86, 88–95. [Google Scholar] [CrossRef] [Green Version]
- Dornez, E.; Gebruers, K.; Joye, I.J.; De Ketelaere, B.; Lenartz, J.; Massaux, C.; Bodson, B.; Delcour, J.A.; Courtin, C.M. Effects of fungicide treatment, N-fertilisation and harvest date on arabinoxylan, endoxylanase activity and endoxylanase inhibitor levels in wheat kernels. J. Cereal Sci. 2008, 47, 190–200. [Google Scholar] [CrossRef]
- Hong, B.H.; Rubenthaler, G.L.; Allan, R.E. Wheat pentosans. I. Cultivar variation and relationship to kernel hardness. Cereal Chem. 1989, 66, 369–373. [Google Scholar]
- Saulnier, L.; Peneau, N.; Thibault, J.-F. Variability in grain extract viscosity and water-soluble arabinoxylan content in wheat. J. Cereal Sci. 1995, 22, 259–264. [Google Scholar] [CrossRef]
- Hernández-Espinosa, N.; Romano, G.P.; Crespo-Herrera, L.; Singh, R.; Guzmán, C.; Ibba, M.I. Endogenous arabinoxylans variability in refined wheat flour and its relationship with quality traits. J. Cereal Sci. 2020, 95, 103062. [Google Scholar] [CrossRef]
- Vignola, M.B.; Moiraghi, M.; Salvucci, E.; Baronia, V.T.; P’erez, G. Whole meal and white flour from argentine wheat genotypes: Mineral and arabinoxylan differences. J. Cereal Sci. 2016, 71, 217–223. [Google Scholar] [CrossRef]
- Lempereur, I.; Rouau, X.; Abecassis, J. Genetic and agronomic variation in arabinoxylan and ferulic acid contents of durum wheat (Triticum durum L.) grain and its milling fractions. J. Cereal Sci. 1997, 25, 103–110. [Google Scholar] [CrossRef]
- Marcotuli, I.; Houston, K.; Waugh, R.; Fincher, G.B.; Burton, R.A.; Blanco, A.; Gadaleta, A. Genome wide association mapping for arabinoxylan content in a collection of tetraploid wheats. PLoS ONE 2015, 10, e0132787. [Google Scholar] [CrossRef] [Green Version]
- Hassan, A.S.; Houston, K.; Lahnstein, J.; Shirley, N.; Schwerdt, J.G.; Gidley, M.J.; Waugh, R.; Little, A.; Burton, R.A. A Genome Wide Association Study of arabinoxylan content in 2-row spring barley grain. PLoS ONE 2017, 12, e0182537. [Google Scholar] [CrossRef] [Green Version]
- Ibba, M.I.; Juliana, P.; Hernández-Espinosa, N.; Posadas-Romano, G.; Dreisigacker, S.; Sehgal, D.; Crespo-Herrera, L.; Singh, R.; Guzmán, C. Genome-wide association analysis for arabinoxylan content in common wheat (T. Aestivum L.) flour. J. Cereal Sci. 2021, 98, 103166. [Google Scholar] [CrossRef]
- Wang, J.; Zhang, Z. GAPIT Version 3: Boosting power and accuracy for genomic association and prediction. Genom. Proteom. Bioinform. 2021, 19, 629–640. [Google Scholar] [CrossRef]
- Turner, S.D. qqman: An R package for visualizing GWAS results using Q-Q and manhattan plots. J. Open Sour. Softw. 2018, 3, 731. [Google Scholar] [CrossRef] [Green Version]
- Wang, Y.; Zhang, W.-Z.; Song, L.-F.; Zou, J.-J.; Su, Z.; Wu, W.-H. Transcriptome analyses show changes in gene expression to accompany pollen germination and tube growth in arabidopsis. Plant Physiol. 2008, 148, 1201–1211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grant, M.R.; Godiard, L.; Straube, E.; Ashfield, T.; Lewald, J.; Sattler, A.; Innes, R.W.; Dangl, J.L. Structure of the Arabidopsis RPM1 gene enabling dual specificity disease resistance. Science 1995, 269, 843–846. [Google Scholar] [CrossRef] [PubMed]
- Mackey, D.; Holt, B.F., III; Wiig, A.; Dangl, J.L. RIN4 interacts with Pseudomonas syringae Type III effector molecules and is required for RPM1-mediated resistance in arabidopsis. Cell 2002, 108, 743–754. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Tian, W.; Tao, F.; Wang, J.; Shang, H.; Chen, X.; Xu, X.; Hu, X. TaRPM1 positively regulates wheat high-temperature seedling-plant resistance to Puccinia striiformis f. sp. tritici. Front. Plant Sci. 2020, 10, 1679. [Google Scholar] [CrossRef] [Green Version]
- Du, D.; Zhang, C.; Xing, Y.; Lu, X.; Cai, L.; Yun, H.; Zhang, Q.; Zhang, Y.; Chen, X.; Liu, M.; et al. The CC-NB-LRR OsRLR1 mediates rice disease resistance through interaction with OsWRKY19. Plant Biotechnol. J. 2021, 19, 1052–1064. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Yan, X.; Li, J.; Yang, Q.; Jamil, W.; Teng, Y.; Bai, S. Genome wide identification and predicted functional analyses of NAC transcription factors in Asian pears. BMC Plant Biol. 2018, 18, 1–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiang, Y.; Tang, N.; Du, H.; Ye, H.; Xiong, L. Characterization of OsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol. 2008, 148, 1938–1952. [Google Scholar] [CrossRef] [Green Version]
- Liang, Y.; Xia, J.; Jiang, Y.; Bao, Y.; Chen, H.; Wang, D.; Zhang, D.; Yu, J.; Cang, J. Genome-Wide Identification and Analysis of bZIP Gene Family and Resistance of TaABI5 (TabZIP96) under Freezing Stress in Wheat (Triticum aestivum). Int. J. Mol. Sci. 2022, 23, 2351. [Google Scholar] [CrossRef] [PubMed]
- Xiaoli, G. Isolation and Function Analysis of Heat Stress Related Transcription Factor Gene TabZIP28 in Wheat (Triticum aestivum). J. Agric. Biotechnol. 2016, 24, 157–167. [Google Scholar]
- Wang, F.; Lin, R.; Li, Y.; Wang, P.; Feng, J.; Chen, W.; Xu, S. TabZIP74 Acts as a Positive Regulator in Wheat Stripe Rust Resistance and Involves Root Development by mRNA Splicing. Front. Plant Sci. 2019, 10, 1551. [Google Scholar] [CrossRef] [PubMed]
- Guo, G.; Yang, Y.; Cao, L.; Liu, W.; Caili, B.I. RF2 Basic Leucine Zipper Transcription Factor TabZIP3 Involved in Salt Stress Response in Wheat. J. Agric. Sci. Technol. 2019, 21, 20. [Google Scholar]
- Bi, C.; Yu, Y.; Dong, C.; Yang, Y.; Zhai, Y.; Du, F.; Xia, C.; Ni, Z.; Kong, X.; Zhang, L. The bZIP transcription factor TabZIP15 improves salt stress tolerance in wheat. Plant Biotechnol. J. 2020, 19, 209–211. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Li, L.; Peng, D.; Liu, M.; Wei, A.; Li, X. TaFDL2-1A interacts with TabZIP8-7A protein to cope with drought stress via the abscisic acid signaling pathway. Plant Sci. 2021, 311, 111022. [Google Scholar] [CrossRef]
- Liu, M.; Wang, Z.; Xiao, H.-M.; Yang, Y. Characterization of TaDREB1 in wheat genotypes with different seed germination under osmotic stress. Hereditas 2018, 155, 26. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, L.; Xia, C.; Gao, L.; Hao, C.; Zhao, G.; Jia, J.; Kong, X. A Novel Wheat C-bZIP Gene, TabZIP14-B, Participates in Salt and Freezing Tolerance in Transgenic Plants. Front. Plant Sci. 2017, 8, 710. [Google Scholar] [CrossRef] [Green Version]
- Miedes, E.; Vanholme, R.; Boerjan, W.; Molina, A. The role of the secondary cell wall in plant resistance to pathogens. Front. Plant Sci. 2014, 5, 358. [Google Scholar] [CrossRef] [Green Version]
- Molina, A.; Miedes, E.; Bacete, L.; Rodríguez, T.; Mélida, H.; Denancé, N.; Sanchez-Vallet, A.; Rivière, M.; López, G.; Freydier, a.; et al. Arabidopsis cell wall composition determines disease resistance specificity and fitness. Proc. Natl. Acad. Sci. USA 2021, 118, e2010243118. [Google Scholar] [CrossRef]
- Min, K.D.; Na Kang, Y.; Kim, C.S.; Choi, C.H.; Kim, A.J.Y. Genetic diversity and population structure of Korean common wheat (Triticum aestivum). Korean J. Breed. Sci. 2021, 53, 277–288. [Google Scholar] [CrossRef]
- Yang, J.; Park, J.; Son, J.-H.; Kim, K.-H.; Kim, K.-M.; Jeong, H.-Y.; Kang, C.-S.; Son, J.-Y.; Park, T.-I.; Choi, C. Protein and Arabinoxylan Contents of Whole Grains from Wheat Genetic Resources Cultivated in Korea. Korean J. Crop Sci. 2021, 66, 29–36. [Google Scholar]
- Allen, A.M.; Winfield, M.O.; Burridge, A.J.; Downie, R.C.; Benbow, H.R.; Barker, G.L.A.; Wilkinson, P.A.; Coghill, J.; Waterfall, C.; Davassi, A.; et al. Characterization of a Wheat Breeders’ Array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol. J. 2017, 15, 390–401. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- CerealsDB. Available online: https://www.faceDb.uk.net (accessed on 20 October 2022).
- Huang, M.; Liu, X.; Zhou, Y.; Summers, R.M.; Zhang, Z. BLINK: A package for the next level of genome-wide association studies with both individuals and markers in the millions. Gigascience 2018, 8, giy154. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- QTLmax 3.0. Available online: https://www.qtlmax.com (accessed on 12 December 2022).
- Zhu, T.; Wang, L.; Rimbert, H.; Rodriguez, J.C.; Deal, K.R.; De Oliveira, R.; Choulet, F.; Keeble-Gagnère, G.; Tibbits, J.; Rogers, J.; et al. Optical maps refine the bread wheat Triticum aestivum cv Chinese Spring genome assembly. Plant J. 2021, 107, 303–314. [Google Scholar] [CrossRef]




| SNP | Chr. | Position | p-Value | RefSeq Sequence | FDR | Gene Name | Alleles |
|---|---|---|---|---|---|---|---|
| AX–95086356 | 1BL | 677224871 | 2.24 × 10−7 | NC_057795.1 | 0.000791 | F-box/FBD/LRR-repeat protein | A/G |
| AX–94502724 | 3DL | 354080813 | 1.90 × 10−8 | NC_057802.1 | 0.001791 | NA | C/T |
| AX–94470319 | 4BL | 609317748 | 1.15 × 10−8 | NC_057804.1 | 0.000369 | Disease resistance protein RPM1/PIK6-NP-like | A/G |
| AX–95019636 | 5BL | 696490490 | 7.41 × 10−7 | NC_057807.1 | 0.00473 | U2 small nuclear ribonucleoprotein A′-like | C/T |
| AX–94534026 | 5DL | 548367051 | 5.47 × 10−6 | NC_057808.1 | 0.021813 | serine/arginine repetitive matrix protein 1-like | C/T |
| AX–94713015 | 5DL | 409878931 | 5.27 × 10−6 | NC_057808.1 | 0.021813 | bZIP transcription factior 29 | A/G |
| AX–95092984 | 7BS | 67265891 | 6.94 × 10−8 | NC_057813.1 | 0.001109 | Carboxypeptidase | A/C |
| AX–94934861 | 7DS | 2522044 | 5.39 × 10−6 | NC_057814.1 | 0.021813 | Disease resistance protein RPP13/PIK6-NP-like | A/G |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, M.H.; Park, J.; Kim, K.-H.; Kim, K.-M.; Kang, C.-S.; Lee, G.E.; Choi, J.Y.; Shon, J.; Ko, J.-M.; Choi, C. Genome-Wide Association Study of Arabinoxylan Content from a 562 Hexaploid Wheat Collection. Plants 2023, 12, 184. https://doi.org/10.3390/plants12010184
Lee MH, Park J, Kim K-H, Kim K-M, Kang C-S, Lee GE, Choi JY, Shon J, Ko J-M, Choi C. Genome-Wide Association Study of Arabinoxylan Content from a 562 Hexaploid Wheat Collection. Plants. 2023; 12(1):184. https://doi.org/10.3390/plants12010184
Chicago/Turabian StyleLee, Myoung Hui, Jinhee Park, Kyeong-Hoon Kim, Kyeong-Min Kim, Chon-Sik Kang, Go Eun Lee, Jun Yong Choi, Jiyoung Shon, Jong-Min Ko, and Changhyun Choi. 2023. "Genome-Wide Association Study of Arabinoxylan Content from a 562 Hexaploid Wheat Collection" Plants 12, no. 1: 184. https://doi.org/10.3390/plants12010184







