Fine Mapping of the Affecting Tillering and Plant Height Gene CHA-1 in Rice
Abstract
:1. Introduction
2. Results
2.1. Phenotypic Analysis of the CHA-1 Mutant
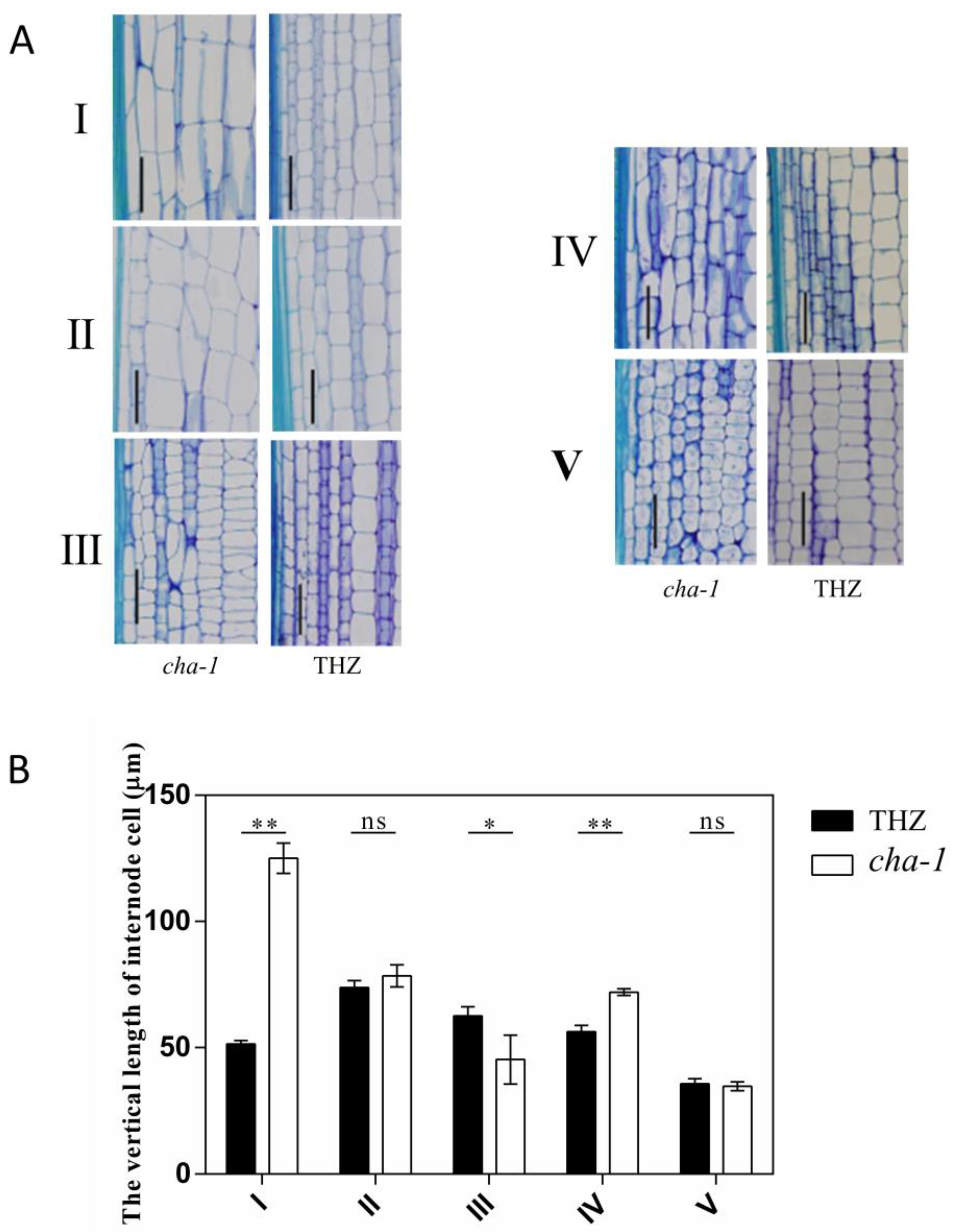
2.2. Microscopic Observation of the CHA-1 Mutant
2.3. Fine-Mapping of CHA-1
2.4. Analysis of Candidate Genes of CHA-1
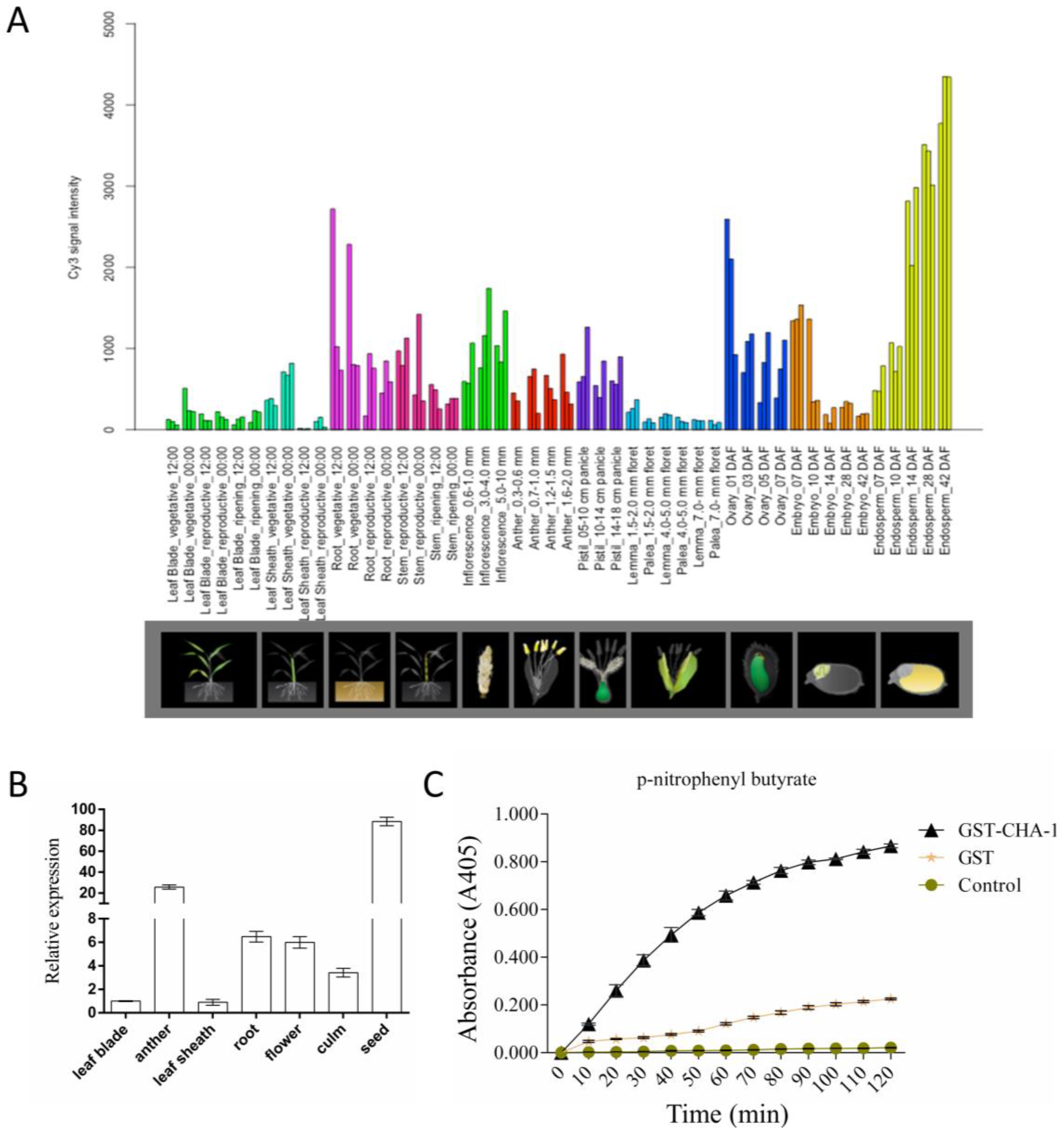
2.5. Expression and Evolutionary Analysis of CHA-1
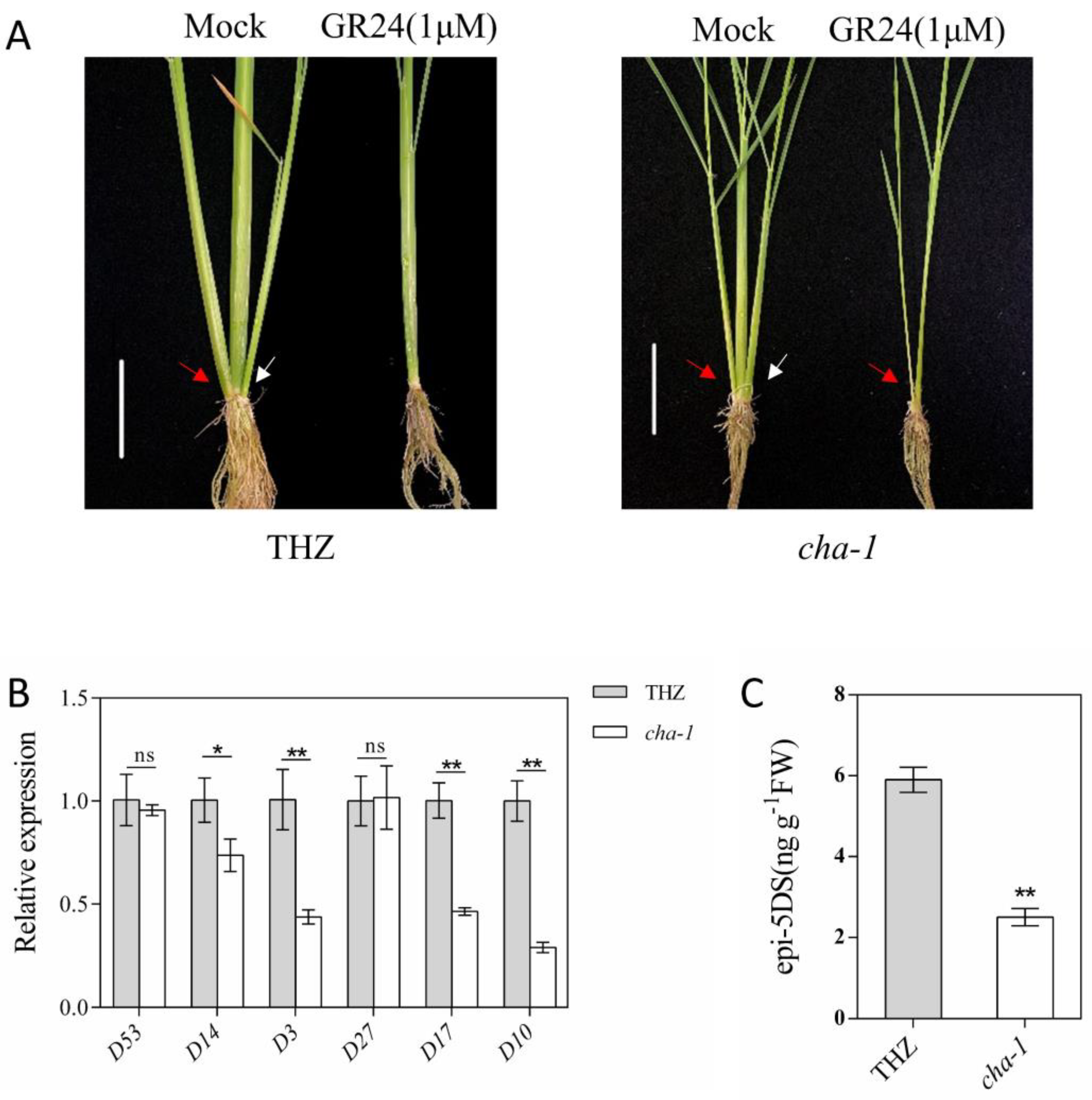
2.6. CHA-1 Affects the SL Biosynthesis
3. Materials and Methods
3.1. Plant Materials and Phenotypic Characterization
3.2. Histological Analysis
3.3. Fine Mapping of CHA-1
3.4. Phylogenetic Tree Construction and Analysis
3.5. Lipase Activity Assay
3.6. RNA Extraction and Quantitative Real-Time PCR (qPCR)
3.7. Vector Construction and Transformation
3.8. GR24 Treatment
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Khush, G.S. What it will take to feed 5.0 billion Rice consumers in 2030. Plant Mol. Biol. 2005, 59, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Godfray, H.; Beddington, J.R.; Crute, I.R.; Haddad, L.; Lawrence, D.; Muir, J.F.; Pretty, J.; Robinson, S.; Thomas, S.M.; Toulmin, C. Food Security: The challenge of feeding 9 billion people. Science 2010, 327, 812–818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bai, S.; Smith, S.M.; Li, J. Rice Plant Architecture: Molecular Basis and Application in Breeding. In Rice Genomics, Genetics and Breeding; Bai, S., Smith, S.M., Li, J., Sasaki, T., Ashikari, M., Eds.; Springer: Cham, Switzerland, 2018; pp. 129–154. [Google Scholar]
- Domagalska, M.A.; Leyser, O. Signal integration in the control of shoot branching. Nat. Rev. Mol. Cell Biol. 2011, 12, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Y.; Qian, Q.; Fu, Z.M.; Wang, Y.H.; Xiong, G.S.; Zeng, D.L.; Wang, X.Q.; Liu, X.F.; Teng, S.; Hiroshi, F.; et al. Control of tillering in rice. Nature 2003, 422, 618–621. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, W.; Ohmori, Y.; Ushijima, T.; Matsusaka, H.; Matsushita, T.; Kumamaru, T.; Kawano, S.; Hirano, H.Y. Axillary meristem formation in rice requires the WUSCHEL ortholog TILLERS ABSENT1. Plant Cell 2015, 27, 1173–1184. [Google Scholar] [CrossRef] [Green Version]
- Lu, Z.; Shao, G.N.; Xiong, J.S.; Jiao, Y.Q.; Wang, J.; Liu, G.F.; Meng, X.B.; Liang, Y.; Xiong, G.S.; Wang, Y.H.; et al. MONOCULM 3, an ortholog of WUSCHEL in rice, is required for tiller bud formation. J. Genet. Genom. 2015, 42, 71–78. [Google Scholar] [CrossRef]
- Shao, G.N.; Lu, Z.F.; Xiong, J.S.; Wang, B.; Jing, Y.H.; Meng, X.B.; Liu, G.F.; Ma, H.Y.; Liang, Y.; Chen, F.; et al. Tiller bud formation regulators MOC1 and MOC3 cooperatively promote tiller bud outgrowth by activating FON1 expression in rice. Mol. Plant 2019, 12, 1090–1102. [Google Scholar] [CrossRef] [Green Version]
- Umehara, M.; Hanada, A.; Yoshida, S.; Akiyama, K.; Arite, T.; Takeda-Kamiya, N.; Magome, H.; Kamiya, Y.; Shirasu, K.; Yoneyama, K.; et al. Inhibition of shoot branching by new terpenoid plant hormones. Nature 2008, 455, 195–200. [Google Scholar] [CrossRef]
- Gomez-Roldan, V.; Fermas, S.; Brewer, P.B.; Puech-Pages, V.; Dun, E.A.; Pillot, J.P.; Letisse, F.; Matusova, R.; Danoun, S.; Portais, J.C.; et al. Strigolactone inhibition of shoot branching. Nature 2008, 455, 189–194. [Google Scholar] [CrossRef]
- Mashiguchi, K.; Seto, Y.; Yamaguchi, S. Strigolactone biosynthesis, transport and perception. Plant J. 2021, 105, 335–350. [Google Scholar] [CrossRef]
- Jia, K.P.; Baz, L.; Al-Babili, S. From carotenoids to strigolactones. J. Exp. Bot. 2018, 69, 2189–2204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, H.; Wang, R.X.; Qian, Q.; Yan, M.X.; Meng, X.B.; Fu, Z.M.; Yan, C.Y.; Jiang, B.; Su, Z.; Li, J.Y.; et al. DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth. Plant Cell 2009, 21, 1512–1525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zou, J.H.; Zhang, S.Y.; Zhang, W.P.; Li, G.; Chen, Z.X.; Zhai, W.X.; Zhao, X.F.; Pan, X.B.; Xie, Q.; Zhu, L.H. The rice HIGH-TILLERING DWARF1 encoding an ortholog of arabidopsis MAX3 is required for negative regulation of the outgrowth of axillary buds. Plant J. 2006, 48, 687–696. [Google Scholar] [CrossRef] [PubMed]
- Alder, A.; Jamil, M.; Marzorati, M.; Bruno, M.; Vermathen, M.; Bigler, P.; Ghisla, S.; Bouwmeester, H.; Beyer, P.; Al-Babili, S. The path from beta-carotene to carlactone, a strigolactone-like plant hormone. Science 2012, 335, 1348–1351. [Google Scholar] [CrossRef] [Green Version]
- Jiang, L.; Liu, X.; Xiong, G.S.; Liu, H.H.; Chen, F.L.; Wang, L.; Meng, X.B.; Liu, G.F.; Yu, H.; Yuan, Y.D.; et al. DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 2013, 504, 401–405. [Google Scholar] [CrossRef] [Green Version]
- Zhou, F.; Lin, Q.B.; Zhu, L.H.; Ren, Y.L.; Zhou, K.N.; Shabek, N.; Wu, F.Q.; Mao, H.B.; Dong, W.; Gan, L.; et al. D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling. Nature 2013, 504, 406–410. [Google Scholar] [CrossRef] [Green Version]
- Arite, T.; Umehara, M.; Ishikawa, S.; Hanada, A.; Maekawa, M.; Yamaguchi, S.; Kyozuka, J. d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol. 2009, 50, 1416–1424. [Google Scholar] [CrossRef] [Green Version]
- Yao, R.F.; Ming, Z.H.; Yan, L.M.; Li, S.H.; Wang, F.; Ma, S.; Yu, C.T.; Yang, M.; Chen, L.; Chen, L.H.; et al. DWARF14 is a non-canonical hormone receptor for strigolactone. Nature 2016, 536, 469–473. [Google Scholar] [CrossRef]
- Song, X.G.; Lu, Z.F.; Yu, H.; Shao, G.N.; Xiong, J.S.; Meng, X.B.; Jing, Y.H.; Liu, G.F.; Xiong, G.S.; Duan, J.B.; et al. IPA1 functions as a downstream transcription factor repressed by D53 in strigolactone signaling in rice. Cell Res. 2017, 27, 1128–1141. [Google Scholar] [CrossRef] [Green Version]
- Li, W.; Ling, H.; Zhang, F.; Yao, H.Y.; Sun, X.F.; Tang, K.X. Analysis of Arabidopsis genes encoding putative class III lipases. J. Plant Biochem. Biot. 2012, 21, 261–267. [Google Scholar] [CrossRef]
- Zhao, H.; Ma, B.; Duan, K.X.; Li, X.K.; Lu, X.; Yin, C.C.; Tao, J.J.; Wei, W.; Zhang, W.K.; Xin, P.Y.; et al. The GDSL lipase MHZ11 modulates ethylene signaling in rice roots. Plant Cell 2020, 32, 1626–1643. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, H.G.; Xue, D.W.; Gao, Z.Y.; Yan, M.X.; Xu, W.Y.; Xing, Z.; Huang, D.N.; Qian, Q.; Xue, Y.B. A putative lipase gene EXTRA GLUME1 regulates both empty-glume fate and spikelet development in rice. Plant J. 2009, 57, 593–605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zeng, D.Y.; Cui, J.; Yin, Y.S.; Zhang, M.; Shan, S.; Liu, M.Y.; Cheng, D.Y.; Lu, W.H.; Sun, Y.Q. Proteomic analysis in different development stages on SP0 generation of rice seeds after space flight. Life Sci. Space Res. 2020, 26, 34–45. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Mishra, A.K.; Mohanta, Y.K.; Al-Harrasi, A. Space Breeding: The Next-Generation Crops. Front. Plant Sci. 2021, 12, 771985. [Google Scholar] [CrossRef]
- Xiao, W.M.; Yang, Q.Y.; Wang, H.; Guo, T.; Liu, Y.Z.; Zhu, X.Y.; Chen, Z.Q. Identification and fine mapping of a resistance gene to magnaporthe oryzae in a space-induced rice mutant. Mol. Breed. 2011, 28, 303–312. [Google Scholar] [CrossRef]
- Li, H.Z.; Wang, H.; Guo, T.; Liu, Y.Z.; Zhang, J.G.; Chen, Z.Q. Genetic analysis and gene mapping of a new dwarf rice CHA-1 induced by space mutation. J. South China Agric. Univ. 2009, 30, 6–9. (In Chinese) [Google Scholar]
- Liu, Y.Z.; Wang, H.; Chen, Z.Q.; Guo, T.; Zhang, J.G. Characters variation of special dwarf mutant CHA-1 induced by space environment in rice. J. South China Agric. Univ. 2005, 26, 1–4. (In Chinese) [Google Scholar]
- Sato, Y.; Antonio, B.A.; Namiki, N.; Takehisa, H.; Minami, H.; Kamatsuki, K.; Sugimoto, K.; Shimizu, Y.; Hirochika, H.; Nagamura, Y. RiceXPro: A platform for monitoring gene expression in japonica rice grown under natural field conditions. Nucleic Acids Res. 2011, 39, D1141–D1148. [Google Scholar] [CrossRef]
- Wang, Y.X.; Shang, L.G.; Yu, H.; Zeng, L.J.; Hu, J.; Ni, S.; Rao, Y.C.; Li, S.F.; Chu, J.F.; Meng, X.B.; et al. A strigolactone biosynthesis gene contributed to the green revolution in rice. Mol. Plant 2020, 13, 923–932. [Google Scholar] [CrossRef]
- Murray, M.G.; Thompson, W.F. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980, 8, 4321–4325. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [Green Version]
- Chen, C.J.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.H.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Long, T.; Wang, Y.F.; Tong, X.H.; Tang, J.; Li, J.L.; Wang, H.M.; Tang, L.Q.; Li, Z.Y.; Shu, Y.Z.; et al. RMS2 encoding a GDSL lipase mediates lipid homeostasis in anthers to determine rice male fertility. Plant Physiol. 2020, 182, 2047–2064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef] [Green Version]
- Zeng, D.Y.; Cui, J.; Yin, Y.S.; Zhang, M.; Shan, S.; Gao, X.; Zhang, Y.C.; Lu, W.H.; Sun, Y.Q. Effects of space flight on expression of key proteins in rice leaves. Rice Sci. 2020, 27, 423–433. [Google Scholar]
- Yu, S.; Luo, H.; Li, J.; Yu, X. Molecular variation and application from aerospace mutagenesis in upland rice huhan 3 and huhan 7. Rice Sci. 2013, 20, 249–258. [Google Scholar] [CrossRef]
- Liu, W.; Zhang, D.C.; Tang, M.F.; Li, D.Y.; Zhu, Y.X.; Zhu, L.H.; Chen, C.Y. THIS1 is a putative lipase that regulates tillering, plant height, and spikelet fertility in rice. J. Exp. Bot. 2013, 64, 4389–4402. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.C.; Yu, H.; Huang, J.; Wang, W.S.; Faruquee, M.; Zhang, F.; Zhao, X.Q.; Fu, B.Y.; Chen, K.; Zhang, H.L.; et al. Towards a deeper haplotype mining of complex traits in rice with RFGB v2.0. Plant Biotechnol. J. 2020, 18, 14–16. [Google Scholar] [CrossRef] [Green Version]
- Arite, T.; Iwata, H.; Ohshima, K.; Maekawa, M.; Nakajima, M.; Kojima, M.; Sakakibara, H.; Kyozuka, J. DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J. 2007, 51, 1019–1029. [Google Scholar] [CrossRef] [PubMed]
- Akoh, C.C.; Lee, G.C.; Liaw, Y.C.; Huang, T.H.; Shaw, J.F. GDSL family of serine esterases/lipases. Prog. Lipid Res. 2004, 43, 534–552. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.S.; Kim, B.K.; Kwon, S.J.; Jin, H.C.; Park, O.K. Arabidopsis GDSL lipase 2 plays a role in pathogen defense via negative regulation of auxin signaling. Biochem. Bioph. Res. Commun. 2009, 379, 1038–1042. [Google Scholar] [CrossRef] [PubMed]
- Hayward, A.; Stirnberg, P.; Beveridge, C.; Leyser, O. Interactions between Auxin and Strigolactone in Shoot Branching Control. Plant Physiol. 2009, 151, 400–412. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.H.; Wang, M.L.; Li, Y.Q.; Yan, W.; Chang, Z.Y.; Ni, H.L.; Chen, Z.F.; Wu, J.X.; Xu, C.J.; Deng, X.W.; et al. GDSL esterase/lipases OsGELP34 and OsGELP110/OsGELP115 are essential for rice pollen development. J. Integr. Plant Biol. 2020, 62, 1574–1593. [Google Scholar] [CrossRef] [PubMed] [Green Version]


| Types | Number of Plants | Ratio | χ2(3:1) χ20.05 = 3.84 |
|---|---|---|---|
| High plant height | 2471 | 3.04:1 | 0.12 |
| Low plant height | 812 |
| Gene ID | Annotation |
|---|---|
| LOC_Os01g54810 | Class III lipase |
| LOC_Os01g54850 | Cyclin-like F-box domain containing protein |
| LOC_Os01g54860 | Enoyl-CoA hydratase/isomerase |
| Marker | Primer Sequences (5′–3′) |
|---|---|
| RM8097-F | GCTGTCACTGACCGAGCGTAGG |
| RM8097-R | TCGAGAGATCCAATCCAGTTTGC |
| RM11668-F | AGTGTCTCTGGAGTTGGGAGTGG |
| RM11668-R | CTGTTCTTCCAGATGGGCTTCC |
| RM11669-F | AAACCGTTCCAGGGAGACTGACC |
| RM11669-R | TCGTCTGATCCATCCATCCATCC |
| DL1-F | AATGCGTGGGGTTTCATCTA |
| DL1-R | TAGAGCATGGATAGACGGGG |
| DL3-F | AGCTATGTGGTTAGGTCC |
| DL3-R | TAGATGAGGAAGCCTAGT |
| DL4-F | AGTGGCTAGTCACTTACA |
| DL4-R | GGAAGCCTAGTATGAAGC |
| DL5-F | ATTCCGGTGGCGTTTTCA |
| DL5-R | CCACCAAAATTGTAGGGAGT |
| DL6-F | CTCATTGTTGCCTATGAG |
| DL6-R | GCACGTACGTAGTGAGAT |
| DL7-F | ATCAGAAGCTCCTGACTCTT |
| DL7-R | GCCGGAGAGGTAGTCGT |
| DL8-F | TCTGAACTGAATGGTTCG |
| DL8-R | TACAGTGGAGTCCTGCTA |
| DL9-F | AGAGCACCCAAGAGTTAATC |
| DL9-R | TGGCTGATATTGGGTATG |
| DL10-F | GCCTTAGAGGAGGATCTTCT |
| DL10-R | TTCCTTGCATCTCACGTAGG |
| DL2-F | AAAAGCCCACTTTGCATGAG |
| DL2-R | AGGTGTAACGAGAAAGCGGA |
| RM11677-F | GTCTTGGAGCTGAGCACCTTGG |
| RM11677-R | GGCCCTCCGTGTAATCCTATTCC |
| qPCR-D53-F | CCAAGCAGTTTGAAGCGAC |
| qPCR-D53-R | CCGCAAGTTTATCAAAGTCAA |
| qPCR-D10-F | CGTGGCGATATCGATGGT |
| qPCR-D10-R | CGACCTCCTCGAACGTCTT |
| qPCR-D14-F | CGCCTTCGTCGGCCACTC |
| qPCR-D14-R | TCGAACCCGCCGTGGTAGTC |
| qPCR-D17-F | CTGTTCTTAGCGGGGTGTTC |
| qPCR-D17-R | GGCGTCGAACTCGTAGAAAG |
| qPCR-D3-F | TTAAGGTGGAGGGTGATTGC |
| qPCR-D3-R | AAGATCCATCTGCCCTGTTG |
| qPCR-D27-F | TCTGGGCTAAAGAATGAAAAG |
| qPCR-D27-R | AGAGCTTGGGTCACAATCTCG |
| Com-CHA-1-F | GAGCTCGGTACCCGGGGATCCTCTTGTAATTTTTGGGTATAAATAGACCC |
| Com-CHA-1-R | ACGACGGCCAGTGCCAAGCTTTGTAAATTTTTGTGGTCCAAAGTGA |
| GST-CHA-1-F | CCGCGTGGATCCCCGGAATTCATGGCGATCGACCTGGCG |
| GST-CHA-1-R | GTCACGATGCGGCCGCTCGAGCTACTTTGCAGGGGGTGACG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, T.; Xiao, W.; Huang, C.; Zhou, D.; Liu, Y.; Guo, T.; Chen, Z.; Wang, H. Fine Mapping of the Affecting Tillering and Plant Height Gene CHA-1 in Rice. Plants 2023, 12, 1507. https://doi.org/10.3390/plants12071507
Chen T, Xiao W, Huang C, Zhou D, Liu Y, Guo T, Chen Z, Wang H. Fine Mapping of the Affecting Tillering and Plant Height Gene CHA-1 in Rice. Plants. 2023; 12(7):1507. https://doi.org/10.3390/plants12071507
Chicago/Turabian StyleChen, Tengkui, Wuming Xiao, Cuihong Huang, Danhua Zhou, Yongzhu Liu, Tao Guo, Zhiqiang Chen, and Hui Wang. 2023. "Fine Mapping of the Affecting Tillering and Plant Height Gene CHA-1 in Rice" Plants 12, no. 7: 1507. https://doi.org/10.3390/plants12071507







