Transcriptome Analysis Reveals Novel Insights into the Hyperaccumulator Phytolacca acinosa Roxb. Responses to Cadmium Stress
Abstract
:1. Introduction
2. Results
2.1. Response of P. acinosa Seeds to Cd2+ Treatment
2.2. Response of P. acinosa Seedlings to Cd2+ Treatment
2.3. P. acinosa Has a High Cd2+ Accumulation Capacity
2.4. De Novo Transcriptome Sequencing and Assembly
2.5. Functional Annotation of Unigenes
2.6. Differentially Expressed Genes (DEGs) Response to Cd2+ Stress in P. acinosa
2.7. GO Enrichment Analysis of DEGs
2.8. KEGG Pathway Enrichment Analysis of DEGs
2.9. Transcription Factors Were Differentially Expressed under Cd2+ Stress
2.10. Cell Wall-Related Genes Were Differentially Expressed under Cd2+ Stress
2.11. Secondary Metabolism-Related Genes Were Differentially Expressed under Cd2+ Stress
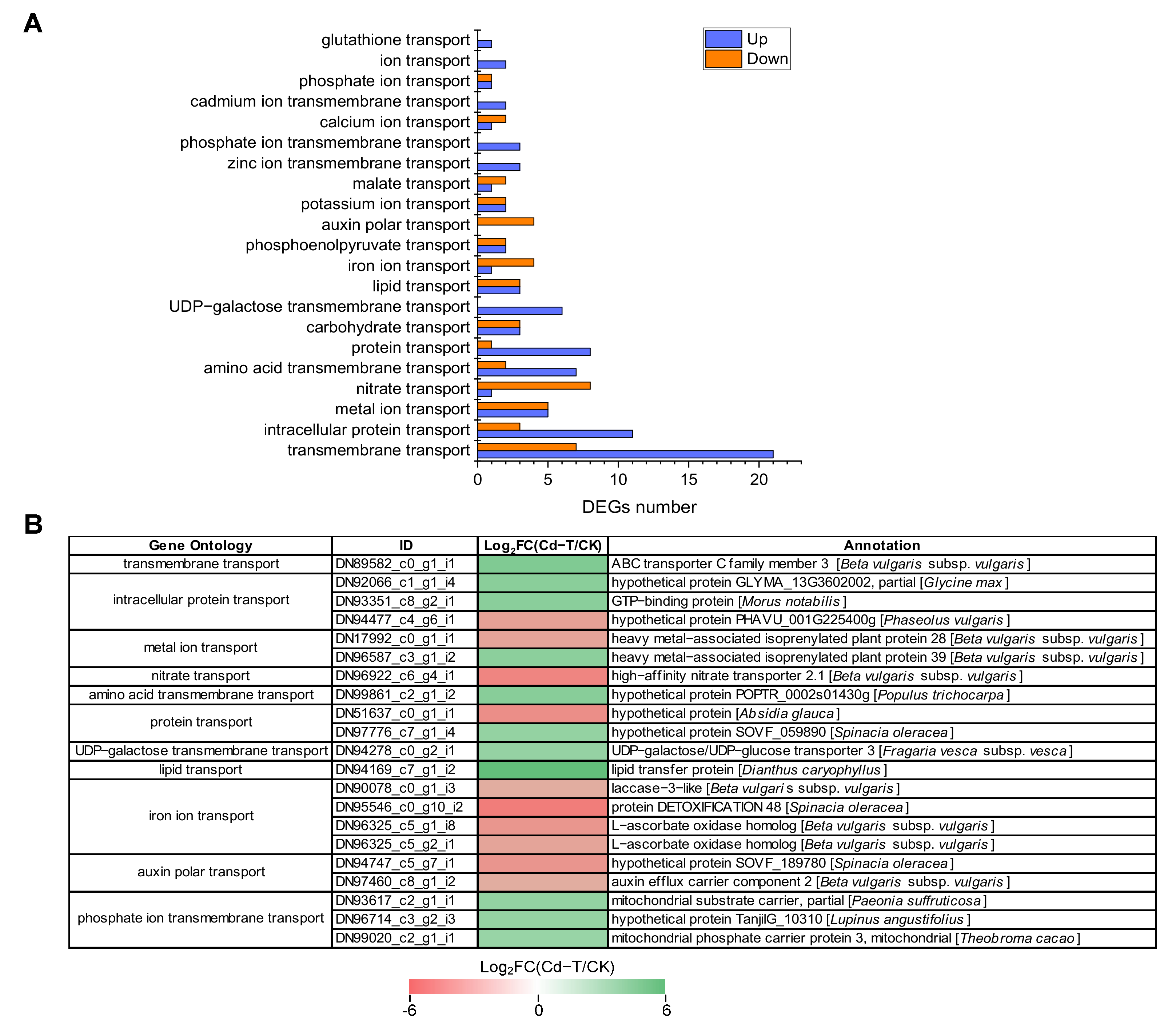
2.12. Membrane Proteins and Ion Transporters Were Differentially Expressed under Cd2+ Stress
2.13. Defense System-Related Genes Were Differentially Expressed under Cd2+ Stress
2.14. Validation of the DEGs by Real-Time PCR Analysis
3. Discussion
3.1. P. acinosa Can Be Used as a Candidate Plant for Cd Phytoremediation
3.2. Numerous TFs Were Involved in P. acinosa Response to Cd2+ Stress
3.3. Multiple Mechanisms Were Involved in Cd2+ Stress Tolerance in P. acinosa
3.4. ABC Transporters Were Involved in Regulating Cd2+ Stress
4. Materials and Methods
4.1. Materials and Cd2+ Treatment
4.2. Analysis of Seedling Traits
4.3. Cd2+ Content Analysis by ICP–MS
4.4. Measurement of MDA Content
4.5. Cd2+ Treatment for Transcriptome Sequencing
4.6. Library Preparation and Next Generation Sequencing
4.7. De Novo Transcriptome Assembly and Functional Annotation
4.8. Differentially Expression Analysis
4.9. Real-Time PCR Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, H.; Wang, Q.; Cui, Y.; Dong, Y.; Christie, P. Slow release chelate enhancement of lead phytoextraction by corn (Zea mays L.) from contaminated soil-a preliminary study. Sci. Total Environ. 2005, 339, 179–187. [Google Scholar] [CrossRef]
- Baryla, A.; Carrier, P.; Franck, F.; Coulomb, C.; Sahut, C.; Havaux, M. Leaf chlorosis in oilseed rape plants (Brassica napus) grown on cadmium-polluted soil: Causes and consequences for photosynthesis and growth. Planta 2001, 212, 696–709. [Google Scholar] [CrossRef] [PubMed]
- Asghar, M.; Habib, S.; Zaman, W.; Hussain, S.; Ali, H.; Saqib, S. Synthesis and characterization of microbial mediated cadmium oxide nanopartciles. Microsc. Res. Tech. 2020, 83, 1574–1584. [Google Scholar] [CrossRef]
- Kaleem, M.; Minhas, L.A.; Hashmi, M.Z.; Ali, M.A.; Mahmoud, R.M.; Saqib, S.; Nazish, M.; Zaman, W.; Mumtaz, A.S. Biosorption of Cadmium and Lead by Dry Biomass of Nostoc sp. MK-11 Kinetic and Isotherm Study. Molecules 2023, 28, 2292. [Google Scholar] [CrossRef]
- Zhou, J.; Cui, H.B.; Liang, J.N.; Xu, L.; Zhu, Z.Q.; Wang, X.X.; Cui, J.; Shang, J. Remediation technologies and current problems of heavy metal contaminated sites with ‘Demonstration project of soil remediation on the Periphery of Guixi Smelter’ as example. Soil 2015, 47, 283–288. [Google Scholar]
- Yuan, L.; Wang, H.; Wu, Y. Effects of cadmium and iron on the some physiological indicators in leaves of tobacco. Acta Sci. Circumstantiae 1990, 12, 147–154. [Google Scholar]
- Miner, G.; Gutierrez, R.; King, L. Soil Factors Affecting Plant Concentrations of Cadmium, Copper, and Zinc on Sludge-Amended Soils. J. Environ. Qual. 1997, 26, 989–994. [Google Scholar] [CrossRef]
- Choi, Y.E.; Harada, E.; Wada, M.; Tsuboi, H.; Morita, Y.; Kusano, T. Detoxification of cadmium in tobacco plants formation and active excretion of crystals containing cadmium and calcium through trichomes. Planta 2001, 213, 45–50. [Google Scholar] [CrossRef]
- Wagner, G.J. Accumulation of Cadmium in Crop Plants and Its Consequences to Human Health. Adv. Agron. 1993, 51, 173–212. [Google Scholar]
- Vido, K.; Spector, D.; Lagniel, G.; Lopez, S.; Toledano, M.B.; Labarre, J. A proteome analysis of the cadmium response in Saccharomyces cerevisiae. J. Biol. Chem. 2001, 276, 8469–8474. [Google Scholar] [CrossRef] [PubMed]
- Luo, Z.B.; He, J.; Polle, A.; Rennenberg, H. Heavy metal accumulation and signal transduction in herbaceous and woody plants: Paving the way for enhancing phytoremediation efficiency. Biotechnol. Adv. 2016, 34, 1131–1148. [Google Scholar] [CrossRef]
- Kashif, M.; Sang, Y.M.; Mo, S.M.; Rehman, S.u.; Khan, S.; Khan, M.R.; He, S.; Jiang, C.J. Deciphering the biodesulfurization pathway employing marine mangrove Bacillus aryabhattai strain NM1-A2 according to whole genome sequencing and transcriptome analyses. Genomics 2023, 115, 110635. [Google Scholar] [CrossRef] [PubMed]
- Nie, S.Q.; Mo, S.M.; Gao, T.W.; Yan, B.; Shen, P.H.; Kashif, M.; Zhang, Z.F.; Li, J.H.; Jiang, C.J. Coupling effects of nitrate reduction and sulfur oxidation in a subtropical marine mangrove ecosystem with Spartina alterniflora invasion. Sci. Total Environ. 2023, 862, 160930. [Google Scholar] [CrossRef]
- Wei, S.H.; Zhou, Q.X.; Mathews, S. A newly found cadmium accumulator-Taraxacum mongolicum. J. Hazard. Mater. 2008, 159, 544–547. [Google Scholar] [CrossRef] [PubMed]
- Saifullah; Meers, E.; Qadir, M.; Caritat, P.D.; Tack, F.M.G.; Laing, G.D.; Zia, M.H. EDTA-assisted Pb phytoextraction. Chemosphere 2009, 74, 1279–1291. [Google Scholar]
- Zhu, Y.; Bi, D.; Yuan, L.; Yin, X. Phytoremediation of Cadmium and Copper Contaminated Soils. In Phytoremediation and Biofortification: Two Sides of One Coin, 1st ed.; Yuan, L., Yin, X., Eds.; Springer: Amsterdam, The Netherlands, 2012; pp. 75–81. [Google Scholar]
- Knight, B.; Zhao, F.; McGrath, S.; Shen, Z. Zinc and cadmium uptake by the hyperaccumulator Thlaspi caerulescens in contaminated soils and its effects on the concentration and chemical speciation of metals in soil solution. Plant Soil 1997, 197, 71–78. [Google Scholar] [CrossRef]
- Lu, L.; Tian, S.; Zhang, M.; Zhang, J.; Yang, X.; Jiang, H. The role of Ca pathway in Cd uptake and translocation by the hyperaccumulator Sedum alfredii. J. Hazard. Mater. 2010, 183, 22–28. [Google Scholar] [CrossRef]
- Liu, W.; Shu, W.; Lan, C. Viola baoshanensis—A new Cd-hyperaccumulator. Chin. Sci. Bull. 2003, 48, 2046–2049. [Google Scholar]
- Wei, S.; Li, Y.; Zhou, Q.; Srivastava, M.; Chiu, S.; Zhan, J.; Wu, Z.; Sun, T. Effect of fertilizer amendments on phytoremediation of Cd-contaminated soil by a newly discovered hyperaccumulator Solanum nigrum L. J. Hazard. Mater. 2010, 176, 269–273. [Google Scholar] [CrossRef]
- Nie, F.H. Cd hyper-accumulator Phytolacca acinosa Roxb. and Cd-accumulative characteristics. Ecol. Environ. 2006, 15, 303–306. [Google Scholar]
- Liu, X.; Peng, K.; Wang, A.; Lian, C.; Shen, Z. Cadmium accumulation and distribution in populations of Phytolacca americana L. and the role of transpiration. Chemosphere 2010, 78, 1136–1141. [Google Scholar] [CrossRef]
- Clemens, S.; Aarts, M.G.M.; Thomine, S.; Verbruggen, N. Plant science: The key to preventing slow cadmium poisoning. Trends Plant Sci. 2013, 18, 92–99. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Li, H.; Ma, C.; Zhang, Y.; Polle, A.; Rennenberg, H.; Cheng, X.; Luo, Z.B. Overexpression of bacterial gamma-glutamylcysteine synthetase mediates changes in cadmium influx, allocation and detoxification in poplar. New Phytol. 2015, 205, 240–254. [Google Scholar] [CrossRef] [PubMed]
- Mills, R.F.; Francini, A.; Ferreira da Rocha, P.S.; Baccarini, P.J.; Aylett, M.; Krijger, G.C.; Williams, L.E. The plant P1B-type ATPase AtHMA4 transports Zn and Cd and plays a role in detoxification of transition metals supplied at elevated levels. FEBS Lett. 2005, 579, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Verret, F.; Gravot, A.; Auroy, P.; Leonhardt, N.; David, P.; Nussaume, L.; Vavasseur, A.; Richaud, P. Overexpression of AtHMA4 enhances root-to-shoot translocation of zinc and cadmium and plant metal tolerance. FEBS Lett. 2004, 576, 306–312. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Huang, Y.; Song, H.; Chen, F.; Cao, S. A MYB4-MAN3-Mannose-MNB1 signaling cascade regulates cadmium tolerance in Arabidopsis. PLoS Genet. 2021, 17, e1009636. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Zhao, H.X.; Wu, L.H.; Liu, A.N.; Zhao, F.J.; Xu, W.Z. Heavy metal ATPase 3 (HMA3) confers cadmium hypertolerance on the cadmium/zinc hyperaccumulator Sedum plumbizincicola. New Phytol. 2017, 215, 687–698. [Google Scholar] [CrossRef]
- Zhou, J.; Wan, H.; He, J.; Lyu, D.; Li, H. Integration of Cadmium Accumulation, Subcellular Distribution, and Physiological Responses to Understand Cadmium Tolerance in Apple Rootstocks. Front. Plant Sci. 2017, 8, 966. [Google Scholar] [CrossRef]
- Montero-Palmero, M.B.; Martin-Barranco, A.; Escobar, C.; Hernandez, L.E. Early transcriptional responses to mercury: A role for ethylene in mercury-induced stress. New Phytol. 2014, 201, 116–130. [Google Scholar] [CrossRef]
- Devaiah, B.N.; Karthikeyan, A.S.; Raghothama, K.G. WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol. 2007, 143, 1789–1801. [Google Scholar] [CrossRef]
- Zhu, N.; Cheng, S.; Liu, X.; Du, H.; Dai, M.; Zhou, D.X.; Yang, W.; Zhao, Y. The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice. Plant Sci. Int. J. Exp. Plant Biol. 2015, 236, 146–156. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, M.; Hao, Y.; Kapoor, A.; Dong, C.H.; Fujii, H.; Zheng, X.; Zhu, J.K. A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance. J. Biol. Chem. 2006, 281, 37636–37645. [Google Scholar] [CrossRef] [PubMed]
- Devaiah, B.N.; Madhuvanthi, R.; Karthikeyan, A.S.; Raghothama, K.G. Phosphate starvation responses and gibberellic acid biosynthesis are regulated by the MYB62 transcription factor in Arabidopsis. Mol. Plant 2009, 2, 43–58. [Google Scholar] [CrossRef] [PubMed]
- Mengiste, T.; Chen, X.; Salmeron, J.; Dietrich, R. The BOTRYTIS SUSCEPTIBLE1 gene encodes an R2R3MYB transcription factor protein that is required for biotic and abiotic stress responses in Arabidopsis. Plant Cell 2003, 15, 2551–2565. [Google Scholar] [CrossRef] [PubMed]
- Zanetti, M.E.; Ripodas, C.; Niebel, A. Plant NF-Y transcription factors: Key players in plant-microbe interactions, root development and adaptation to stress. Biochim. Biophys. Acta. Gene Regul. Mech. 2017, 1860, 645–654. [Google Scholar] [CrossRef] [PubMed]
- Leyva-Gonzalez, M.A.; Ibarra-Laclette, E.; Cruz-Ramirez, A.; Herrera-Estrella, L. Functional and transcriptome analysis reveals an acclimatization strategy for abiotic stress tolerance mediated by Arabidopsis NF-YA family members. PLoS ONE 2012, 7, e48138. [Google Scholar] [CrossRef]
- Xu, Z.; Dong, M.; Peng, X.; Ku, W.; Zhao, Y.; Yang, G. New insight into the molecular basis of cadmium stress responses of wild paper mulberry plant by transcriptome analysis. Ecotoxicol. Environ. Saf. 2019, 171, 301–312. [Google Scholar] [CrossRef]
- Ogo, Y.; Itai, R.N.; Nakanishi, H.; Kobayashi, T.; Takahashi, M.; Mori, S.; Nishizawa, N.K. The rice bHLH protein OsIRO2 is an essential regulator of the genes involved in Fe uptake under Fe-deficient conditions. Plant J. Cell Mol. Biol. 2007, 51, 366–377. [Google Scholar] [CrossRef]
- Chen, H.; Li, Y.; Ma, X.; Guo, L.; He, Y.; Ren, Z.; Kuang, Z.; Zhang, X.; Zhang, Z. Analysis of potential strategies for cadmium stress tolerance revealed by transcriptome analysis of upland cotton. Sci. Rep. 2019, 9, 86. [Google Scholar] [CrossRef]
- Cosio, C.; Martinoia, E.; Keller, C. Hyperaccumulation of cadmium and zinc in Thlaspi caerulescens and Arabidopsis halleri at the leaf cellular level. Plant Physiol. 2004, 134, 716–725. [Google Scholar] [CrossRef]
- Dimitrov, D.; Yoncheva, T.; Haygarov, V. Evaluation the Risk of Toxic Compounds Formation in Grape and Fruit Brandies. J. Microbiol. Biotechnol. Food Sci. 2016, 6, 681–684. [Google Scholar] [CrossRef]
- Cataldo, S.; Gianguzza, A.; Pettignano, A.; Piazzese, D.; Sammartano, S. Complex Formation of Copper(II) and Cadmium(II) with Pectin and Polygalacturonic Acid in Aqueous Solution. An ISE-H+ and ISE-Me2+ Electrochemical Study. Int. J. Electrochem. Sci. 2012, 7, 6722–6737. [Google Scholar] [CrossRef]
- Eghbaliferiz, S.; Iranshahi, M. Prooxidant Activity of Polyphenols, Flavonoids, Anthocyanins and Carotenoids: Updated Review of Mechanisms and Catalyzing Metals. Phytother. Res. 2016, 30, 1379–1391. [Google Scholar] [CrossRef]
- Zhao, H.; Wei, Y.; Wang, J.; Chai, T. Isolation and expression analysis of cadmium-induced genes from Cd/Mn hyperaccumulator Phytolacca americana in response to high Cd exposure. Plant Biol. 2019, 21, 15–24. [Google Scholar] [CrossRef]
- Nadeem, S.; Muhammad, I.; Rashid, S.M.; Wajid, I.; Asif, K.M.; Amar, M.; Abdur, R.; Saddam, H. Phytoremediation strategies for soils contaminated with heavy metals: Modifications and future perspectives. Chemosphere 2017, 171, 710–721. [Google Scholar]
- Liang, X.A.; Yan, W.A.; Wei, L.A.; Jin, W.B.; Xz, C.; Kz, B.; Ry, A.; Rw, A.; Yang, X.A.; Wei, Z.A. De novo sequencing of root transcriptome reveals complex cadmium-responsive regulatory networks in radish (Raphanus sativus L.). Plant Sci. Int. J. Exp. Plant Biol. 2015, 236, 313–323. [Google Scholar]
- Dubos, C.; Stracke, R.; Grotewold, E.; Weisshaar, B.; Martin, C.; Lepiniec, L. MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010, 15, 573–581. [Google Scholar] [CrossRef] [PubMed]
- Van de Mortel, J.E.; Schat, H.; Moerland, P.D.; Ver Loren van Themaat, E.; Van der Ent, S.; Blankestijn, H.; Ghandilyan, A.; Tsiatsiani, S.; Aarts, M.G. Expression differences for genes involved in lignin, glutathione and sulphate metabolism in response to cadmium in Arabidopsis thaliana and the related Zn/Cd-hyperaccumulator Thlaspi caerulescens. Plant Cell Environ. 2008, 31, 301–324. [Google Scholar] [CrossRef]
- Wang, F.Z.; Chen, M.X.; Yu, L.J.; Xie, L.J.; Yuan, L.B.; Qi, H.; Xiao, M.; Guo, W.; Chen, Z.; Yi, K.; et al. OsARM1, an R2R3 MYB Transcription Factor, Is Involved in Regulation of the Response to Arsenic Stress in Rice. Front. Plant Sci. 2017, 8, 1868. [Google Scholar] [CrossRef]
- Li, S.; Wang, W.; Gao, J.; Yin, K.; Wang, R.; Wang, C.; Petersen, M.; Mundy, J.; Qiu, J.L. MYB75 Phosphorylation by MPK4 Is Required for Light-Induced Anthocyanin Accumulation in Arabidopsis. Plant Cell 2016, 28, 2866–2883. [Google Scholar] [CrossRef]
- Farinati, S.; DalCorso, G.; Varotto, S.; Furini, A. The Brassica juncea BjCdR15, an ortholog of Arabidopsis TGA3, is a regulator of cadmium uptake, transport and accumulation in shoots and confers cadmium tolerance in transgenic plants. New Phytol. 2010, 185, 964–978. [Google Scholar] [CrossRef]
- Wang, Y.; Gao, C.; Liang, Y.; Wang, C.; Yang, C.; Liu, G. A novel bZIP gene from Tamarix hispida mediates physiological responses to salt stress in tobacco plants. J. Plant Physiol. 2010, 167, 222–230. [Google Scholar] [CrossRef]
- Chmielowska-Bak, J.; Lefevre, I.; Lutts, S.; Deckert, J. Short term signaling responses in roots of young soybean seedlings exposed to cadmium stress. J. Plant Physiol. 2013, 170, 1585–1594. [Google Scholar] [CrossRef]
- Ogawa, I.; Nakanishi, H.; Mori, S.; Nishizawa, N.K. Time course analysis of gene regulation under cadmium stress in rice. Plant Soil 2009, 325, 97–108. [Google Scholar] [CrossRef]
- Yamaguchi, H.; Fukuoka, H.; Arao, T.; Ohyama, A.; Nunome, T.; Miyatake, K.; Negoro, S. Gene expression analysis in cadmium-stressed roots of a low cadmium-accumulating solanaceous plant, Solanum torvum. J. Exp. Bot. 2010, 61, 423–437. [Google Scholar] [CrossRef] [PubMed]
- Sundaramoorthy, P.; Chidambaram, A.; Ganesh, K.S.; Unnikannan, P.; Baskaran, L. Chromium stress in paddy: (i) Nutrient status of paddy under chromium stress; (ii) Phytoremediation of chromium by aquatic and terrestrial weeds. Comptes Rendus Biol. 2010, 333, 597–607. [Google Scholar] [CrossRef]
- Zhu, H.; Ai, H.; Cao, L.; Sui, R.; Ye, H.; Du, D.; Sun, J.; Yao, J.; Chen, K.; Chen, L. Transcriptome analysis providing novel insights for Cd-resistant tall fescue responses to Cd stress. Ecotoxicol. Environ. Saf. 2018, 160, 349–356. [Google Scholar] [CrossRef] [PubMed]
- Leng, Y. Molecular Mechanism of Mung Bean Seedlings in Response to Cadmium Stress; Lanzhou Jiaotong University: Lanzhou, China, 2021. [Google Scholar]
- Nocito, F.F.; Lancilli, C.; Crema, B.; Fourcroy, P.; Davidian, J.C.; Sacchi, G.A. Heavy metal stress and sulfate uptake in maize roots. Plant Physiol. 2006, 141, 1138–1148. [Google Scholar] [CrossRef]
- Babula, P.; Klejdus, B.; Kovacik, J.; Hedbavny, J.; Hlavna, M. Lanthanum rather than cadmium induces oxidative stress and metabolite changes in Hypericum perforatum. J. Hazard. Mater. 2015, 286, 334–342. [Google Scholar] [CrossRef] [PubMed]
- Kovacik, J.; Klejdus, B. Dynamics of phenolic acids and lignin accumulation in metal-treated Matricaria chamomilla roots. Plant Cell Rep. 2008, 27, 605–615. [Google Scholar] [CrossRef]
- Gutsch, A.; Keunen, E.; Guerriero, G.; Renaut, J.; Cuypers, A.; Hausman, J.F.; Sergeant, K. Long-term cadmium exposure influences the abundance of proteins that impact the cell wall structure in Medicago sativa stems. Plant Biol. 2018, 20, 1023–1035. [Google Scholar] [CrossRef]
- Greger, M.; Kabir, A.H.; Landberg, T.; Maity, P.J.; Lindberg, S. Silicate reduces cadmium uptake into cells of wheat. Environ. Pollut. 2016, 211, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Li, Q.; Liu, Z. Genome-wide identification, characterization and phylogenetic analysis of 50 catfish ATP-binding cassette (ABC) transporter genes. PLoS ONE 2013, 8, e63895. [Google Scholar] [CrossRef]
- Park, J.; Song, W.Y.; Ko, D.; Eom, Y.; Hansen, T.H.; Schiller, M.; Lee, T.G.; Martinoia, E.; Lee, Y. The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J. Cell Mol. Biol. 2012, 69, 278–288. [Google Scholar] [CrossRef] [PubMed]
- Brunetti, P.; Zanella, L.; De Paolis, A.; Di Litta, D.; Cecchetti, V.; Falasca, G.; Barbieri, M.; Altamura, M.M.; Costantino, P.; Cardarelli, M. Cadmium-inducible expression of the ABC-type transporter AtABCC3 increases phytochelatin-mediated cadmium tolerance in Arabidopsis. J. Exp. Bot. 2015, 66, 3815–3829. [Google Scholar] [CrossRef] [PubMed]
- Teschner, J.; Lachmann, N.; Schulze, J.; Geisler, M.; Selbach, K.; Santamaria-Araujo, J.; Balk, J.; Mendel, R.R.; Bittner, F. A novel role for Arabidopsis mitochondrial ABC transporter ATM3 in molybdenum cofactor biosynthesis. Plant Cell 2010, 22, 468–480. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.Y.; Bovet, L.; Maeshima, M.; Martinoia, E.; Lee, Y. The ABC transporter AtPDR8 is a cadmium extrusion pump conferring heavy metal resistance. Plant J. Cell Mol. Biol. 2007, 50, 207–218. [Google Scholar] [CrossRef]
- Fu, S.; Lu, Y.; Zhang, X.; Yang, G.; Chao, D.; Wang, Z.; Shi, M.; Chen, J.; Chao, D.Y.; Li, R.; et al. The ABC transporter ABCG36 is required for cadmium tolerance in rice. J. Exp. Bot. 2019, 70, 5909–5918. [Google Scholar] [CrossRef]
- Jan, B.; Bhat, M.A.; Bhat, T.A.; Nazir, A.; Dar, K.A.; Kamran, M.; Soufan, W.; Abdelhamid, M.T.; Sabagh, A.E. Measuring the Influence of Seedling Age and Nutrient Sources on the Performance of Sweet Corn (Zea mays L.) under Temperate Climatic Conditions. ACS Omega 2023, 8, 18981–18989. [Google Scholar] [CrossRef]
- Sui, M.F.; Wei, X.H.; Xin, X.Q.; Yue, K.; Zhao, Y.; Han, T.; Ma, W.J.; Luo, Q. Exogenous cGMP regulates seed germination of ryegrass under salt stress. Acta Ecol. Sin. 2018, 38, 6171–6179. [Google Scholar]
- Xie, Q.; Ma, L.Y.; Tan, P.; Deng, W.T.; Huang, C.; Liu, D.M.; Lin, W.H.; Su, Y. Multiple High-Affinity K+ Transporters and ABC Transporters Involved in K+ Uptake/Transport in the Potassium-Hyperaccumulator Plant Phytolacca acinosa Roxb. Plants 2020, 9, 470. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zhi, J.; Zhang, H.; Li, J.; Zhao, Q.; Xu, J. Transcriptome analysis of Phytolacca americana L. in response to cadmium stress. PLoS ONE 2017, 12, e0184681. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Xu, J.; Huang, K.; Su, Y.; Tong, J.; Huang, Z.; Huang, C.; Wei, M.; Lin, W.; Xiao, L. Dynamic formation and transcriptional regulation mediated by phytohormones during chalkiness formation in rice. BMC Plant Biol. 2021, 21, 308. [Google Scholar] [CrossRef] [PubMed]
- Patel, R.K.; Mukesh, J.; Liu, Z. NGS QC Toolkit: A toolkit for quality control of next generation sequencing data. PLoS ONE 2012, 7, e30619. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Amit, I. Trinity: Reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat. Biotechnol. 2013, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Simon, A.; Theodor, P.P.; Wolfgang, H. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with topHat and cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, 182–185. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]











| Sample | Raw Reads Number | Raw Bases Number | Clean Reads Number | Clean Bases Number | Valid Bases (%) | Q20 (%) | Q30 (%) | GC (%) |
|---|---|---|---|---|---|---|---|---|
| CK1 | 34,397,898 | 5,159,684,700 | 34,004,184 | 5,075,043,936 | 98.35% | 98.57% | 95.56% | 44.00% |
| CK2 | 42,980,760 | 6,447,114,000 | 42,566,702 | 6,353,506,726 | 98.54% | 98.65% | 95.74% | 44.00% |
| CK3 | 43,298,432 | 6,494,764,800 | 42,821,964 | 6,391,566,154 | 98.41% | 98.63% | 95.67% | 44.00% |
| Cd-T1 | 34,587,638 | 5,188,145,700 | 34,262,482 | 5,123,597,064 | 98.75% | 98.60% | 95.58% | 44.00% |
| Cd-T2 | 38,393,362 | 4,759,004,300 | 38,182,134 | 4,682,860,231 | 98.40% | 98.57% | 95.55% | 44.00% |
| Cd-T3 | 46,707,780 | 7,006,167,000 | 46,290,484 | 6,919,976,366 | 98.76% | 98.65% | 95.73% | 44.00% |
| Type | Assembled Transcripts | Predicted Unigenes |
|---|---|---|
| Total sequence number | 255,468 | 124,408 |
| Total sequence base | 299,181,020 | 113,753,609 |
| Largest length (bp) | 17,759 | 17,759 |
| Smallest length (bp) | 301 | 301 |
| Average length (bp) | 1171.11 | 914.36 |
| N50 length (bp) | 1781 | 1282 |
| ID | Log2FC (Cd-T/CK) | Regulate | Annotation |
|---|---|---|---|
| cell wall biogenesis | |||
| DN88120_c0_g1_i1 | −2.01 | Down | xyloglucan endotransglucosylase/hydrolase 2-like |
| DN92891_c3_g4_i1 | 1.66 | Up | xyloglucan endotransglucosylase/hydrolase protein 23 |
| DN92891_c3_g6_i1 | −2.72 | Down | xyloglucan endotransglucosylase/hydrolase protein22-like isoform X1 |
| DN95917_c1_g1_i1 | −2.41 | Down | xyloglucan endotransglucosylase/hydrolase protein 23 |
| DN96589_c1_g1_i2 | 2.41 | Up | xyloglucan endotransglucosylase/hydrolase protein 6 |
| DN96589_c1_g5_i1 | −1.62 | Down | xyloglucan endotransglucosylase/hydrolase protein 6 |
| DN97081_c2_g1_i11 | −1.21 | Down | beta-arabinofuranosyltransferase RAY1 isoform X3 |
| DN97099_c2_g1_i1 | −1.47 | Down | xyloglucan endotransglucosylase/hydrolase protein 7 |
| DN98746_c2_g1_i4 | −1.46 | Down | xyloglucan endotransglucosylase/hydrolase 3 |
| DN99640_c5_g1_i1 | −1.05 | Down | hypothetical protein SOVF_125280 |
| DN88120_c0_g2_i1 | −1.58 | Down | hypothetical protein SOVF_073730 |
| cell wall organization | |||
| DN100083_c6_g2_i4 | −1.11 | Down | hypothetical protein SOVF_144400 |
| DN108829_c0_g1_i1 | 1.62 | Up | beta-1,3-galactosyltransferase 6 |
| DN89571_c0_g1_i2 | 2.11 | Up | rhamnogalacturonate lyase B |
| DN91042_c0_g1_i1 | 1.52 | Up | polygalacturonase-like |
| DN91881_c0_g1_i1 | −1.66 | Down | hypothetical protein SOVF_175910 |
| DN95973_c4_g1_i3 | −1.26 | Down | glucan endo-1,3-beta-glucosidase 13 |
| DN95989_c1_g3_i3 | 1.28 | Up | NAC transcription factor |
| DN96070_c1_g1_i10 | 1.97 | Up | hypothetical protein BVRB_5g098360 isoform B |
| DN96441_c2_g2_i2 | −1.32 | Down | omega-hydroxypalmitate O-feruloyl transferase |
| DN96517_c3_g4_i1 | −6.13 | Down | Putative UDP-glucuronate:xylan alpha-glucuronosyltransferase 4 |
| DN96603_c7_g3_i1 | −2.65 | Down | hypothetical protein BVRB_9g215820 |
| DN96627_c2_g1_i1 | −2.86 | Down | hypothetical protein BVRB_9g215820 |
| DN97041_c1_g1_i1 | −1.09 | Down | leucine-rich repeat extensin-like protein 4 |
| DN98038_c4_g1_i5 | −1.03 | Down | beta-1,4-xylosyltransferase IRX9 |
| DN91881_c0_g2_i1 | −2.95 | Down | casparian strip membrane protein 3 |
| DN92988_c2_g2_i1 | −1.20 | Down | GATA transcription factor 12 |
| DN92954_c10_g1_i1 | −1.18 | Down | uncharacterized protein LOC104819621 |
| cell wall modification | |||
| DN92674_c4_g1_i2 | −1.39 | Down | pectinesterase/pectinesterase inhibitor 21 |
| DN92816_c2_g3_i2 | −5.14 | Down | pectinesterase/pectinesterase inhibitor 59 |
| DN94343_c1_g1_i1 | 1.25 | Up | KMT00424|hypothetical protein BVRB_9g217270 |
| DN94343_c1_g2_i1 | 2.30 | Up | putative pectinesterase 11 |
| DN99628_c2_g1_i2 | −1.35 | Down | casparian strip membrane protein 2 |
| DN88314_c0_g1_i1 | −2.32 | Down | pectinesterase/pectinesterase inhibitor 12 |
| cellulose biosynthetic process | |||
| DN94431_c0_g5_i2 | 1.95 | Up | cellulose synthase-like protein G2 |
| DN94789_c0_g4_i1 | −5.27 | Down | protein trichome birefringence-like 42 |
| DN96659_c4_g1_i6 | 1.02 | Up | hypothetical protein SOVF_137370 |
| DN98575_c9_g1_i2 | 1.16 | Up | hypothetical protein SOVF_143010 |
| DN99723_c4_g4_i4 | −1.43 | Down | protein trichome birefringence-like 3 |
| DN94431_c0_g6_i2 | 1.67 | Up | hypothetical protein SOVF_076670 |
| DN97763_c1_g5_i1 | 1.13 | Up | hypothetical protein BVRB_5g111110 |
| DN100016_c2_g3_i2 | 1.52 | Up | hypothetical protein BVRB_6g142540 |
| DN100016_c2_g2_i1 | 3.83 | Up | hypothetical protein BVRB_6g142540 |
| cellulose catabolic process | |||
| DN94190_c4_g2_i1 | −1.28 | Down | endoglucanase 11 |
| DN95890_c3_g1_i4 | −1.23 | Down | endoglucanase 6 |
| DN96141_c3_g1_i4 | 1.19 | Up | uncharacterized protein LOC104893418 |
| DN95568_c3_g5_i1 | −1.87 | Down | endoglucanase 12 |
| DN86253_c0_g1_i1 | −2.54 | Down | hypothetical protein SOVF_009160 |
| ID | Log2FC (Cd-T/CK) | Regulate | Annotation |
|---|---|---|---|
| Flavonoid biosynthesis | |||
| DN110312_c0_g1_i1 | −3.55 | Down | chalcone synthase 2-like |
| DN91008_c0_g1_i2 | −1.63 | Down | protein SRG1-like |
| DN92655_c9_g1_i4 | 1.39 | Up | hypothetical protein SOVF_156310 |
| DN92655_c9_g6_i1 | 1.32 | Up | hypothetical protein SOVF_156310 |
| DN93138_c8_g4_i2 | 1.78 | Up | hypothetical protein SOVF_168370 |
| DN93301_c1_g1_i3 | −2.93 | Down | chalcone synthase 2-like |
| DN93562_c6_g3_i1 | 1.83 | Up | chalcone synthase |
| DN95283_c6_g3_i1 | 1.53 | Up | hypothetical protein SOVF_192090 |
| DN95456_c3_g5_i1 | 6.37 | Up | unnamed protein product |
| DN95974_c2_g2_i1 | 1.55 | Up | hypothetical protein SOVF_200110 |
| DN96124_c1_g1_i1 | −1.15 | Down | UDP-glycosyltransferase 79B6-like |
| DN96124_c1_g6_i2 | 1.60 | Up | hypothetical protein SOVF_025340 |
| DN96124_c1_g7_i2 | 1.28 | Up | hypothetical protein SOVF_025340 |
| DN96763_c2_g1_i1 | 2.09 | Up | chalcone synthase |
| DN99228_c1_g1_i1 | 1.55 | Up | flavanone-3-hydroxylase |
| DN99283_c4_g1_i1 | −1.82 | Down | hypothetical protein SOVF_168360 |
| phenylpropanoid biosynthesis | |||
| DN94161_c5_g1_i3 | 1.19 | Up | cinnamyl alcohol dehydrogenase 1 |
| DN98963_c5_g4_i1 | −2.27 | Down | hypothetical protein BVRB_8g195860 isoform B |
| DN74041_c0_g1_i1 | −6.91 | Down | class III peroxidase |
| DN86278_c0_g1_i1 | 4.72 | Up | peroxidase 4 |
| DN93922_c5_g2_i1 | 4.30 | Up | hypothetical protein PRUPE_6G205600 |
| DN96687_c1_g3_i2 | 1.06 | Up | phenylalanine ammonia-lyase, partial |
| DN98188_c1_g4_i4 | 1.07 | Up | phenylalanine ammonia-lyase |
| DN98963_c5_g4_i1 | −2.27 | Down | hypothetical protein BVRB_8g195860 isoform B |
| Monoterpenoid biosynthesis | |||
| DN93319_c3_g1_i1 | 1.81 | Up | (+)-neomenthol dehydrogenase-like, partial |
| DN96631_c4_g3_i1 | 2.21 | Up | (+)-neomenthol dehydrogenase isoform X2 |
| Cutin, suberine and wax biosynthesis | |||
| DN96918_c3_g5_i1 | 2.40 | Up | hypothetical protein SOVF_091970 |
| DN93789_c3_g1_i1 | −1.67 | Down | long chain acyl-CoA synthetase 1 |
| DN95362_c6_g2_i1 | −2.06 | Down | hypothetical protein BVRB_1g012280 |
| DN99249_c1_g1_i1 | 2.44 | Up | protein ECERIFERUM 3 |
| DN99249_c1_g3_i4 | 3.35 | Up | protein ECERIFERUM 3 |
| DN98976_c3_g1_i1 | −4.86 | Down | omega-hydroxypalmitate O-feruloyl transferase |
| carotenoid biosynthesis | |||
| DN98063_c4_g3_i1 | 1.04 | Up | cyclo-DOPA 5-O-glucosyltransferase |
| DN20061_c0_g1_i1 | 2.16 | Up | 9-cis-epoxycarotenoid dioxygenase NCED5, chloroplastic |
| Tropane, piperidine and pyridine alkaloid biosynthesis | |||
| DN98183_c5_g1_i1 | 1.22 | Up | hypothetical protein SOVF_178490 isoform A |
| ID | Primer Sequences 5′-3′ |
|---|---|
| DN92930_c2_g1_i1 | F: CCCTTTCACCCAGCCTAGTT |
| R: TTGTACCAACGCCCCAAAGA | |
| DN100451_c6_g2_i1 | F: TGGCTTCATGACAATCCGCT |
| R: AACGGAAGCCTAACCACTCG | |
| DN97551_c2_g1_i1 | F: GCGTGGACGGTATATGGGTT |
| R: GCTACTGGCCCCATGAAGTT | |
| DN99005_c3_g2_i1 | F: TGCAACCGTAAGCGGATACT |
| R: ATCCCCTCTAAACCCGTCCA | |
| DN99585_c3_g1_i2 | F: TTGCAGCCGCTTTCTATGGT |
| R: GCCCTCTTCTGGTATGTGGG | |
| DN95604_c7_g2_i2 | F: TCATGTTGGCTTGTGCTGGA |
| R: AGTTGAGCGCACAGGATTGA | |
| DN97835_c5_g5_i1 | F: AGAATGCGAAAGGCTTGGGA |
| R: TGTGCTTGGTCCACACTCAG | |
| DN97835_c5_g7_i3 | F: GTAACCTGGGTCATCGGCTC |
| R: TGGCTGCAATCGCATACCTT | |
| DN96587_c3_g1_i2 | F: AGTGTGGGGAGCCTAGTGAT |
| R: GGTCTAAAGGACCCACACG | |
| DN97830_c6_g5_i4 | F: GTAACACCTCCATCTCCGCC |
| R: AAAAGCCACCGGAAGAAGC | |
| DN89582_c0_g1_i1 | F: CCGAGACCCGTGTTTGTGTA |
| R: CTGTTTTGGGCTTCACCAGC | |
| DN95554_c2_g4_i1 | F: TGTGGCCCACTCAACTAAGG |
| R: AGATCCCCACCTGGATCAGA | |
| Actin | F: TTGAGCAGGAATCGGAG |
| R: TGCTGCTTCCATACCTATC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, Q.; Deng, W.; Su, Y.; Ma, L.; Yang, H.; Yao, F.; Lin, W. Transcriptome Analysis Reveals Novel Insights into the Hyperaccumulator Phytolacca acinosa Roxb. Responses to Cadmium Stress. Plants 2024, 13, 297. https://doi.org/10.3390/plants13020297
Xie Q, Deng W, Su Y, Ma L, Yang H, Yao F, Lin W. Transcriptome Analysis Reveals Novel Insights into the Hyperaccumulator Phytolacca acinosa Roxb. Responses to Cadmium Stress. Plants. 2024; 13(2):297. https://doi.org/10.3390/plants13020297
Chicago/Turabian StyleXie, Qin, Wentao Deng, Yi Su, Liying Ma, Haijun Yang, Feihong Yao, and Wanhuang Lin. 2024. "Transcriptome Analysis Reveals Novel Insights into the Hyperaccumulator Phytolacca acinosa Roxb. Responses to Cadmium Stress" Plants 13, no. 2: 297. https://doi.org/10.3390/plants13020297





