QTLs and Candidate Genes for Seed Protein Content in Two Recombinant Inbred Line Populations of Soybean
Abstract
:1. Introduction
2. Results
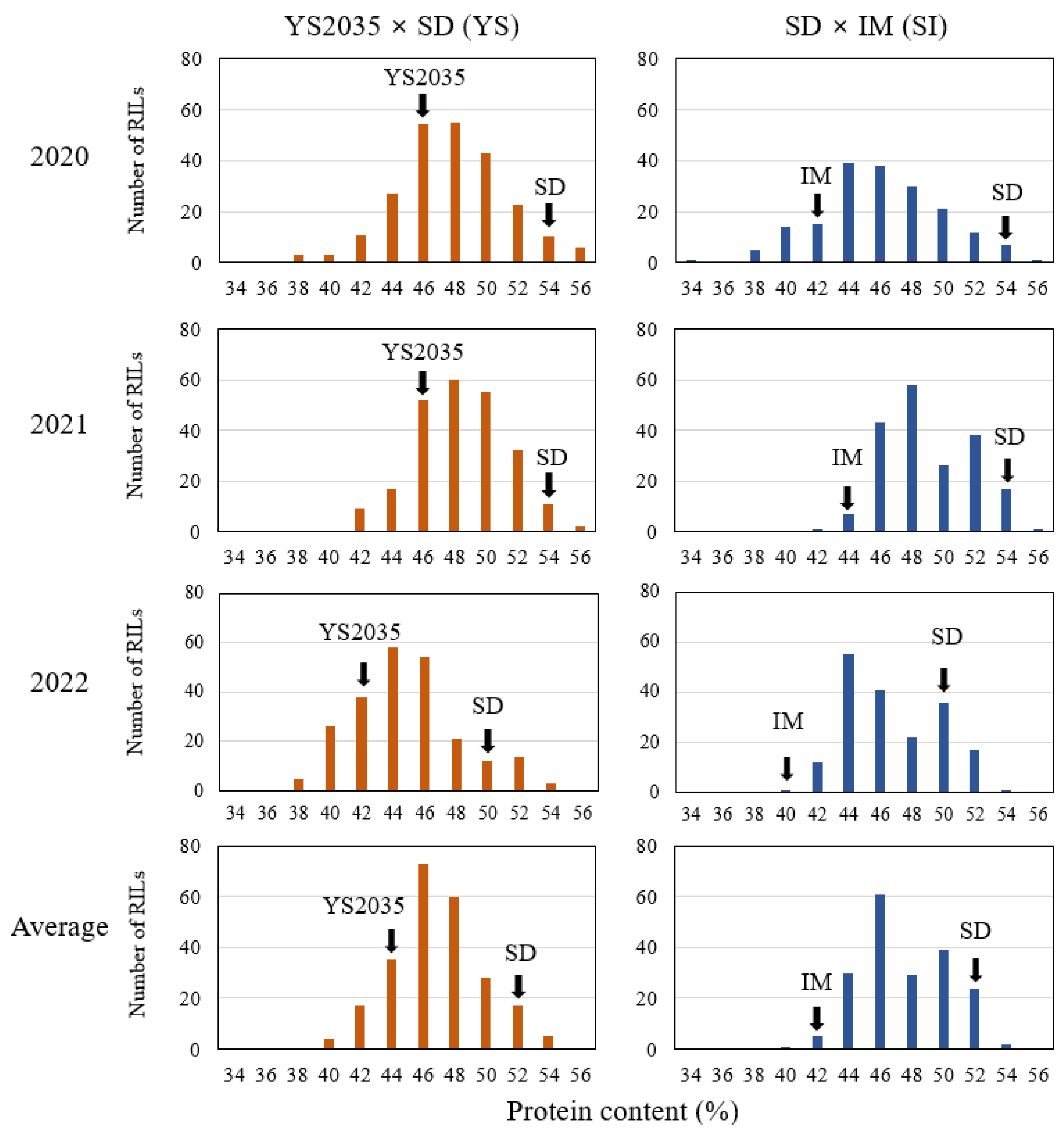
2.1. Phenotypic Variation in the Seed Protein Content
2.2. Linkage Map Construction
2.3. QTL Analysis
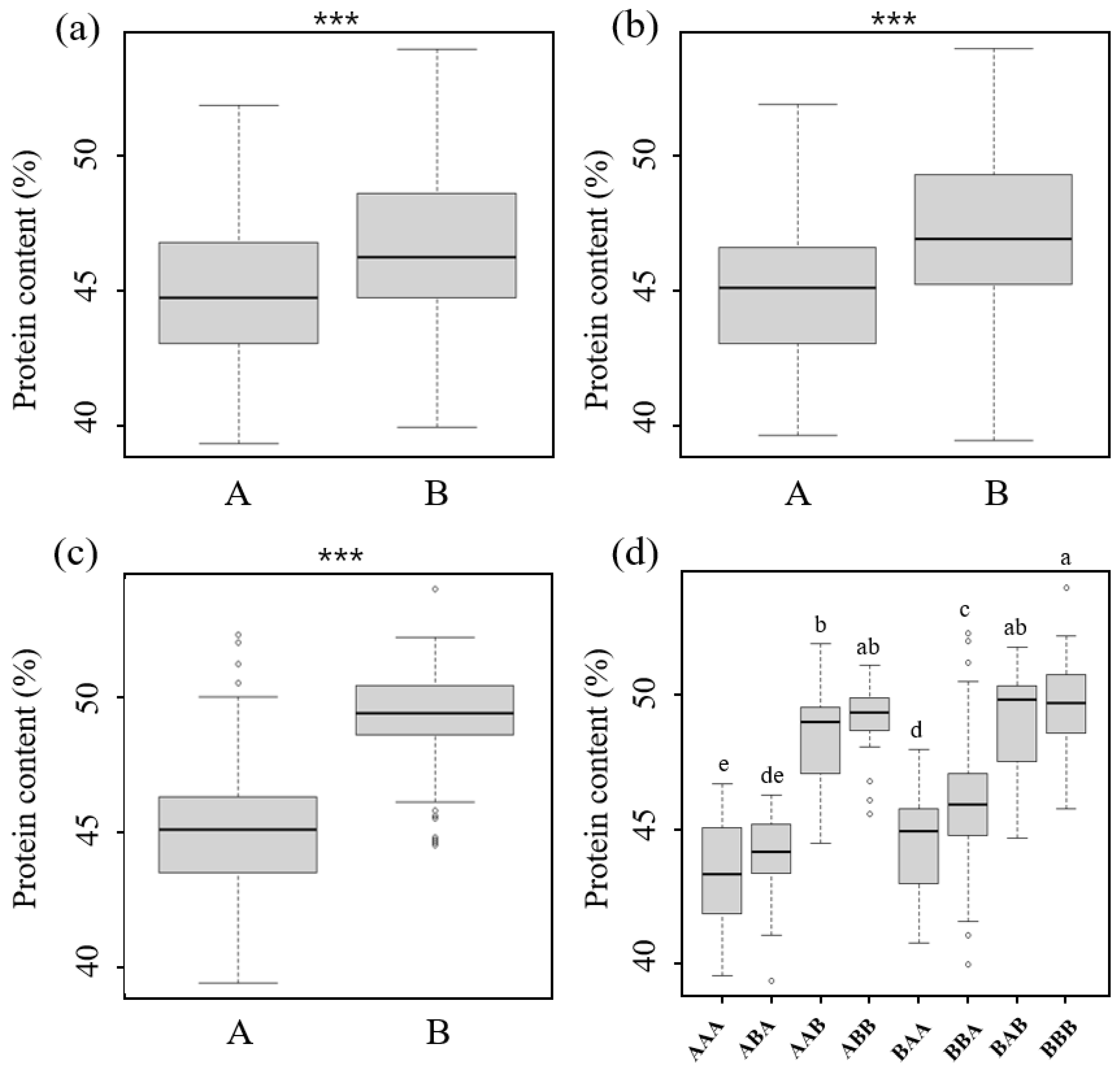
2.4. Phenotypic Variation According to the Allele Patterns
2.5. SNP Variation Analysis and Variant Annotation
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Analysis of Crude Seed Protein Concentrations
4.3. Genomic DNA Extraction and Genotyping
4.4. Genetic Linkage Map Construction and QTL Analysis
4.5. Prediction of Novel Candidate QTL and Genes
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Henchion, M.; Hayes, M.; Mullen, A.M.; Fenelon, M.; Tiwari, B. Future Protein Supply and Demand: Strategies and Factors Influencing a Sustainable Equilibrium. Foods 2017, 6, 53. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.K.; Manjaya, J.G. Advances in improvement of soybean seed composition traits using genetic, genomic and biotechnological approaches. Euphytica 2022, 218, 99. [Google Scholar] [CrossRef]
- Qin, P.; Wang, T.; Luo, Y. A review on plant-based proteins from soybean: Health benefits and soy product development. J. Agric. Food Res. 2022, 7, 100265. [Google Scholar] [CrossRef]
- Liu, S.; Liu, Z.; Hou, X.; Li, X. Genetic mapping and functional genomics of soybean seed protein. Mol. Breed. 2023, 43, 29. [Google Scholar] [CrossRef] [PubMed]
- Thrane, M.; Paulsen, P.V.; Orcutt, M.W.; Krieger, T.M. Soy Protein: Impacts, Production, and Applications. In Sustainable Protein Sources; Elsevier: Amsterdam, The Netherlands, 2017; pp. 23–45. ISBN 9780128027769. [Google Scholar]
- Soystats. 2023 SOYSTATS: A Reference Guide to Important Soybean Facts and Figures; American Soybean Association: St. Louis, MO, USA, 2023. [Google Scholar]
- Caputo, V.; Sogari, G.; Van Loo, E.J. Do plant-based and blend meat alternatives taste like meat? A combined sensory and choice experiment study. Appl. Econ. Perspect. Policy 2022, 45, 86–105. [Google Scholar] [CrossRef]
- Zhang, T.; Dou, W.; Zhang, X.; Zhao, Y.; Zhang, Y.; Jiang, L.; Sui, X. The Development History and Recent Updates on Soy Protein-Based Meat Alternatives. Trends Food Sci. Technol. 2021, 109, 702–710. [Google Scholar] [CrossRef]
- Mariotti, F.; Gardner, C.D. Dietary Protein and Amino Acids in Vegetarian Diets-A Review. Nutrients 2019, 11, 2661. [Google Scholar] [CrossRef]
- Detzel, A.; Kruger, M.; Busch, M.; Blanco-Gutierrez, I.; Varela, C.; Manners, R.; Bez, J.; Zannini, E. Life cycle assessment of animal-based foods and plant-based protein-rich alternatives: An environmental perspective. J. Sci. Food Agric. 2022, 102, 5098–5110. [Google Scholar] [CrossRef]
- Bolon, Y.T.; Joseph, B.; Cannon, S.B.; Graham, M.A.; Diers, B.W.; Farmer, A.D.; May, G.D.; Muehlbauer, G.J.; Specht, J.E.; Tu, Z.J.; et al. Complementary Genetic and Genomic Approaches Help Characterize the Linkage Group I Seed Protein QTL in Soybean. BMC Plant Biol. 2010, 10, 41. [Google Scholar] [CrossRef]
- Hwang, E.-Y.; Song, Q.; Jia, G.; Specht, J.E.; Hyten, D.L.; Costa, J.; Cregan, P.B. A Genome-Wide Associa-tion Study of Seed Protein and Oil Content in Soybean. BMC Genom. 2014, 15, 1. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef] [PubMed]
- Marsh, J.I.; Hu, H.; Petereit, J.; Bayer, P.E.; Valliyodan, B.; Batley, J.; Nguyen, H.T.; Edwards, D. Haplotype mapping uncovers unexplored variation in wild and domesticated soybean at the major protein locus cqProt-003. Theor. Appl. Genet. 2022, 135, 1443–1455. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.J.; Kang, B.H.; Moon, C.Y.; Kang, S.; Shin, S.; Chowdhury, S.; Choi, M.S.; Park, S.K.; Moon, J.K.; Ha, B.K. Quantitative Trait Loci (QTL) Analysis of Seed Protein and Oil Content in Wild Soybean (Glycine soja). Int. J. Mol. Sci. 2023, 24, 4077. [Google Scholar] [CrossRef]
- Diers, B.W.; Keim, P.; Fehr, W.R.; Shoemaker, R.C. RFLP Analysis of Soybean Seed Protein and Oil Content. Theor. Appl. Genet. 1992, 83, 608–612. [Google Scholar] [CrossRef]
- Kim, M.; Schultz, S.; Nelson, R.L.; Diers, B.W. Identification and Fine Mapping of a Soybean Seed Protein QTL from PI 407788A on Chromosome 15. Crop Sci. 2016, 56, 219–225. [Google Scholar] [CrossRef]
- Brzostowski, L.F.; Diers, B.W. Agronomic Evaluation of a High Protein Allele from PI407788A on Chromosome 15 across Two Soybean Backgrounds. Crop Sci. 2017, 57, 2972–2978. [Google Scholar] [CrossRef]
- Seo, J.-H.; Kim, K.-S.; Ko, J.-M.; Choi, M.-S.; Kang, B.-K.; Kwon, S.-W.; Jun, T.-H.; Singh, R. Quantitative trait locus analysis for soybean (Glycine max) seed protein and oil concentrations using selected breeding populations. Plant Breed. 2019, 138, 95–104. [Google Scholar] [CrossRef]
- Warrington, C.V.; Abdel-Haleem, H.; Hyten, D.L.; Cregan, P.B.; Orf, J.H.; Killam, A.S.; Bajjalieh, N.; Li, Z.; Boerma, H.R. QTL for seed protein and amino acids in the Benning x Danbaekkong soybean population. Theor. Appl. Genet. 2015, 128, 839–850. [Google Scholar] [CrossRef]
- Chung, J.; Babka, H.L.; Graef, G.L.; Staswick, P.E.; Lee, D.J.; Cregan, P.B.; Shoemaker, R.C.; Specht, J.E. The Seed Protein, Oil, and Yield QTL on Soybean Linkage Group I. Crop Sci. 2003, 43, 1053–1067. [Google Scholar] [CrossRef]
- Pandurangan, S.; Pajak, A.; Molnar, S.J.; Cober, E.R.; Dhaubhadel, S.; Hernandez-Sebastia, C.; Kaiser, W.M.; Nelson, R.L.; Huber, S.C.; Marsolais, F. Relationship between asparagine metabolism and protein concentration in soybean seed. J. Exp. Bot. 2012, 63, 3173–3184. [Google Scholar] [CrossRef]
- Mao, T.; Jiang, Z.; Han, Y.; Teng, W.; Zhao, X.; Li, W.; Morris, B. Identification of quantitative trait loci underlying seed protein and oil contents of soybean across multi-genetic backgrounds and environments. Plant Breed. 2013, 132, 630–641. [Google Scholar] [CrossRef]
- Fliege, C.E.; Ward, R.A.; Vogel, P.; Nguyen, H.; Quach, T.; Guo, M.; Viana, J.P.G.; Dos Santos, L.B.; Specht, J.E.; Clemente, T.E.; et al. Fine mapping and cloning of the major seed protein quantitative trait loci on soybean chromosome 20. Plant J. 2022, 110, 114–128. [Google Scholar] [CrossRef] [PubMed]
- Goettel, W.; Zhang, H.; Li, Y.; Qiao, Z.; Jiang, H.; Hou, D.; Song, Q.; Pantalone, V.R.; Song, B.H.; Yu, D.; et al. POWR1 is a domestication gene pleiotropically regulating seed quality and yield in soybean. Nat. Commun. 2022, 13, 3051. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Wang, W.; He, Q.; Xiang, S.; Tian, D.; Zhao, T.; Gai, J. Identifying a wild allele conferring small seed size, high protein content and low oil content using chromosome segment substitution lines in soybean. Theor. Appl. Genet. 2019, 132, 2793–2807. [Google Scholar] [CrossRef] [PubMed]
- Wilcox, J.R.; Guodong, Z. Relationships between Seed Yield and Seed Protein in Determinate and Indeterminate Soybean Populations. Crop Sci. 1997, 37, 361–364. [Google Scholar] [CrossRef]
- Cober, E.R.; Voldeng, H.D. Developing High-Protein, High-Yield Soybean Populations and Lines. Crop Sci. 2000, 40, 39–42. [Google Scholar] [CrossRef]
- Miao, L.; Yang, S.; Zhang, K.; He, J.; Wu, C.; Ren, Y.; Gai, J.; Li, Y. Natural variation and selection in GmSWEET39 affect soybean seed oil content. N. Phytol. 2020, 225, 1651–1666. [Google Scholar] [CrossRef]
- Wang, S.; Liu, S.; Wang, J.; Yokosho, K.; Zhou, B.; Yu, Y.C.; Liu, Z.; Frommer, W.B.; Ma, J.F.; Chen, L.Q.; et al. Simultaneous changes in seed size, oil content and protein content driven by selection of SWEET homologues during soybean domestication. Natl. Sci. Rev. 2020, 7, 1776–1786. [Google Scholar] [CrossRef]
- Zhang, H.; Goettel, W.; Song, Q.; Jiang, H.; Hu, Z.; Wang, M.L.; An, Y.C. Selection of GmSWEET39 for oil and protein improvement in soybean. PLoS Genet. 2020, 16, e1009114. [Google Scholar] [CrossRef]
- Duan, Z.; Li, Q.; Wang, H.; He, X.; Zhang, M. Genetic regulatory networks of soybean seed size, oil and protein contents. Front. Plant Sci. 2023, 14, 1160418. [Google Scholar] [CrossRef]
- Patil, G.; Mian, R.; Vuong, T.; Pantalone, V.; Song, Q.; Chen, P.; Shannon, G.J.; Carter, T.C.; Nguyen, H.T. Molecular mapping and genomics of soybean seed protein: A review and perspective for the future. Theor. Appl. Genet. 2017, 130, 1975–1991. [Google Scholar] [CrossRef] [PubMed]
- Van, K.; McHale, L.K. Meta-Analyses of QTLs Associated with Protein and Oil Contents and Compositions in Soybean [Glycine max (L.) Merr.] Seed. Int. J. Mol. Sci. 2017, 18, 1180. [Google Scholar] [CrossRef] [PubMed]
- Brzostowski, L.F.; Pruski, T.I.; Specht, J.E.; Diers, B.W. Impact of seed protein alleles from three soybean sources on seed composition and agronomic traits. Theor. Appl. Genet. 2017, 130, 2315–2326. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Choi, M.-S.; Kim, H.-T.; Yun, H.-T.; Lee, B.; Chung, Y.-S.; Kim, R.W.; Choi, H.-K. Soybean [Glycine max (L.) Merrill]: Importance as A Crop and Pedigree Reconstruction of Korean Varieties. Plant Breed. Biotechnol. 2015, 3, 179–196. [Google Scholar] [CrossRef]
- Lee, J.S.; Kim, S.-M.; Kang, S. Fine mapping of quantitative trait loci for sucrose and oligosaccharide contents in soybean [Glycine max (L.) Merr.] using 180 K Axiom® SoyaSNP genotyping platform. Euphytica 2015, 208, 195–203. [Google Scholar] [CrossRef]
- Wang, Z.; Yu, D.; Morota, G.; Dhakal, K.; Singer, W.; Lord, N.; Huang, H.; Chen, P.; Mozzoni, L.; Li, S.; et al. Genome-wide association analysis of sucrose and alanine contents in edamame beans. Front. Plant Sci. 2022, 13, 1086007. [Google Scholar] [CrossRef]
- Arias, C.L.; Quach, T.; Huynh, T.; Nguyen, H.; Moretti, A.; Shi, Y.; Guo, M.; Rasoul, A.; Van, K.; McHale, L.; et al. Expression of AtWRI1 and AtDGAT1 during soybean embryo development influences oil and carbohydrate metabolism. Plant Biotechnol. J. 2022, 20, 1327–1345. [Google Scholar] [CrossRef]
- Xu, W.; Wang, Q.; Zhang, W.; Zhang, H.; Liu, X.; Song, Q.; Zhu, Y.; Cui, X.; Chen, X.; Chen, H. Using transcriptomic and metabolomic data to investigate the molecular mechanisms that determine protein and oil contents during seed development in soybean. Front. Plant Sci. 2022, 13, 1012394. [Google Scholar] [CrossRef]
- Wang, J.; Mao, L.; Zeng, Z.; Yu, X.; Lian, J.; Feng, J.; Yang, W.; An, J.; Wu, H.; Zhang, M.; et al. Genetic mapping high protein content QTL from soybean ‘Nanxiadou 25′ and candidate gene analysis. BMC Plant Biol. 2021, 21, 388. [Google Scholar] [CrossRef]
- Du, J.; Wang, S.; He, C.; Zhou, B.; Ruan, Y.L.; Shou, H. Identification of regulatory networks and hub genes controlling soybean seed set and size using RNA sequencing analysis. J. Exp. Bot. 2017, 68, 1955–1972. [Google Scholar] [CrossRef]
- Hooker, J.C.; Nissan, N.; Luckert, D.; Zapata, G.; Hou, A.; Mohr, R.M.; Glenn, A.J.; Barlow, B.; Daba, K.A.; Warkentin, T.D.; et al. GmSWEET29 and Paralog GmSWEET34 Are Differentially Expressed between Soybeans Grown in Eastern and Western Canada. Plants 2022, 11, 2337. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.T.; Ko, J.M.; Baek, I.Y.; Jeon, M.K.; Han, W.Y.; Park, K.Y.; Lee, B.W.; Lee, Y.H.; Jung, C.S.; Oh, K.W.; et al. Soybean Cultivar for Tofu, ‘Saedanbaek’ with Disease Resistance, and High Protein Content. Korean J. Breed. Sci. 2014, 46, 295–301. [Google Scholar] [CrossRef]
- Natarajan, S.; Luthria, D.; Bae, H.; Lakshman, D.; Mitra, A. Transgenic soybeans and soybean protein analysis: An overview. J. Agric. Food Chem. 2013, 61, 11736–11743. [Google Scholar] [CrossRef] [PubMed]
- Bandillo, N.; Jarquin, D.; Song, Q.; Nelson, R.; Cregan, P.; Specht, J.; Lorenz, A. A Population Structure and Genome-Wide Association Analysis on the USDA Soybean Germplasm Collection. Plant Genome 2015, 8, e0024. [Google Scholar] [CrossRef]
- Song, Q.; Hyten, D.L.; Jia, G.; Quigley, C.V.; Fickus, E.W.; Nelson, R.L.; Cregan, P.B. Fingerprinting Soybean Germplasm and Its Utility in Genomic Research. G3 2015, 5, 1999–2006. [Google Scholar] [CrossRef]
- Brummer, E.C.; Graef, G.L.; Orf, J.; Wilcox, J.R.; Shoemaker, R.C. Mapping QTL for Seed Protein and Oil Content in Eight Soybean Populations. Crop Sci. 1997, 37, 370–378. [Google Scholar] [CrossRef]
- Li, L.; Zheng, W.; Zhu, Y.; Ye, H.; Tang, B.; Arendsee, Z.W.; Jones, D.; Li, R.; Ortiz, D.; Zhao, X.; et al. QQS orphan gene regulates carbon and nitrogen partitioning across species via NF-YC interactions. Proc. Natl. Acad. Sci. USA 2015, 112, 14734–14739. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, W.; Lin, Y.; Zhang, L.; Wang, C.; Xu, R. Construction of a high-density genetic map and mapping of QTLs for soybean (Glycine max) agronomic and seed quality traits by specific length amplified fragment sequencing. BMC Genom. 2018, 19, 641. [Google Scholar] [CrossRef]
- Kambhampati, S.; Aznar-Moreno, J.A.; Bailey, S.R.; Arp, J.J.; Chu, K.L.; Bilyeu, K.D.; Durrett, T.P.; Allen, D.K. Temporal changes in metabolism late in seed development affect biomass composition. Plant Physiol. 2021, 186, 874–890. [Google Scholar] [CrossRef]
- Shin, D.C.; Baek, I.Y.; Kang, S.T.; Song, S.B.; Hur, S.O.; Kwack, Y.H.; Lim, M.S. A New Disease and Lodging Resistance, High Yielding Soybean Variety “Ilmikong”. Korean J. Breed. Sci. 1998, 30, 397. [Google Scholar]
- RDA (Rural Development Administration). Agricultural Science Technology Standards for Investigation of Research; Rural Development Administration: Jeonju, Republic of Korea, 2012.
- Saint-Denis, T.; Goupy, J. Optimization of a nitrogen analyser based on the Dumas method. Anal. Chim. Acta 2004, 515, 191–198. [Google Scholar] [CrossRef]
- Dhungana, S.K.; Seo, J.-H.; Kang, B.-K.; Park, J.-H.; Kim, J.-H.; Sung, J.-S.; Back, I.-Y.; Shin, S.-O.; Jung, C.-S. Protein, Amino Acid, Oil, Fatty Acid, Sugar, Anthocyanin, Isoflavone, Lutein, and Antioxidant Variations in Colored Seed-Coated Soybeans. Plants 2021, 10, 1765. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.G.; Jeong, N.; Kim, J.H.; Lee, K.; Kim, K.H.; Pirani, A.; Ha, B.K.; Kang, S.T.; Park, B.S.; Moon, J.K.; et al. Development, validation and genetic analysis of a large soybean SNP genotyping array. Plant J. 2015, 81, 625–636. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Li, H.; Zhang, L.; Wang, J. QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 2015, 3, 269–283. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. 2021. Available online: https://www.r-project.org (accessed on 20 May 2023).
- Toker, C. Estimates of broad-sense heritability for seed yield and yield criteria in faba bean (Vicia faba L.). Hereditas 2004, 140, 222–225. [Google Scholar] [CrossRef]
- Dhungana, S.K.; Park, J.-H.; Oh, J.-H.; Kang, B.-K.; Seo, J.-H.; Sung, J.-S.; Kim, H.-S.; Shin, S.-O.; Back, I.-Y.; Jung, C.-S. Quantitative Trait Locus Mapping for Drought Tolerance in Soybean Recombinant Inbred Line Population. Plants 2021, 10, 1816. [Google Scholar] [CrossRef]

| Population 1 | Marker 2 | Chr 3 | Genetic Position (cM) | Physical Position of Markers (bp) 4 | Year | Gene Name | Gene No. | LOD 5 | PVE 6 (%) | Add 7 | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Y × S | qPSD15-1 | 15 | 305 | 7,930,801–8,678,412 | 2020 2021 Average 8 | Glyma.15g101800–Glyma.15g110600 | 89 | 14.0 14.6 12.3 | 17.5 17.1 13.8 | −2.7 −2.1 −1.7 | |
| Y × S | qPSD18-1 | 18 | 75 | 46,911,930–47,526,734 | 2022 Average | Glyma.18g193300–Glyma.18g197100 | 39 | 6.7 5.9 | 7.0 5.5 | −0.9 −0.6 | [38,39,40] |
| Y × S | qPSD20-1 | 20 | 96 | 31,781,045–31,961,695 | 2020 2021 2022 Average | Glyma.20g085100–Glyma.20g085700 | 7 | 21.1 24.7 20.9 30.6 | 22.5 29.1 24.7 35.4 | −1.8 −1.6 −1.8 −1.6 | [14,24,25] |
| S × I | qPSD20-2 | 20 | 68 | 30,395,400–31,781,045 | 2020 2021 2022 Average | Glyma.20g081000–Glyma.20g085450 | 46 | 23.0 48.2 55.0 52.7 | 34.1 59.7 66.0 61.5 | 2.4 2.1 2.4 2.3 | [14,15,19,24,25,41] |
| Top 20 RILs with High Protein Content | Genotype of the Marker Linked to the QTLs | Protein Content (%) | Bottom 20 RILs with Low Protein Content | Genotype of the Marker Linked to the QTLs | Protein Content (%) | ||||
|---|---|---|---|---|---|---|---|---|---|
| qPSD15-1 | qPSD18-1 | qPSD20-1 | qPSD15-1 | qPSD18-1 | qPSD20-1 | ||||
| YS-196 | B | B | A | 52.3 | YS-229 | A | B | A | 39.4 |
| SI-400 | B | B | B | 52.1 | SI-465 | A | A | A | 39.6 |
| YS-068 | B | B | B | 52.1 | YS-111 | A | A | A | 39.8 |
| YS-080 | B | A | B | 52.0 | YS-209 | A | A | A | 39.9 |
| YS-005 | B | A | B | 52.0 | YS-036 | B | B | A | 40.0 |
| SI-317 | A | A | B | 51.9 | YS-037 | A | A | A | 40.3 |
| YS-190 | B | A | B | 51.8 | YS-117 | A | A | A | 40.4 |
| YS-109 | B | B | B | 51.7 | SI-337 | A | A | A | 40.6 |
| YS-199 | B | B | B | 51.6 | YS-043 | B | A | A | 40.8 |
| SI-428 | B | B | B | 51.5 | YS-118 | B | A | A | 40.9 |
| YS-173 | B | A | B | 51.4 | SI-361 | B | B | A | 41.1 |
| SI-423 | B | A | B | 51.4 | YS-063 | A | B | A | 41.1 |
| YS-205 | B | A | B | 51.3 | YS-227 | B | A | A | 41.2 |
| SI-445 | B | B | B | 51.3 | YS-048 | B | A | A | 41.2 |
| YS-090 | B | B | B | 51.3 | SI-473 | B | A | A | 41.2 |
| YS-008 | B | A | B | 51.2 | YS-015 | B | A | A | 41.4 |
| SI-348 | B | B | B | 51.2 | YS-235 | A | A | A | 41.6 |
| YS-202 | B | B | B | 51.2 | YS-100 | B | B | A | 41.6 |
| SI-326 | B | B | B | 51.1 | SI-502 | A | A | A | 41.6 |
| SI-345 | A | B | B | 51.1 | YS-045 | B | A | A | 41.6 |
| Population | Marker | Gene ID | Annotation Description | Biological Process | Reference | SNP Type |
|---|---|---|---|---|---|---|
| Y × S | qPSD15-1 | Glyma.15g102100 | Alpha/Beta hydrolase domain-containing protein | NA | Stop gain | |
| Glyma.15g102202 | Elongation factor Tu GTP binding domain | Translational elongation | Frameshift variant | |||
| Glyma.15g102252 | Elongation factor Tu C-terminal domain | Translational elongation | Frameshift variant | |||
| Glyma.15g102800 | Mediator of RNA polymerase II transcription subunit 33a | Phenylpropanoid metabolic process | Stop gain | |||
| Glyma.15g103100 | Mitochondrial editing factor 18 | RNA modification | Frameshift variant | |||
| Glyma.15g107200 | GPI-anchored protein | Biological process | Stop gain | |||
| Glyma.15g108000 | Starch/carbohydrate-binding module (family 53) | Starch biosynthetic process | Frameshift variant | |||
| Glyma.15g108900 | Glycosyl hydrolases family 17 | Carbohydrate metabolic process | Frameshift variant | |||
| Glyma.15g109800 | Peroxisomal membrane protein 2 | Biological process | Frameshift variant | |||
| Glyma.15g109900 | F-BOX protein with a domain protein | NA | Frameshift variant | |||
| qPSD18-1 | Glyma.18g193300 | Laccase | Iron ion transport | Frameshift variant | ||
| Glyma.18g193600 | Fructose-1,6-bisphosphatase, N-terminal domain | Sucrose metabolic process | [38] | Frameshift variant | ||
| Glyma.18g194700 | NA | NA | Stop gain | |||
| Glyma.18g194900 | NA | NA | Frameshift variant | |||
| Glyma.18g195000 | NA | Biological process | Frameshift variant | |||
| Glyma.18g195700 | Alpha-carboxyltransferase aCT-1 precursor | Fatty acid biosynthesis | [39,40] | Missense variant | ||
| Glyma.18g195900 | Carboxyl transferase domain | Fatty acid biosynthesis | [39,40] | Missense variant | ||
| Glyma.18g196000 | Carboxyl transferase domain | Fatty acid biosynthesis | [39,40] | Missense variant | ||
| Glyma.18g196600 | NA | NA | Stop gain | |||
| Glyma.18g197100 | NA | NA | Frameshift variant | |||
| qPSD20-1 | Glyma.20g085100 | POWR1 CCT motif family protein | Biological process | [14,24,25] | Missense variant | |
| Glyma.20g085700 | Unknown protein | NA | [15] | Stop gain | ||
| S × I | qPSD20-2 | Glyma.20g081500 | Lipase containing protein | Lipid catabolic process | Missense variant | |
| Glyma.20g082450 | Ammonium transporter 1 | Ammonium transport | [15] | Missense variant | ||
| Glyma.20g082700 | Sugar efflux transporter SWEET52 | Carbohydrate transport | [42,43] | Missense variant | ||
| Glyma.20g084000 | Small nuclear ribonucleoprotein F | Spliceosomal snRNP assembly | [15] | Missense variant | ||
| Glyma.20g084051 | Far1-relate | Regulation of transcription | [15] | Missense variant | ||
| Glyma.20G084500 | WD40 repeat protein | Innate immune response | [15] | Missense variant | ||
| Glyma.20g085100 | POWR1 CCT motif family protein | Biological process | [14,24,25] | Missense variant |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, H.R.; Seo, J.H.; Kang, B.K.; Kim, J.H.; Heo, S.V.; Choi, M.S.; Ko, J.Y.; Kim, C.S. QTLs and Candidate Genes for Seed Protein Content in Two Recombinant Inbred Line Populations of Soybean. Plants 2023, 12, 3589. https://doi.org/10.3390/plants12203589
Park HR, Seo JH, Kang BK, Kim JH, Heo SV, Choi MS, Ko JY, Kim CS. QTLs and Candidate Genes for Seed Protein Content in Two Recombinant Inbred Line Populations of Soybean. Plants. 2023; 12(20):3589. https://doi.org/10.3390/plants12203589
Chicago/Turabian StylePark, Hye Rang, Jeong Hyun Seo, Beom Kyu Kang, Jun Hoi Kim, Su Vin Heo, Man Soo Choi, Jee Yeon Ko, and Choon Song Kim. 2023. "QTLs and Candidate Genes for Seed Protein Content in Two Recombinant Inbred Line Populations of Soybean" Plants 12, no. 20: 3589. https://doi.org/10.3390/plants12203589






