Genome-Wide Identification and Analysis of Catharanthus roseus Receptor-like Kinase 1-like Proteins in Eggplant
Abstract
:1. Introduction
2. Results
2.1. Identification of CrRLK1L Proteins in Eggplant
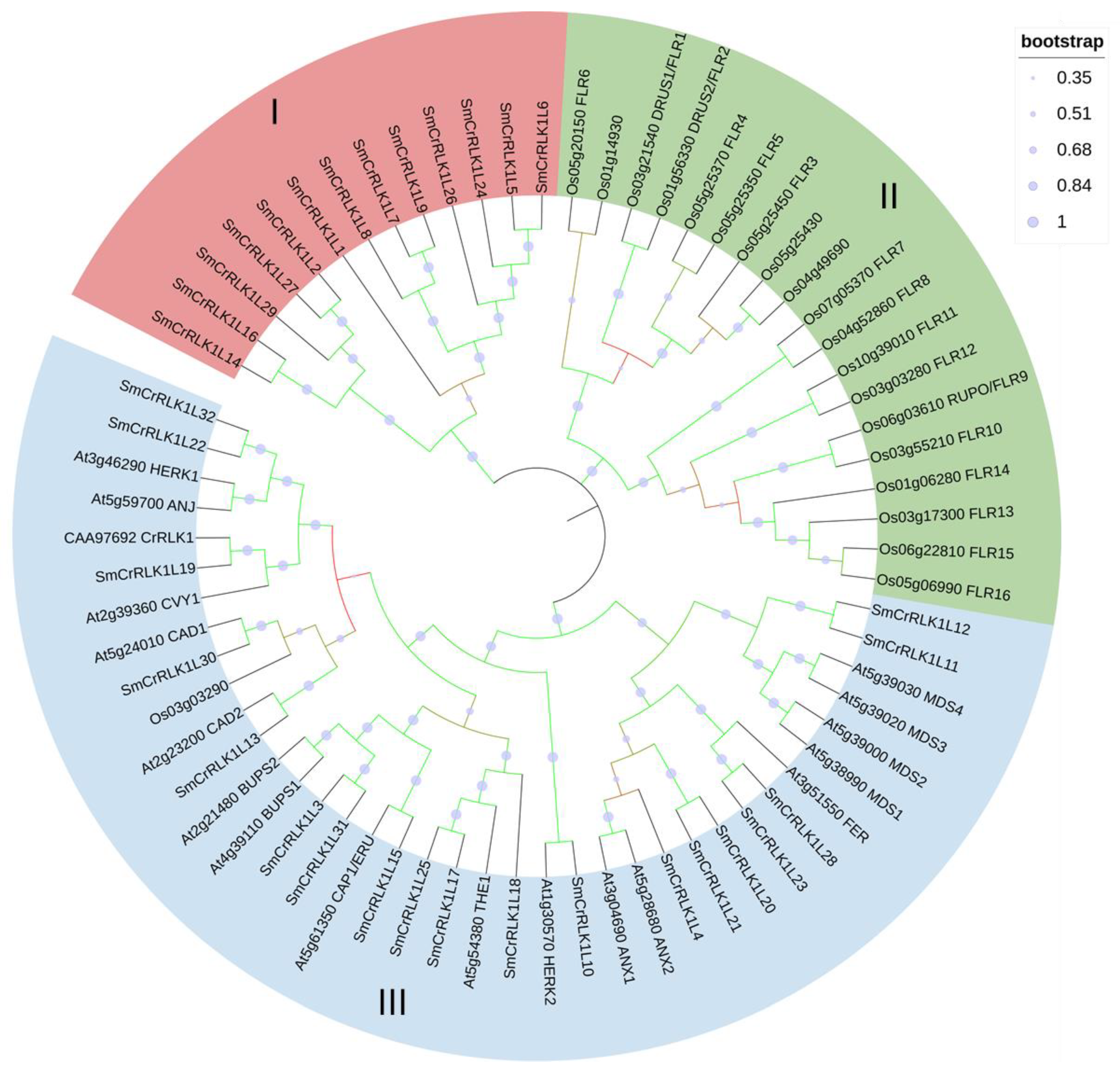
2.2. Phylogenetic Analysis of the SmCrRLK1Ls
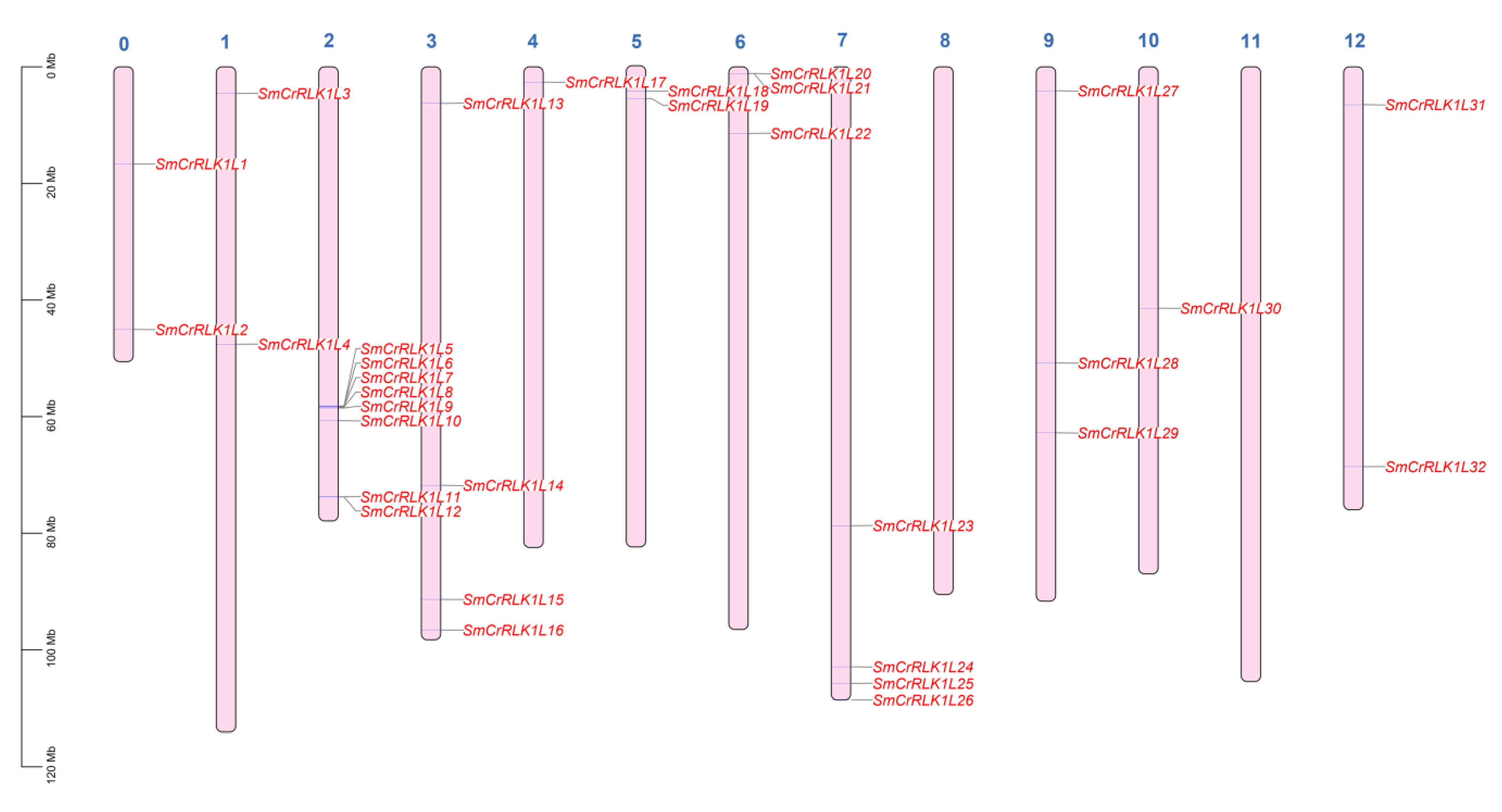
2.3. Chromosome Distribution of SmCrRLK1Ls
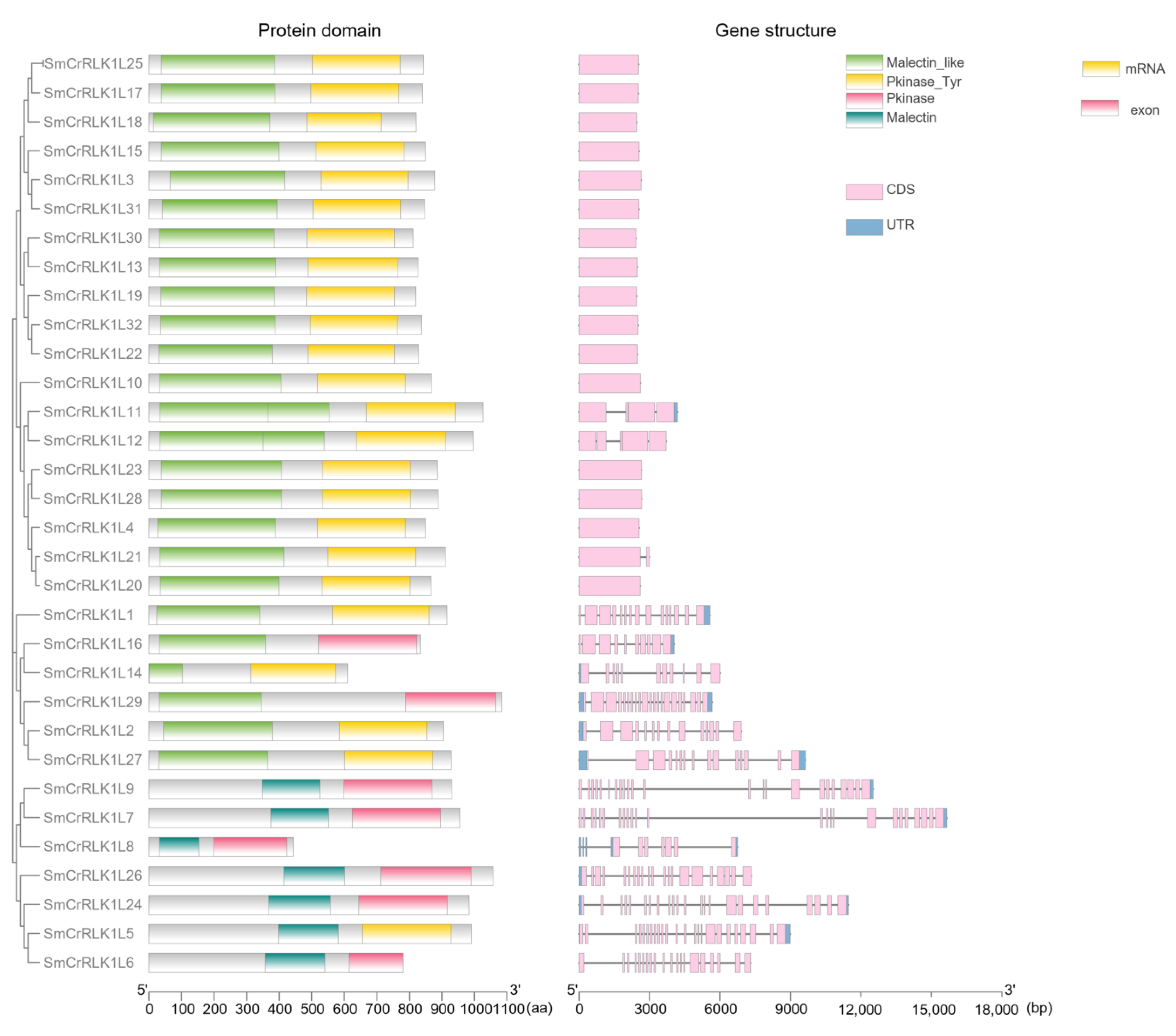
2.4. Protein Domain and Gene Structure Analysis of SmCrRLK1L
2.5. Conserved Motifs and Subcellular Localization of SmCrRLK1L Prediction
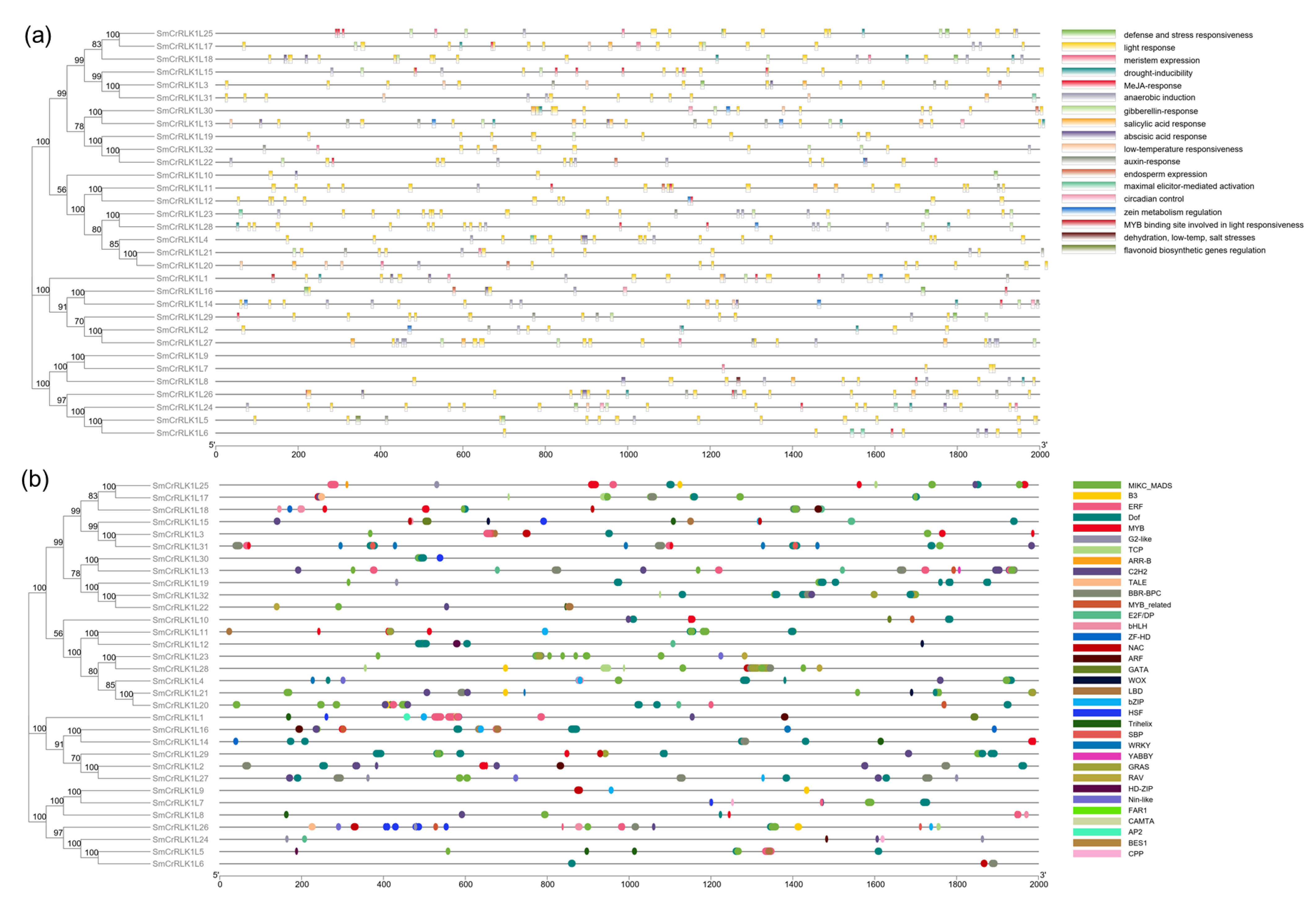
2.6. SmCrRLK1L Gene Promoter Analysis
2.7. SmCrRLK1L Gene Duplication Events
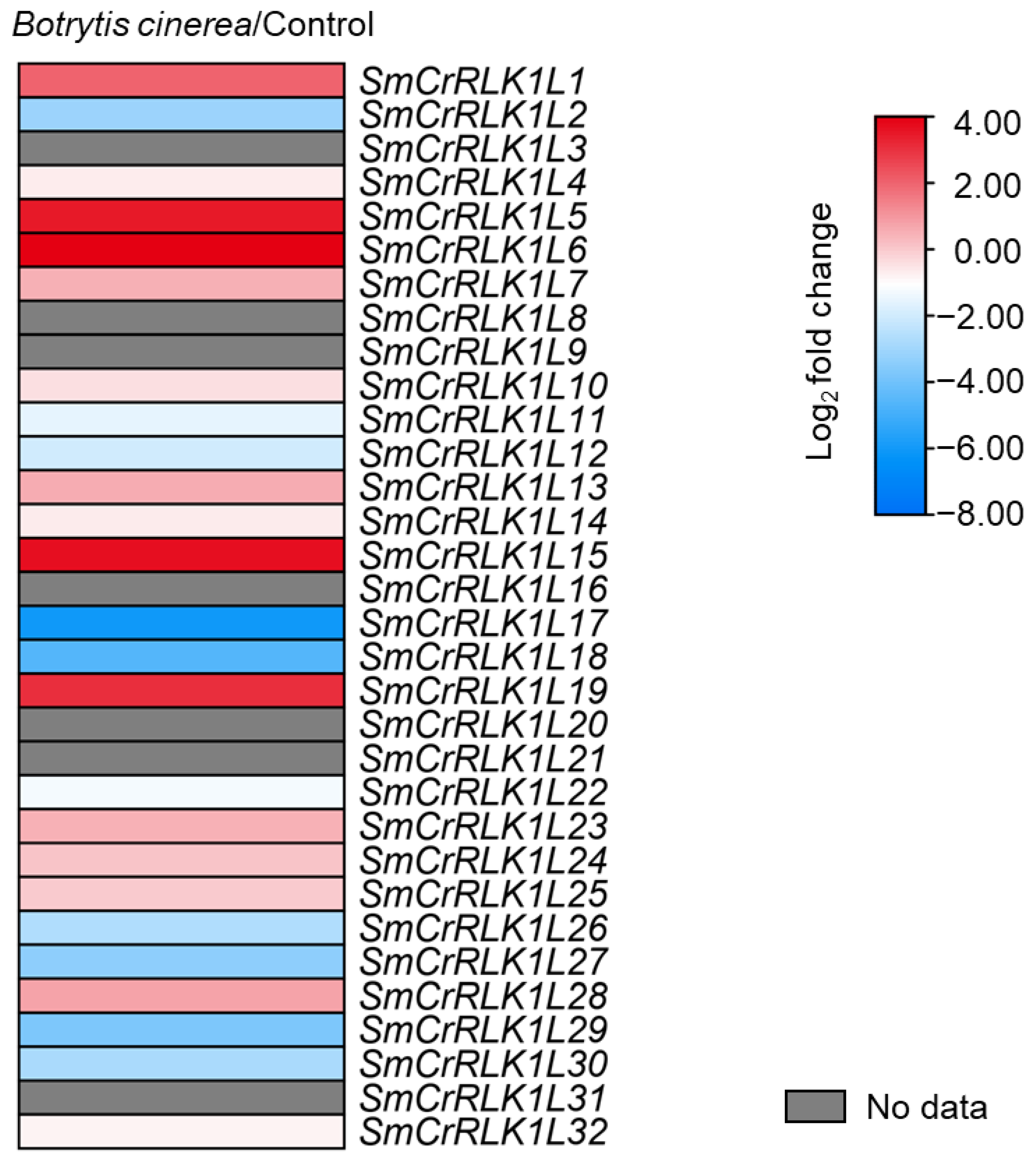
2.8. Expression Analysis of SmCrRLK1L Genes in Response to Botrytis cinerea Infection
3. Discussion
4. Materials and Methods
4.1. SmCrRLK1L Members, Physicochemical Property Identification
4.2. Phylogenetic Analysis
4.3. Gene Location and Collinearity Analysis
4.4. Protein Domain and Gene Structure Analysis
4.5. Subcellular Localization Prediction
4.6. Protein Conserved Motifs Analysis
4.7. Promoter Analysis
4.8. Transcriptomic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Ji, D.; Chen, T.; Zhang, Z.; Li, B.; Tian, S. Versatile Roles of the Receptor-like Kinase Feronia in Plant Growth, Development and Host-Pathogen Interaction. Int. J. Mol. Sci. 2020, 21, 7881. [Google Scholar] [CrossRef]
- Zhu, S.; Fu, Q.; Xu, F.; Zheng, H.; Yu, F. New Paradigms in Cell Adaptation: Decades of Discoveries on the CrRLK1L Receptor Kinase Signalling Network. New Phytol. 2021, 232, 1168–1183. [Google Scholar] [CrossRef]
- Schulze-Muth, P.; Irmler, S.; Schröder, G.; Schröder, J. Novel Type of Receptor-like Protein Kinase from a Higher Plant (Catharanthus roseus). J. Biol. Chem. 1996, 271, 26684–26689. [Google Scholar] [CrossRef]
- Franck, C.M.; Westermann, J.; Boisson-Dernier, A. Plant Malectin-like Receptor Kinases: From Cell Wall Integrity to Immunity and Beyond. Annu. Rev. Plant. Biol. 2018, 69, 301–328. [Google Scholar] [CrossRef]
- Schallus, T.; Jaeckh, C.; Fehér, K.; Palma, A.S.; Liu, Y.; Simpson, J.C.; Mackeen, M.; Stier, G.; Gibson, T.J.; Feizi, T.; et al. Malectin: A Novel Carbohydrate-Binding Protein of the Endoplasmic Reticulum and a Candidate Player in the Early Steps of Protein N-Glycosylation. Mol. Biol. Cell 2008, 19, 3404–3414. [Google Scholar] [CrossRef]
- Escobar-Restrepo, J.; Huck, N.; Kessler, S.; Gagliardini, V.; Gheyselinck, J.; Yang, W.; Grossniklaus, U. The FERONIA Receptor-like Kinase Mediates Male-Female Interactions during Pollen Tube Reception. Science 2007, 317, 656–660. [Google Scholar] [CrossRef]
- Duan, Q.; Kita, D.; Li, C.; Cheung, A.Y.; Wu, H.M. FERONIA Receptor-like Kinase Regulates RHO GTPase Signaling of Root Hair Development. Proc. Natl. Acad. Sci. USA 2010, 107, 17821–17826. [Google Scholar] [CrossRef]
- Haruta, M.; Sabat, G.; Stecker, K.; Minkoff, B.B.; Sussman, M.R. A Peptide Hormone and Its Receptor Protein Kinase Regulate Plant Cell Expansion. Science 2014, 343, 408–411. [Google Scholar] [CrossRef]
- Yang, T.; Wang, L.; Li, C.; Liu, Y.; Zhu, S.; Qi, Y.; Liu, X.; Lin, Q.; Luan, S.; Yu, F. Receptor Protein Kinase FERONIA Controls Leaf Starch Accumulation by Interacting with Glyceraldehyde-3-Phosphate Dehydrogenase. Biochem. Biophys. Res. Commun. 2015, 465, 77–82. [Google Scholar] [CrossRef]
- Li, C.; Wu, H.M.; Cheung, A.Y. FERONIA and Her Pals: Functions and Mechanisms. Plant Physiol. 2016, 171, 2379–2392. [Google Scholar] [CrossRef]
- Stegmann, M.; Monaghan, J.; Smakowska-Luzan, E.; Rovenich, H.; Lehner, A.; Holton, N.; Belkhadir, Y.; Zipfel, C. The Receptor Kinase FER Is a RALF-Regulated Scaffold Controlling Plant Immune Signaling. Science 2017, 355, 287–289. [Google Scholar] [CrossRef]
- Ge, Z.; Bergonci, T.; Zhao, Y.; Zou, Y.; Du, S.; Liu, M.; Luo, X.; Ruan, H.; García-Valencia, L.E.; Zhong, S.; et al. Arabidopsis Pollen Tube Integrity and Sperm Release Are Regulated by RALF-Mediated Signaling. Science 2017, 358, 1596–1600. [Google Scholar] [CrossRef]
- Huck, N.; Moore, J.M.; Federer, M.; Grossniklaus, U. The Arabidopsis Mutant Feronia Disrupts the Female Gametophytic Control of Pollen Tube Reception. Development 2003, 130, 2149–2159. [Google Scholar] [CrossRef]
- Zhang, X.; Yang, Z.; Wu, D.; Yu, F. RALF–FERONIA Signaling: Linking Plant Immune Response with Cell Growth. Plant Commun. 2020, 1, 100084. [Google Scholar] [CrossRef]
- Kessler, S.A.; Shimosato-Asano, H.; Keinath, N.F.; Wuest, S.E.; Ingram, G.; Panstruga, R.; Grossniklaus, U. Conserved Molecular Components for Pollen Tube Reception and Fungal Invasion. Science 2010, 330, 968–971. [Google Scholar] [CrossRef]
- Guo, H.; Nolan, T.M.; Song, G.; Liu, S.; Xie, Z.; Chen, J.; Schnable, P.S.; Walley, J.W.; Yin, Y. FERONIA Receptor Kinase Contributes to Plant Immunity by Suppressing Jasmonic Acid Signaling in Arabidopsis thaliana. Curr. Biol. 2018, 28, 3316–3324. [Google Scholar] [CrossRef] [PubMed]
- Deslauriers, S.D.; Larsen, P.B. FERONIA Is a Key Modulator of Brassinosteroid and Ethylene Responsiveness in Arabidopsis Hypocotyls. Mol. Plant 2010, 3, 626–640. [Google Scholar] [CrossRef]
- Mao, D.; Yu, F.; Li, J.; Van de Poel, B.; Tan, D.; Li, J.; Liu, Y.; Li, X.; Dong, M.; Chen, L.; et al. FERONIA Receptor Kinase Interacts with S-Adenosylmethionine Synthetase and Suppresses S-Adenosylmethionine Production and Ethylene Biosynthesis in Arabidopsis. Plant Cell Environ. 2015, 38, 2566–2574. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yu, F.; Liu, Y.; Du, C.; Li, X.; Zhu, S.; Wang, X.; Lan, W.; Rodriguez, P.L.; Liu, X.; et al. FERONIA Interacts with ABI2-Type Phosphatases to Facilitate Signaling Cross-Talk between Abscisic Acid and RALF Peptide in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, E5519–E5527. [Google Scholar] [CrossRef] [PubMed]
- Masachis, S.; Segorbe, D.; Turrà, D.; Leon-Ruiz, M.; Fürst, U.; El Ghalid, M.; Leonard, G.; López-Berges, M.S.; Richards, T.A.; Felix, G.; et al. A Fungal Pathogen Secretes Plant Alkalinizing Peptides to Increase Infection. Nat. Microbiol. 2016, 1, 16043. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Cui, X.; Qin, G.; Chen, T.; Tian, S. SlFERL Interacts with S-Adenosylmethionine Synthetase to Regulate Fruit Ripening. Plant Physiol. 2020, 184, 2168–2181. [Google Scholar] [CrossRef]
- Yu, Y.; Chakravorty, D.; Assmann, S.M. The G Protein β-Subunit, AGB1, Interacts with FERONIA in RALF1-Regulated Stomatal Movement. Plant Physiol. 2018, 176, 2426–2440. [Google Scholar] [CrossRef] [PubMed]
- Ge, Z.; Dresselhaus, T.; Qu, L.J. How CrRLK1L Receptor Complexes Perceive RALF Signals. Trends Plant Sci. 2019, 24, 978–981. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Stegmann, M.; Han, Z.; DeFalco, T.A.; Parys, K.; Xu, L.; Belkhadir, Y.; Zipfel, C.; Chai, J. Mechanisms of RALF Peptide Perception by a Heterotypic Receptor Complex. Nature 2019, 572, 270–274. [Google Scholar] [CrossRef] [PubMed]
- Campbell, L.; Turner, S.R. A Comprehensive Analysis of RALF Proteins in Green Plants Suggests There Are Two Distinct Functional Groups. Front. Plant Sci. 2017, 8, 37. [Google Scholar] [CrossRef]
- Dünser, K.; Gupta, S.; Herger, A.; Feraru, M.I.; Ringli, C.; Kleine-Vehn, J. Extracellular Matrix Sensing by FERONIA and Leucine-Rich Repeat Extensins Controls Vacuolar Expansion during Cellular Elongation in Arabidopsis thaliana. EMBO J. 2019, 38, 1–12. [Google Scholar] [CrossRef]
- Du, C.; Li, X.; Chen, J.; Chen, W.; Li, B.; Li, C.; Wang, L.; Li, J.; Zhao, X.; Lin, J.; et al. Receptor Kinase Complex Transmits RALF Peptide Signal to Inhibit Root Growth in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, E8326–E8334. [Google Scholar] [CrossRef]
- Campos, W.F.; Dressano, K.; Ceciliato, P.H.O.; Guerrero-Abad, J.C.; Silva, A.L.; Fiori, C.S.; Morato do Canto, A.; Bergonci, T.; Claus, L.A.N.; Silva-Filho, M.C.; et al. Arabidopsis thaliana Rapid Alkalinization Factor 1–Mediated Root Growth Inhibition Is Dependent on Calmodulin-like Protein 38. J. Biol. Chem. 2018, 293, 2159–2171. [Google Scholar] [CrossRef]
- Li, C.; Liu, X.; Qiang, X.; Li, X.; Li, X.; Zhu, S.; Wang, L.; Wang, Y.; Liao, H.; Luan, S.; et al. EBP1 Nuclear Accumulation Negatively Feeds Back on FERONIA-Mediated RALF1 Signaling. PLoS Biol. 2018, 16, e2006340. [Google Scholar] [CrossRef]
- Yu, M.; Li, R.; Cui, Y.; Chen, W.; Li, B.; Zhang, X.; Bu, Y.; Cao, Y.; Xing, J.; Jewaria, P.K.; et al. The RALF1-FERONIA Interaction Modulates Endocytosis to Mediate Control of Root Growth in Arabidopsis. Development 2020, 147, dev189902. [Google Scholar] [CrossRef]
- Zhu, S.; Estévez, J.M.; Liao, H.; Zhu, Y.; Yang, T.; Li, C.; Wang, Y.; Li, L.; Liu, X.; Pacheco, J.M.; et al. The RALF1-FERONIA Complex Phosphorylates EIF4E1 to Promote Protein Synthesis and Polar Root Hair Growth. Mol. Plant 2020, 13, 698–716. [Google Scholar] [CrossRef] [PubMed]
- Hématy, K.; Sado, P.-E.; Van Tuinen, A.; Rochange, S.; Desnos, T.; Balzergue, S.; Pelletier, S.; Renou, J.-P.; Höfte, H. A Receptor-like Kinase Mediates the Response of Arabidopsis Cells to the Inhibition of Cellulose Synthesis. Curr. Biol. 2007, 17, 922–931. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Li, L.; Ye, H.; Yu, X.; Algreen, A.; Yin, Y. Three Related Receptor-like Kinases Are Required for Optimal Cell Elongation in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2009, 106, 7648–7653. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, S.; Murata, T.; Sakurai-Ozato, N.; Kubo, M.; Demura, T.; Fukuda, H.; Hasebe, M. ANXUR1 and 2, Sister Genes to FERONIA/SIRENE, Are Male Factors for Coordinated Fertilization. Curr. Biol. 2009, 19, 1327–1331. [Google Scholar] [CrossRef]
- Richter, J.; Watson, J.M.; Stasnik, P.; Borowska, M.; Neuhold, J.; Berger, M.; Stolt-Bergner, P.; Schoft, V.; Hauser, M.T. Multiplex Mutagenesis of Four Clustered CrRLK1L with CRISPR/Cas9 Exposes Their Growth Regulatory Roles in Response to Metal Ions. Sci. Rep. 2018, 8, 12182. [Google Scholar] [CrossRef]
- Barchi, L.; Rabanus-Wallace, M.T.; Prohens, J.; Toppino, L.; Padmarasu, S.; Portis, E.; Rotino, G.L.; Stein, N.; Lanteri, S.; Giuliano, G. Improved Genome Assembly and Pan-genome Provide Key Insights into Eggplant Domestication and Breeding. Plant J. 2021, 107, 579–596. [Google Scholar] [CrossRef]
- Wei, Q.; Wang, J.; Wang, W.; Hu, T.; Hu, H.; Bao, C. A High-Quality Chromosome-Level Genome Assembly Reveals Genetics for Important Traits in Eggplant. Hortic. Res. 2020, 7, 153. [Google Scholar] [CrossRef] [PubMed]
- Lindner, H.; Müller, L.M.; Boisson-Dernier, A.; Grossniklaus, U. CrRLK1L Receptor-like Kinases: Not Just Another Brick in the Wall. Curr. Opin. Plant Biol. 2012, 15, 659–669. [Google Scholar] [CrossRef]
- Nguyen, Q.-N.; Lee, Y.-S.; Cho, L.-H.; Jeong, H.-J.; An, G.; Jung, K.-H. Genome-Wide Identification and Analysis of Catharanthus Roseus RLK1-like Kinases in Rice. Planta 2015, 241, 603–613. [Google Scholar] [CrossRef]
- Zuo, C.; Zhang, W.; Ma, Z.; Chu, M.; Mao, J.; An, Z.; Chen, B. Genome-Wide Identification and Expression Analysis of the CrRLK1L Gene Family in Apple (Malus domestica). Plant Mol. Biol. Rep. 2018, 36, 844–857. [Google Scholar] [CrossRef]
- Zhang, Q.; Jia, M.; Xing, Y.; Qin, L.; Li, B.; Jia, W. Genome-Wide Identification and Expression Analysis of MRLK Family Genes Associated with Strawberry (Fragaria Vesca) Fruit Ripening and Abiotic Stress Responses. PLoS ONE 2016, 11, e0163647. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.-Q.; Yu, T.-F.; Sun, G.-Z.; Zheng, J.-C.; Chen, J.; Zhou, Y.-B.; Chen, M.; Ma, Y.-Z.; Wei, W.-L.; Xu, Z.-S. Genome-Wide Analysis of the Catharanthus roseus RLK1-Like in Soybean and GmCrRLK1L20 Responds to Drought and Salt Stresses. Front. Plant Sci. 2021, 12, 614909. [Google Scholar] [CrossRef]
- Luo, C.; Sun, Q.; Zhang, F.; Zhang, D.; Liu, C.; Wu, Q.; Shu, B. Genome-Wide Identification and Expression Analysis of the Citrus Malectin Domain-Containing Receptor-like Kinases in Response to Arbuscular Mycorrhizal Fungi Colonization and Drought. Hortic. Environ. Biotechnol. 2020, 61, 891–901. [Google Scholar] [CrossRef]
- Niu, E.; Cai, C.; Zheng, Y.; Shang, X.; Fang, L.; Guo, W. Genome-Wide Analysis of CrRLK1L Gene Family in Gossypium and Identification of Candidate CrRLK1L Genes Related to Fiber Development. Mol. Genet. Genom. 2016, 291, 1137–1154. [Google Scholar] [CrossRef] [PubMed]
- Kou, X.; Qi, K.; Qiao, X.; Yin, H.; Liu, X.; Zhang, S.; Wu, J. Evolution, Expression Analysis, and Functional Verification of Catharanthus roseus RLK1-like Kinase (CrRLK1L) Family Proteins in Pear (Pyrus bretchneideri). Genomics 2017, 109, 290–301. [Google Scholar] [CrossRef]
- Li, X.; Guo, C.; Wang, Q.; Li, Z.; Cai, J.; Wu, D.; Li, Y.; Yang, A.; Guo, Y.; Gao, J.; et al. Systematic Analysis of Tobacco CrRLK1L Family Genes and Functional Identification of NtCrRLK1L47 in Environmental Stresses. Front. Plant Sci. 2022, 13, 838857. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.-F.; Zhang, W.-N.; Kang, Y.-C.; Fan, Y.-L.; Yang, X.-Y.; Shi, M.-F.; Zhang, R.-Y.; Zhang, J.-L.; Qin, S.-H. Genome-Wide Identification and Expression Patterns in Response to Signals from Phytophthora infestans of CrRLK1Ls Gene Family in Potato. Acta Agron. Sin. 2021, 48, 249–258. [Google Scholar] [CrossRef]
- Ma, W.; Liu, X.; Chen, K.; Yu, X.; Ji, D. Genome-Wide Re-Identification and Analysis of CrRLK1Ls in Tomato. Int. J. Mol. Sci. 2023, 24, 3142. [Google Scholar] [CrossRef]
- Sakamoto, T.; Deguchi, M.; Brustolini, O.J.B.; Santos, A.A.; Silva, F.F.; Fontes, E.P.B. The Tomato RLK Superfamily: Phylogeny and Functional Predictions about the Role of the LRRII-RLK Subfamily in Antiviral Defense. BMC Plant Biol. 2012, 12, 229. [Google Scholar] [CrossRef]
- Guruprasad, K.; Reddy, B.V.B.; Pandit, M.W. Correlation between Stability of a Protein and Its Dipeptide Composition: A Novel Approach for Predicting in Vivo Stability of a Protein from Its Primary Sequence. Protein Eng. Des. Sel. 1990, 4, 155–161. [Google Scholar] [CrossRef]
- Dean, R.; Van Kan, J.A.L.; Pretorius, Z.A.; Hammond-Kosack, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The Top 10 Fungal Pathogens in Molecular Plant Pathology. Mol. Plant Pathol. 2012, 13, 414–430. [Google Scholar] [CrossRef] [PubMed]
- Fillinger, S.; Elad, Y. Botrytis—The Fungus, the Pathogen and Its Management in Agricultural Systems; Fillinger, S., Elad, Y., Eds.; Springer International Publishing: Cham, Switzerland, 2016; ISBN 978-3-319-23370-3. [Google Scholar]
- Ji, D.; Liu, W.; Cui, X.; Liu, K.; Liu, Y.; Huang, X.; Li, B.; Qin, G.; Chen, T.; Tian, S. A Receptor-like Kinase SlFERL Mediates Immune Responses of Tomato to Botrytis cinerea by Recognizing BcPG1 and Fine-tuning MAPK Signaling. New Phytol. 2023, nph.19210. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.J.; Zhang, X.S.; Gao, X.-Q. ROS in the Male–Female Interactions During Pollination: Function and Regulation. Front. Plant Sci. 2020, 11, 177. [Google Scholar] [CrossRef]
- Jia, M.; Du, P.; Ding, N.; Zhang, Q.; Xing, S.; Wei, L.; Zhao, Y.; Mao, W.; Li, J.; Li, B.; et al. Two FERONIA-like Receptor Kinases Regulate Apple Fruit Ripening by Modulating Ethylene Production. Front. Plant Sci. 2017, 8, 1406. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, L.; Cui, Y.; He, L.; Qi, Y.; Zhang, J.; Lin, J.; Liao, H.; Lin, Q.; Yang, T.; et al. Two FERONIA-like Receptor (FLR) Genes Are Required to Maintain Architecture, Fertility, and Seed Yield in Rice. Mol. Breed. 2016, 36, 151. [Google Scholar] [CrossRef]
- Pu, C.X.; Han, Y.F.; Zhu, S.; Song, F.Y.; Zhao, Y.; Wang, C.; Zhang, Y.; Yang, Q.; Wang, J.; Bu, S.-L.; et al. The Rice Receptor-like Kinases DWARF AND RUNTISH SPIKELET1 and 2 Repress Cell Death and Affect Sugar Utilization during Reproductive Development. Plant Cell 2017, 29, 70–89. [Google Scholar] [CrossRef]
- Liu, L.; Zheng, C.; Kuang, B.; Wei, L.; Yan, L.; Wang, T. Receptor-like Kinase RUPO Interacts with Potassium Transporters to Regulate Pollen Tube Growth and Integrity in Rice. PLoS Genet. 2016, 12, e1006085. [Google Scholar] [CrossRef]
- Wang, D.; Liang, X.; Bao, Y.; Yang, S.; Zhang, X.; Yu, H.; Zhang, Q.; Xu, G.; Feng, X.; Dou, D. A Malectin-like Receptor Kinase Regulates Cell Death and Pattern-triggered Immunity in Soybean. EMBO Rep. 2020, 21, e50442. [Google Scholar] [CrossRef]
- Rao, S.; Wu, X.; Zheng, H.; Lu, Y.; Peng, J.; Wu, G.; Chen, J.; Yan, F. Genome-Wide Identification and Analysis of Catharanthus Roseus RLK1-like Kinases in Nicotiana benthamiana. BMC Plant Biol. 2021, 21, 425. [Google Scholar] [CrossRef]
- Jiang, W.; Li, C.; Li, L.; Li, Y.; Wang, Z.; Yu, F.; Yi, F.; Zhang, J.; Zhu, J.-K.; Zhang, H.; et al. Genome-Wide Analysis of CqCrRLK1L and CqRALF Gene Families in Chenopodium quinoa and Their Roles in Salt Stress Response. Front. Plant Sci. 2022, 13, 918594. [Google Scholar] [CrossRef]
- Li, X.; Guo, C.; Ahmad, S.; Wang, Q.; Yu, J.; Liu, C.; Guo, Y. Systematic Analysis of MYB Family Genes in Potato and Their Multiple Roles in Development and Stress Responses. Biomolecules 2019, 9, 317. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Madeira, F.; Pearce, M.; Tivey, A.R.N.; Basutkar, P.; Lee, J.; Edbali, O.; Madhusoodanan, N.; Kolesnikov, A.; Lopez, R. Search and Sequence Analysis Tools Services from EMBL-EBI in 2022. Nucleic Acids Res. 2022, 50, W276–W279. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.-S.; Chen, Y.-C.; Lu, C.-H.; Hwang, J.-K. Prediction of Protein Subcellular Localization. Proteins 2006, 64, 643–651. [Google Scholar] [CrossRef]


| Protein Name | SGN Locus | Protein Length | MW | Theoretical pI | Instability Index | Aliphatic Index | GRAVY |
|---|---|---|---|---|---|---|---|
| SmCrRLK1L1 | SMEL4.1_00g002980.1.01 | 916 | 102,981.91 | 9.05 | 38.81 | 89.29 | −0.244 |
| SmCrRLK1L2 | SMEL4.1_00g011070.1.01 | 904 | 100,649.88 | 5.78 | 40.63 | 84.55 | −0.284 |
| SmCrRLK1L3 | SMEL4.1_01g004620.1.01 | 878 | 96,435.51 | 6.03 | 42.89 | 84.54 | −0.164 |
| SmCrRLK1L4 | SMEL4.1_01g027970.1.01 | 850 | 94,767.29 | 6.48 | 44.05 | 80.29 | −0.256 |
| SmCrRLK1L5 | SMEL4.1_02g011520.1.01 | 990 | 111,146.47 | 8.55 | 35.88 | 89.65 | −0.252 |
| SmCrRLK1L6 | SMEL4.1_02g011530.1.01 | 780 | 86,927.27 | 8.05 | 27.96 | 88.64 | −0.27 |
| SmCrRLK1L7 | SMEL4.1_02g011560.1.01 | 956 | 106,793.65 | 5.85 | 36.2 | 90.76 | −0.232 |
| SmCrRLK1L8 | SMEL4.1_02g011630.1.01 | 443 | 49,500.78 | 9.02 | 37.46 | 90.52 | −0.258 |
| SmCrRLK1L9 | SMEL4.1_02g011650.1.01 | 930 | 103,516.67 | 7.27 | 35.74 | 92.49 | −0.151 |
| SmCrRLK1L10 | SMEL4.1_02g013460.1.01 | 868 | 96,482.78 | 5.66 | 40.89 | 81.12 | −0.273 |
| SmCrRLK1L11 | SMEL4.1_02g026370.1.01 | 1026 | 114,710.11 | 5.84 | 40.63 | 81.01 | −0.273 |
| SmCrRLK1L12 | SMEL4.1_02g026380.1.01 | 997 | 112,040.66 | 6.38 | 41.89 | 84.84 | −0.259 |
| SmCrRLK1L13 | SMEL4.1_03g004220.1.01 | 827 | 92,723.16 | 5.58 | 40.19 | 86.09 | −0.207 |
| SmCrRLK1L14 | SMEL4.1_03g014350.1.01 | 610 | 67,859.65 | 5.85 | 31.72 | 96.05 | −0.06 |
| SmCrRLK1L15 | SMEL4.1_03g027050.1.01 | 850 | 93,804.86 | 6.03 | 38.84 | 86.24 | −0.09 |
| SmCrRLK1L16 | SMEL4.1_03g032580.1.01 | 834 | 93,418.44 | 5.78 | 32.53 | 94.21 | −0.14 |
| SmCrRLK1L17 | SMEL4.1_04g002370.1.01 | 840 | 91,907.43 | 5.27 | 37.89 | 94.46 | 0.025 |
| SmCrRLK1L18 | SMEL4.1_05g003720.1.01 | 820 | 90,378.18 | 6.53 | 37.67 | 89.37 | −0.065 |
| SmCrRLK1L19 | SMEL4.1_05g004710.1.01 | 819 | 91,513.23 | 5.65 | 34.6 | 90.13 | −0.086 |
| SmCrRLK1L20 | SMEL4.1_06g000710.1.01 | 866 | 95,410.66 | 6.42 | 38.33 | 80.25 | −0.298 |
| SmCrRLK1L21 | SMEL4.1_06g000720.1.01 | 911 | 100,056.89 | 6.11 | 34.63 | 82.52 | −0.266 |
| SmCrRLK1L22 | SMEL4.1_06g004680.1.01 | 829 | 90,489.55 | 5.8 | 36.94 | 85.11 | −0.077 |
| SmCrRLK1L23 | SMEL4.1_07g012220.1.01 | 885 | 96,959.49 | 5.77 | 43.66 | 79.51 | −0.209 |
| SmCrRLK1L24 | SMEL4.1_07g020590.1.01 | 983 | 108,300.31 | 8.38 | 30.1 | 94.31 | −0.108 |
| SmCrRLK1L25 | SMEL4.1_07g023150.1.01 | 843 | 92,227.57 | 5.51 | 37.69 | 89.18 | −0.078 |
| SmCrRLK1L26 | SMEL4.1_07g026240.1.01 | 1058 | 116,977.8 | 6.3 | 31.28 | 92.28 | −0.139 |
| SmCrRLK1L27 | SMEL4.1_09g003310.1.01 | 928 | 102,821.64 | 5.61 | 35.6 | 90.15 | −0.181 |
| SmCrRLK1L28 | SMEL4.1_09g010930.1.01 | 888 | 97,206.73 | 5.69 | 42.59 | 80.01 | −0.207 |
| SmCrRLK1L29 | SMEL4.1_09g012150.1.01 | 1084 | 120,775.58 | 5.77 | 46.93 | 97.1 | −0.09 |
| SmCrRLK1L30 | SMEL4.1_10g010500.1.01 | 812 | 90,741.61 | 6.99 | 32.04 | 86.51 | −0.118 |
| SmCrRLK1L31 | SMEL4.1_12g003870.1.01 | 847 | 93,940.44 | 5.44 | 35.97 | 81.89 | −0.188 |
| SmCrRLK1L32 | SMEL4.1_12g012720.1.01 | 837 | 91,980.58 | 6.39 | 35.59 | 82.75 | −0.125 |
| Protein Name | Signal Peptide Number | TMhelix Number | Most Likely Location |
|---|---|---|---|
| SmCrRLK1L1 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L2 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L3 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L4 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L5 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L6 | 1 | 1 | Mitochondrial |
| SmCrRLK1L7 | 1 | 1 | Cytoplasmic |
| SmCrRLK1L8 | 0 | 1 | Plasma Membrane |
| SmCrRLK1L9 | 0 | 1 | Plasma Membrane |
| SmCrRLK1L10 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L11 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L12 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L13 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L14 | 0 | 1 | Plasma Membrane |
| SmCrRLK1L15 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L16 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L17 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L18 | 0 | 1 | Plasma Membrane |
| SmCrRLK1L19 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L20 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L21 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L22 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L23 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L24 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L25 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L26 | 0 | 1 | Plasma Membrane |
| SmCrRLK1L27 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L28 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L29 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L30 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L31 | 1 | 1 | Plasma Membrane |
| SmCrRLK1L32 | 1 | 1 | Plasma Membrane |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, W.; Du, J.; Yu, X.; Chen, K.; Ming, Y.; Jiang, L.; Chen, T.; Ji, D. Genome-Wide Identification and Analysis of Catharanthus roseus Receptor-like Kinase 1-like Proteins in Eggplant. Plants 2023, 12, 3379. https://doi.org/10.3390/plants12193379
Ma W, Du J, Yu X, Chen K, Ming Y, Jiang L, Chen T, Ji D. Genome-Wide Identification and Analysis of Catharanthus roseus Receptor-like Kinase 1-like Proteins in Eggplant. Plants. 2023; 12(19):3379. https://doi.org/10.3390/plants12193379
Chicago/Turabian StyleMa, Wenpeng, Juan Du, Xinlong Yu, Kai Chen, Yucheng Ming, Libo Jiang, Tong Chen, and Dongchao Ji. 2023. "Genome-Wide Identification and Analysis of Catharanthus roseus Receptor-like Kinase 1-like Proteins in Eggplant" Plants 12, no. 19: 3379. https://doi.org/10.3390/plants12193379







