Identification of AREB/ABF Gene Family Involved in the Response of ABA under Salt and Drought Stresses in Jute (Corchorus olitorius L.)
Abstract
1. Introduction
2. Results
2.1. Identification and Characteristics of AREB/ABFs in Jute
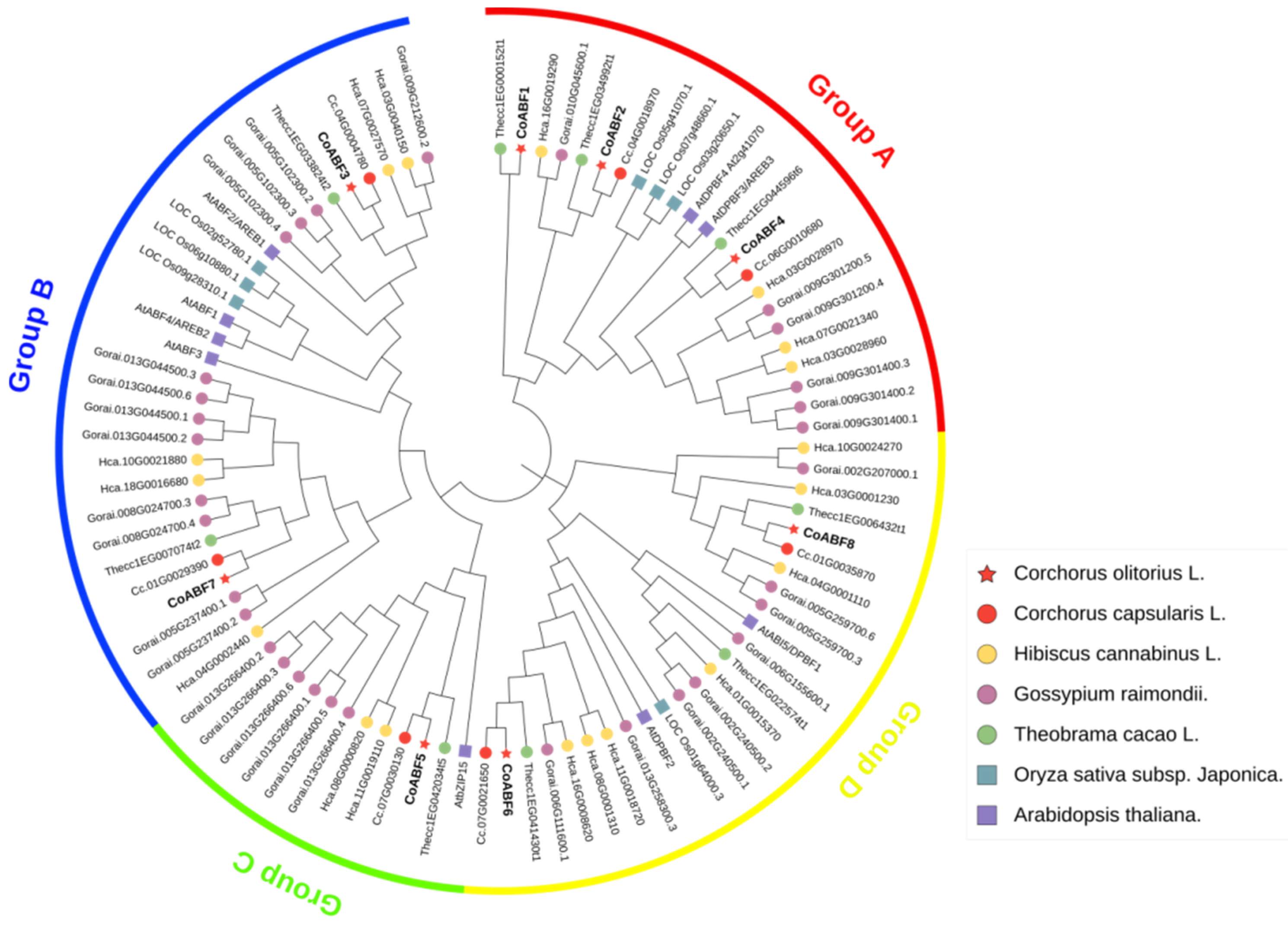
2.2. Phylogenetic Relationship and Sequence Analysis of AREB/ABFs in C. olitorius
2.3. Gene Structure and Conserved Motif Analyses of AREB/ABF Family Members in Jute
2.4. Chromosomal Distribution and Gene Duplication Analysis of CoABF Genes
2.5. cis-Element Analysis of AREB/ABF Gene Family in C. olitorius
2.6. Analysis of AREB/ABF Protein Network Interactions and Gene Ontology Annotation
2.7. Homology Modeling of CoABF Proteins
2.8. Expression Pattern of CoABF Genes under ABA Treatment Using Quantitative Real-Time PCR(qRT-PCR)
2.9. Expression Levels of CoABF3 and CoABF7 under PEG and PEG+ABA Treatments
2.10. Expression Levels of CoABF3 and CoABF7 under ST and ST+ABA Treatments
3. Discussion
4. Materials and Methods
4.1. Sequencing Analysis and Identification of AREB/ABF Gene Family in Jute
4.2. Protein Characterization and Chromosome Distribution Map of CoABF Genes
4.3. Gene Structure and Conserved Motifs Analysis of CoABF Genes
4.4. Gene Duplication and Ka (Nonsynonymous)/Ks (Synonymous) Ratio Analysis of CoABF Genes
4.5. Gene Ontology (GO) Enrichment, Protein Interaction Network, and cis-Elements Analysis of CoABF Genes
4.6. Homology Modeling of CoABF Proteins
4.7. Plant Materials and Growing Conditions
4.8. qRT-PCR Validation
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhang, L.; Ma, X.; Zhang, X.; Xu, Y.; Ibrahim, A.; Yao, J.; Huang, H.; Chen, S.; Liao, Z.; Zhang, Q. Reference genomes of the two cultivated jute species. Plant Biotechnol. J. 2021, 19, 2235–2248. [Google Scholar] [CrossRef]
- Chase, M.; Christenhusz, M.; Fay, M.; Byng, J.; Judd, W.; Soltis, D.; Mabberley, D.; Sennikov, A.; Soltis, P.; Stevens, P. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 2016, 181, 1–20. [Google Scholar]
- Christenhusz, M.J.; Byng, J.W. The number of known plants species in the world and its annual increase. Phytotaxa 2016, 261, 201–217. [Google Scholar] [CrossRef]
- Heidari, P.; Faraji, S.; Poczai, P. Magnesium transporter gene family: Genome-wide identification and characterization in Theobroma cacao, Corchorus capsularis, and Gossypium hirsutum of family Malvaceae. Agronomy 2021, 11, 1651. [Google Scholar] [CrossRef]
- Sarkar, D.; Mahato, A.K.; Satya, P.; Kundu, A.; Singh, S.; Jayaswal, P.K.; Singh, A.; Bahadur, K.; Pattnaik, S.; Singh, N.; et al. The draft genome of Corchorus olitorius cv. JRO-524 (Navin). Genom. Data 2017, 12, 151–154. [Google Scholar] [CrossRef]
- Islam, M.; Saito, J.; Emdad, E.; Ahmed, B.; Islam, M.; Halim, A.; Hossen, Q.; Hossain, M.; Ahmed, R.; Hossain, M. Comparative genomics of two jute species and insight into fiber biogenesis. Nat. Plants 2017, 3, 1–7. [Google Scholar] [CrossRef]
- Islam, M. Biochemistry, medicinal and food values of jute (Corchorus capsularis L. and C. olitorius L.) leaf: A review. Int. J. Enhanc. Res. Sci. Technol. Eng. 2013, 2, 135–144. [Google Scholar]
- Dhar, P.; Ojha, D.; Kar, C.; Mitra, J. Differential response of tossa jute (Corchorus olitorius) submitted to water deficit stress. Ind. Crops Prod. 2018, 112, 141–150. [Google Scholar] [CrossRef]
- Sheheli, S.; Roy, B. Constraints and opportunities of raw jute production: A household level analysis in Bangladesh. Progress. Agric. 2014, 25, 38–46. [Google Scholar] [CrossRef]
- Rahman, K.; Ahmed, N.; Raihan, M.; Hossain, R.; Nowroz, F.; Jannat, F.; Rahman, M.; Hasanuzzaman, M. Jute responses and tolerance to abiotic stress: Mechanisms and approaches. Plants 2021, 10, 1595. [Google Scholar] [CrossRef]
- Munns, R.; Termaat, A. Whole-plant responses to salinity. Funct. Plant Biol. 1986, 13, 143–160. [Google Scholar] [CrossRef]
- Zhu, J. Abiotic stress signaling and responses in plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, X.; Zhang, R.; Yuan, H.; Wang, M.; Yang, H.; Ma, H.; Liu, D.; Jiang, C.; Liang, Z. Root damage under alkaline stress is associated with reactive oxygen species accumulation in rice (Oryza sativa L.). Front. Plant Sci. 2017, 8, 1580. [Google Scholar] [CrossRef]
- Ali, Y.; Aslam, Z.; Ashraf, M.; Tahir, G. Effect of salinity on chlorophyll concentration, leaf area, yield and yield components of rice genotypes grown under saline environment. Int. J. Environ. Sci. Technol. 2004, 1, 221–225. [Google Scholar] [CrossRef]
- Sah, S.; Reddy, K.; Li, L. Abscisic acid and abiotic stress tolerance in crop plants. Front. Plant Sci. 2016, 7, 571. [Google Scholar] [CrossRef]
- Savvides, A.; Ali, S.; Tester, M.; Fotopoulos, V. Chemical priming of plants against multiple abiotic stresses: Mission possible? Trends Plant Sci. 2016, 21, 329–340. [Google Scholar] [CrossRef]
- Wei, L.; Lv, B.; Li, X.; Wang, M.; Ma, H.; Yang, H.; Yang, R.; Piao, Z.; Wang, Z.; Lou, J. Priming of rice (Oryza sativa L.) seedlings with abscisic acid enhances seedling survival, plant growth, and grain yield in saline-alkaline paddy fields. Field Crops Res. 2017, 203, 86–93. [Google Scholar] [CrossRef]
- Zhu, J. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu. Rev. Plant Biol. 2006, 57, 781–803. [Google Scholar] [CrossRef]
- de Zelicourt, A.; Colcombet, J.; Hirt, H. The role of MAPK modules and ABA during abiotic stress signaling. Trends Plant Sci. 2016, 21, 677–685. [Google Scholar] [CrossRef]
- Boneh, U.; Biton, I.; Schwartz, A.; Ben-Ari, G. Characterization of the ABA signal transduction pathway in Vitis vinifera. Plant Sci. 2012, 187, 89–96. [Google Scholar] [CrossRef]
- Boudsocq, M.; Droillard, M.; Barbier-Brygoo, H.; Laurière, C. Different phosphorylation mechanisms are involved in the activation of sucrose non-fermenting 1 related protein kinases 2 by osmotic stresses and abscisic acid. Plant Mol. Biol. 2007, 63, 491–503. [Google Scholar] [CrossRef]
- Yoshida, T.; Mogami, J.; Yamaguchi-Shinozaki, K. ABA-dependent and ABA-independent signaling in response to osmotic stress in plants. Curr. Opin. Plant Biol. 2014, 21, 133–139. [Google Scholar] [CrossRef] [PubMed]
- Van Ha, C.; Leyva-González, M.A.; Osakabe, Y.; Tran, U.T.; Nishiyama, R.; Watanabe, Y.; Tanaka, M.; Seki, M.; Yamaguchi, S.; Dong, N.V.; et al. Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc. Natl. Acad. Sci. USA 2014, 111, 851–856. [Google Scholar]
- Yoshida, T.; Fujita, Y.; Maruyama, K.; Mogami, J.; Todaka, D.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Four Arabidopsis AREB/ABF transcription factors function predominantly in gene expression downstream of SnRK2 kinases in abscisic acid signalling in response to osmotic stress. Plant Cell Environ. 2015, 38, 35–49. [Google Scholar] [CrossRef]
- Fujita, Y.; Yoshida, T.; Yamaguchi-Shinozaki, K. Pivotal role of the AREB/ABF-SnRK2 pathway in ABRE-mediated transcription in response to osmotic stress in plants. Physiol. Plant 2013, 147, 15–27. [Google Scholar] [CrossRef]
- Zandkarimi, H.; Ebadi, A.; Salami, S.A.; Alizade, H.; Baisakh, N. Analyzing the expression profile of AREB/ABF and DREB/CBF genes under drought and salinity stresses in grape (Vitis vinifera L.). PLoS ONE 2015, 10, e0134288. [Google Scholar] [CrossRef]
- Guiltinan, M.J.; Marcotte, W.R.; Quatrano, R.S. A plant leucine zipper protein that recognizes an abscisic acid response element. Science 1990, 250, 267–271. [Google Scholar] [CrossRef]
- Uno, Y.; Furihata, T.; Abe, H.; Yoshida, R.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc. Natl. Acad. Sci. USA 2000, 97, 11632–11637. [Google Scholar] [CrossRef]
- Venkatesh, J.; Yu, J.W.; Park, S.W. Genome-wide analysis and expression profiling of the Solanum tuberosum aquaporins. Plant Physiol. Biochem. 2013, 73, 392–404. [Google Scholar] [CrossRef]
- Busk, P.K.; Pagès, M. Regulation of abscisic acid-induced transcription. Plant Mol. Biol. 1998, 37, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Miyazono, K.; Koura, T.; Kubota, K.; Yoshida, T.; Fujita, Y.; Yamaguchi-Shinozaki, K.; Tanokura, M. Purification, crystallization and preliminary X-ray analysis of OsAREB8 from rice, a member of the AREB/ABF family of bZIP transcription factors, in complex with its cognate DNA. Acta Crystallogr. Sect. F. Struct. Biol. Cryst. Commun. 2012, 68, 491–494. [Google Scholar] [CrossRef]
- Rikiishi, K.; Matsuura, T.; Maekawa, M. TaABF1, ABA response element binding factor 1, is related to seed dormancy and ABA sensitivity in wheat (Triticum aestivum L.) seeds. J. Cereal Sci. 2010, 52, 236–238. [Google Scholar] [CrossRef]
- Liu, T.; Zhou, T.; Lian, M.; Liu, T.; Hou, J.; Ijaz, R.; Song, B. Genome-wide identification and characterization of the AREB/ABF/ABI5 subfamily members from Solanum tuberosum. Int. J. Mol. Sci. 2019, 20, 311. [Google Scholar] [CrossRef]
- Wang, W.; Qiu, X.; Yang, Y.; Kim, H.S.; Jia, X.; Yu, H.; Kwak, S. SweetpotatobZIP transcription factor IbABF4 confers tolerance to multiple abiotic stresses. Front. Plant Sci. 2019, 10, 630. [Google Scholar] [CrossRef]
- Kerr, T.C.; Abdel-Mageed, H.; Aleman, L.; Lee, J.; Payton, P.; Cryer, D.; Allen, R.D. Ectopic expression of two AREB/ABF orthologs increases drought tolerance in cotton (Gossypium hirsutum). Plant Cell Environ. 2018, 41, 898–907. [Google Scholar] [CrossRef] [PubMed]
- Ma, Q.J.; Sun, M.H.; Lu, J.; Liu, Y.J.; You, C.X.; Hao, Y.J. An apple CIPK protein kinase targets a novel residue of AREB transcription factor for ABA-dependent phosphorylation. Plant Cell Environ. 2017, 40, 2207–2219. [Google Scholar] [CrossRef]
- Li, D.; Mou, W.; Luo, Z.; Li, L.; Limwachiranon, J.; Mao, L.; Ying, T. Developmental and stress regulation on expression of a novel miRNA, Fan-miR73 and its target ABI5 in strawberry. Sci. Rep. 2016, 6, 28385. [Google Scholar] [CrossRef]
- Yong, X.; Zheng, T.; Zhuo, X.; Ahmad, S.; Li, L.; Li, P.; Yu, J.; Wang, J.; Cheng, T.; Zhang, Q. Genome-wide identification, characterization, and evolution of ABF/AREB subfamily in nine Rosaceae species and expression analysis in mei (Prunus mume). PeerJ 2021, 9, e10785. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Feng, Z.; Ding, Y.; Qi, Y.; Jiang, S.; Li, Z.; Wang, Y.; Qi, J.; Song, C.; Yang, S.; et al. RAF22, ABI1 and OST1 form a dynamic interactive network that optimizes plant growth and responses to drought stress in Arabidopsis. Mol. Plant 2022, 15, 1192–1210. [Google Scholar] [CrossRef]
- Pan, W.; Lin, B.; Yang, X.; Liu, L.; Xia, R.; Li, J.; Wu, Y.; Xie, Q. The UBC27–AIRP3 ubiquitination complex modulates ABA signaling by promoting the degradation of ABI1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2020, 117, 27694–27702. [Google Scholar] [CrossRef] [PubMed]
- Saleem, M.H.; Rehman, M.; Zahid, M.; Imran, M.; Xiang, W.; Liu, L. Morphological changes and antioxidative capacity of jute (Corchorus capsularis, Malvaceae) under different color light-emitting diodes. Rev. Bras. Bot. 2019, 42, 581–590. [Google Scholar] [CrossRef]
- Naik, M.R.; Barman, D.; Maruthi, R.T.; Babu, V.R.; Mandal, U.K.; Kundu, D.K. Assessment of salinity tolerance based upon morpho-physiological attributes in white jute (Corchorus capsularis L.). J. Environ. Biol. 2019, 40, 377–383. [Google Scholar] [CrossRef]
- Vishwakarma, K.; Upadhyay, N.; Kumar, N.; Yadav, G.; Singh, J.; Mishra, R.K.; Kumar, V.; Verma, R.; Upadhyay, R.G.; Pandey, M.; et al. Abscisic acid signaling and abiotic stress tolerance in plants: A review on current knowledge and future prospects. Front. Plant Sci. 2017, 8, 161. [Google Scholar] [CrossRef] [PubMed]
- Raghavendra, A.S.; Gonugunta, V.K.; Christmann, A.; Grill, E. ABA perception and signalling. Trends Plant Sci. 2010, 15, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Ma, H.; Liu, C.; Li, Z.; Ran, Q.; Xie, G.; Wang, B.; Fang, S.; Chu, J.; Zhang, J. ZmbZIP4 contributes to stress resistance in maize by regulating ABA synthesis and root development. Plant Physiol. 2018, 178, 753–770. [Google Scholar] [CrossRef]
- Mirzaei, K.; Bahramnejad, B.; Fatemi, S. Genome-wide identification and characterization of the bZIP gene family in potato (Solanum tuberosum). Plant Gene 2020, 24, 100257. [Google Scholar] [CrossRef]
- Bensmihen, S.; Rippa, S.; Lambert, G.; Jublot, D.; Pautot, V.; Granier, F.; Giraudat, J.; Parcy, F. The homologous ABI5 and EEL transcription factors function antagonistically to fine-tune gene expression during late embryogenesis. Plant Cell 2002, 14, 1391–1403. [Google Scholar] [CrossRef]
- De Storme, N.; Mason, A. Plant speciation through chromosome instability and ploidy change: Cellular mechanisms, molecular factors and evolutionary relevance. Curr. Plant Biol. 2014, 1, 10–33. [Google Scholar] [CrossRef]
- Soltis, D.E.; Soltis, P.S. Polyploidy: Recurrent formation and genome evolution. Trends Ecol. Evol. 1999, 14, 348–352. [Google Scholar] [CrossRef]
- Doyle, J.J.; Flagel, L.E.; Paterson, A.H.; Rapp, R.A.; Soltis, D.E.; Soltis, P.S.; Wendel, J.F. Evolutionary genetics of genome merger and doubling in plants. Annu. Rev. Genet. 2008, 42, 443–461. [Google Scholar] [CrossRef]
- Yoshida, T.; Fujita, Y.; Sayama, H.; Kidokoro, S.; Maruyama, K.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1, AREB2, and ABF3 are master transcription factors that cooperatively regulate ABRE-dependent ABA signaling involved in drought stress tolerance and require ABA for full activation. Plant J. 2010, 61, 672–685. [Google Scholar] [CrossRef]
- Ji, L.; Wang, J.; Ye, M.; Li, Y.; Guo, B.; Chen, Z.; Li, H.; An, X. Identification and characterization of the Populus AREB/ABF subfamily. J. Integr. Plant Biol. 2013, 55, 177–186. [Google Scholar] [CrossRef]
- Wei, K.; Chen, J.; Wang, Y.; Chen, Y.; Chen, S.; Lin, Y.; Pan, S.; Zhong, X.; Xie, D. Genome-wide analysis of bZIP-encoding genes in maize. DNA Res. 2012, 19, 463–476. [Google Scholar] [CrossRef]
- Choi, H.; Hong, J.; Ha, J.; Kang, J.; Kim, S.Y. ABFs, a family of ABA-responsive element binding factors. J. Biol. Chem. 2000, 275, 1723–1730. [Google Scholar] [CrossRef]
- Puranik, S.; Sahu, P.P.; Srivastava, P.S.; Prasad, M. NAC proteins: Regulation and role in stress tolerance. Trends Plant Sci. 2012, 17, 369–381. [Google Scholar] [CrossRef]
- Pickett, F.B.; Meeks-Wagner, D.R. Seeing double: Appreciating genetic redundancy. Plant Cell 1995, 7, 1347. [Google Scholar]
- Nowak, M.A.; Boerlijst, M.C.; Cooke, J.; Smith, J.M. Evolution of genetic redundancy. Nature 1997, 388, 167–171. [Google Scholar] [CrossRef]
- Lambie, E.J.; Kimble, J. Two homologous regulatory genes, lin-12 and glp-1, have overlapping functions. Development 1991, 112, 231–240. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef]
- Zhang, J. Positive selection, not negative selection, in the pseudogenization of rcsA in Yersinia pestis. Proc. Natl. Acad. Sci. USA 2008, 105, E69. [Google Scholar] [CrossRef] [PubMed]
- Foley, R.C.; Grossman, C.; Ellis, J.G.; Llewellyn, D.J.; Dennis, E.S.; Peacock, W.J.; Singh, K.B. Isolation of a maize bZIP protein subfamily: Candidates for the OCS-element transcription factor. Plant J. 1993, 3, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Wittkopp, P.J.; Kalay, G. Cis-regulatory elements: Molecular mechanisms and evolutionary processes underlying divergence. Nat. Rev. Genet. 2012, 13, 59–69. [Google Scholar] [CrossRef]
- Yamaguchi-Shinozaki, K.; Shinozaki, K. Organization of cis-acting regulatory elements in osmotic-and cold-stress-responsive promoters. Trends Plant Sci. 2005, 10, 88–94. [Google Scholar] [CrossRef]
- Feng, R.J.; Ren, M.Y.; Lu, L.F.; Peng, M.; Guan, X.; Zhou, D.B.; Zhang, M.Y.; Qi, D.F.; Li, K.; Tang, W.; et al. Involvement of abscisic acid-responsive element-binding factors in cassava (Manihot esculenta) dehydration stress response. Sci. Rep. 2019, 9, 12661. [Google Scholar] [CrossRef]
- Nakashima, K.; Ito, Y.; Yamaguchi-Shinozaki, K. Transcriptional regulatory networks in response to abiotic stresses in Arabidopsis and grasses. Plant Physiol. 2009, 149, 88–95. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Alhaithloul, H.A.S.; Abu-Elsaoud, A.M.; Soliman, M.M. Abiotic Stress Tolerance in Crop Plants: Role of Phytohormones. In Abiotic Stress in Plants; Intechopen: London, UK, 2020; p. 233. [Google Scholar]
- Bhuyan, M.; Hassan, K.M.; Lipi, N.; Uddin, M.; Islam, M.; Ferdous, M.; Antor, N.; Das, P. Screening of jute and kenaf varieties for salinity tolerance. Int. J. Adv. Geosci. 2018, 6, 214–218. [Google Scholar] [CrossRef]
- Endo, A.; Koshiba, T.; Kamiya, Y.; Nambara, E. The vascular system is a node of systemic stress responses: The competence of the cell to synthesize abscisic acid and its responsiveness to external cues. Plant Signal. Behav. 2008, 3, 1138–1140. [Google Scholar] [CrossRef]
- North, H.M.; Almeida, A.D.; Boutin, J.P.; Frey, A.; To, A.; Botran, L.; Sotta, B.; Marion-Poll, A. The Arabidopsis ABA-deficient mutant aba4 demonstrates that the major route for stress-induced ABA accumulation is via neoxanthin isomers. Plant J. 2007, 50, 810–824. [Google Scholar] [CrossRef]
- Osakabe, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.S.P. ABA control of plant macroelement membrane transport systems in response to water deficit and high salinity. New Phytol. 2014, 202, 35–49. [Google Scholar] [CrossRef]
- Yu, Y.; Weng, Q.; Zhou, B. Effects of exogenous ABA on contents of lycopene and endogenous hormone in tomato pericarp. Br. Biotechnol. J. 2016, 16, 1–5. [Google Scholar] [CrossRef]
- Tan, L.; Liu, Q.; Song, Y.; Zhou, G.; Luan, L.; Weng, Q.; He, C. Differential function of endogenous and exogenous abscisic acid during bacterial pattern-induced production of reactive oxygen species in Arabidopsis. Int. J. Mol. Sci. 2019, 20, 2544. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis acrosscomputing platforms. Mol. Biol. Evol. 2018, 35, 1547. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.; Jin, H.; Marler, B.; Guo, H. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 138, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Carbone, I.; Ramirez-Prado, J.H.; Jakobek, J.L.; Horn, B.W. Gene duplication, modularity, and adaptation in the evolution of the aflatoxin gene cluster. BMC Evol. Biol. 2007, 7, 111. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modeling, prediction, and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef] [PubMed]
- Geourjon, C.; Deleage, G. SOPMA: Significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Bioinformatics 1995, 11, 681–684. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Cheng, Z. Hoagland nutrient solution promotes the growth of cucumber seedlings under light-emitting diode light. Acta Agric. Scand. B Soil Plant Sci. 2015, 65, 74–82. [Google Scholar] [CrossRef]
- Wang, F.B.; Wan, C.Z.; Niu, H.F.; Qi, M.Y.; Gang, L.I.; Zhang, F.; Hu, L.B.; Ye, Y.X.; Wang, Z.X.; Pei, B.L.; et al. OsMas1, a novel maspardin protein gene, confers tolerance to salt and drought stresses by regulating ABA signaling in rice. J. Integr. Agric. 2023, 22, 341–359. [Google Scholar] [CrossRef]
- Zhang, Y.; Sun, T.; Liu, S.; Dong, L.; Liu, C.; Song, W.; Liu, J.; Gai, S. MYC cis-elements in PsMPT promoter is involved in chilling response of Paeonia suffruticosa. PLoS ONE 2016, 11, e0155780. [Google Scholar] [CrossRef]
- Livak, K.; Schmittgen, T. Analysis of relative gene expression data using real-time quantitive PCR and the2-ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]






| Gene | Gene Id | Theoretical pI | Molecular Mass (kDa) | Protein Length (aa) | Coding Sequence (CDS) Length | Sub-Cellular Location |
|---|---|---|---|---|---|---|
| CoABF1 | Co.02G0034250 | 9.86 | 27.67 | 253 | 762 | Nuclear |
| CoABF2 | Co.04G0023530 | 6.67 | 33.45 | 295 | 885 | Nuclear |
| CoABF3 | Co.04G0028660 | 9.36 | 49.05 | 457 | 1374 | Nuclear |
| CoABF4 | Co.06G0010990 | 8.8 | 35.80 | 321 | 966 | Nuclear |
| CoABF5 | Co.07G0004410 | 9.52 | 23.62 | 215 | 648 | Nuclear |
| CoABF6 | Co.07G0011920 | 9.1 | 41.88 | 378 | 1137 | Nuclear |
| CoABF7 | Co.v20117110 | 9.19 | 53.94 | 493 | 1482 | Nuclear |
| CoABF8 | Co.v20124440 | 8.48 | 63.11 | 575 | 1728 | Nuclear |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fiallos-Salguero, M.S.; Li, J.; Li, Y.; Xu, J.; Fang, P.; Wang, Y.; Zhang, L.; Tao, A. Identification of AREB/ABF Gene Family Involved in the Response of ABA under Salt and Drought Stresses in Jute (Corchorus olitorius L.). Plants 2023, 12, 1161. https://doi.org/10.3390/plants12051161
Fiallos-Salguero MS, Li J, Li Y, Xu J, Fang P, Wang Y, Zhang L, Tao A. Identification of AREB/ABF Gene Family Involved in the Response of ABA under Salt and Drought Stresses in Jute (Corchorus olitorius L.). Plants. 2023; 12(5):1161. https://doi.org/10.3390/plants12051161
Chicago/Turabian StyleFiallos-Salguero, Manuel Sebastian, Jing Li, Yunqing Li, Jiantang Xu, Pingping Fang, Yankun Wang, Liwu Zhang, and Aifen Tao. 2023. "Identification of AREB/ABF Gene Family Involved in the Response of ABA under Salt and Drought Stresses in Jute (Corchorus olitorius L.)" Plants 12, no. 5: 1161. https://doi.org/10.3390/plants12051161
APA StyleFiallos-Salguero, M. S., Li, J., Li, Y., Xu, J., Fang, P., Wang, Y., Zhang, L., & Tao, A. (2023). Identification of AREB/ABF Gene Family Involved in the Response of ABA under Salt and Drought Stresses in Jute (Corchorus olitorius L.). Plants, 12(5), 1161. https://doi.org/10.3390/plants12051161





