Abstract
Brassinazole resistant (BZR) genes act downstream of the brassinosteroid signaling pathway regulating plant growth and development and participating in plant stress responses. However, the BZR gene family has not systematically been characterized in potato. We identified eight BZR genes in Solanum tuberosum, which were distributed among seven chromosomes unequally and were classified into three subgroups. Potato and tomato BZR proteins were shown to be closely related with high levels of similarity. The BZR gene family members in each subgroup contained similar conserved motifs. StBZR genes exhibited tissue-specific expression patterns, suggesting their functional differentiation during evolution. StBZR4, StBZR7, and StBZR8 were highly expressed under white light in microtubers. StBZR1 showed a progressive up-regulation from 0 to 6 h and a progressive down-regulation from 6 to 24 h after drought and salt stress. StBZR1, StBZR2, StBZR4, StBZR5, StBZR6, StBZR7 and StBZR8 were significantly induced from 0 to 3 h under BR treatment. This implied StBZR genes are involved in phytohormone and stress response signaling pathways. Our results provide a theoretical basis for understanding the functional mechanisms of BZR genes in potato.
1. Introduction
A variety of endogenous and exogenous signals regulate plant growth and development. Plant hormones are major endogenous signaling molecules that can respond rapidly to environmental stimuli [1]. Brassinosteroids (BRs), a class of plant steroid hormones, play an important role in regulating plant growth and development, along with biotic and abiotic stress responses [2,3]. BRs fully stimulate the anti-stress potential of plants, alleviating damage caused when encountering various adverse external environmental cues through internal regulation [4]. Brassinazole resistant (BZR) is a family of transcription factors (TFs) that acts downstream of the BR signaling pathway, regulating plant growth and development and participating in plant stress responses by regulating the expression levels of some BR-responsive genes [5,6,7,8,9].
BZR proteins contain a nuclear localization sequence (NLS) at the N-terminal, a highly conserved DNA binding domain, a phosphorylation domain (which can be phosphorylated by BRASSINOSTEROID INSENSITIVE2-BIN2), a PEST sequence and a C-terminal domain [10]. BZR1 and Brassinosteroid insensitive 1-ethyl methanesulfonate-suppressor 1 (BES1) are two downstream TFs belonging to the BZR TF family [2]. BZR1 and BES1 share 88% amino acid sequence similarity, and the sequence consistency of the DNA binding domain is 97% [11]. They can be dephosphorylated by protein phosphatase when BR is detected [12]. Dephosphorylated BZR1 and BES1 accumulate in the nucleus and directly bind to cis-elements to regulate plant growth and development [13]. BZR1 and BES1 mediate crosstalk between BR and diverse signals such as other phytohormones, light, and stress, thereby regulating plant development and environment adaptability [5]. Both BZR1 and BES1 have a basic helix-loop-helix (bHLH) DNA binding motif in the N-terminal domain that is highly conserved across the whole family, although their functions have diverged [10]. BZR1 binds to a BR-Response Element (CGTGT/CG motif) to suppress the expression of BR-biosynthetic genes [14], while BES1 binds to an E box (CANNTG sequence) to activate BR induced gene expression [15].
The BZR gene family was first discovered and identified in Arabidopsis thaliana [10]. Research on BZR TFs in plant species has demonstrated their involvement in the regulation of cell elongation and division, plant morphology, flowering and fertility, quality improvement, and fruit ripening [16,17,18,19,20,21,22,23,24]. BZR1 directly regulates C-repeat binding factor 1 (CBF1) and CBF2 expression in Arabidopsis, thereby regulating freezing tolerance [25]. It was reported that AtBES1 plays a negative role in drought responses [26]. On the contrary, TaBZR2 positively regulates drought responses by activating TaGST1 [8]. BZR1/BES1 can directly bind to the promoters of several giberellin biosynthetic genes and control their expression in Arabidopsis and rice (Oryza sativa L.) [27,28]. Furthermore, TaBZR2 confers resistance to wheat stripe rust through the activation of chitinase Cht20.2 transcription [29]. SlBZR1 regulation of cold tolerance in tomato is related to its levels of phosphorylation [30]. BZR1 is a positive regulator of the BR signaling pathway and cooperates with light signal TFs to regulate cell elongation and plant growth [31,32]. BZR can also regulate the expression of drought-responsive glutathione s-transferase 1 (GST1) and interact with RESPONSIVE TO DESICCATION 26 (RD26) and WRKY TFs to modulate plant responses to drought, high temperature and freezing stress [33,34]. The Solanaceae lycopersicum BZR/BES TF SlBZR1 positively regulates BR signaling and salt stress tolerance in tomato and Arabidopsis [35]. In Pyrus bretschneideri, the BZR gene PbBZR1 also acts as a transcriptional repressor of lignin biosynthetic genes in fruits [36]. Overexpression of BpBZR1 enhances salt tolerance in Betula platyphylla [37]. GhBZR3 suppresses cotton fiber elongation by inhibiting very-long-chain fatty acid biosynthesis [38]. TOPLESS (TPL) mediates brassinosteroid-induced transcriptional repression through interaction with BZR1 [39]. Nine BES1 genes localized on eight chromosomes were identified in potato [40]. Exogenous BRs application increased both the number and the total weight of potato tuber [41]. StBRI1 is a functional potato BR receptor and has a novel function for brassinosteroid signaling in controlling tuberization [42]. The activation of reactive oxygen metabolism and phenylpropanoid metabolism by BR could accelerate the wound healing of potato tubers [43]. Transcription analysis during the sprouting process suggested that three BR signaling pathway genes, delta 24 sterol reductase (DWF1), brassinosteroid c-6 oxidase (BRD1) and brassinosteroid-insensitive 1 (BRI1), are all upregulated in potato [33], sterol metabolism linked with BZR genes in potato has not been reported. BRs accelerate the conversion of starch into soluble sugar in tubers, contributing to sprouting in potato [44].
Potato is the fourth-largest major crop in the world and a critical component of the human diet in some countries [45]. The importance of potatoes in securing food and nutritional security has been identified by the Food and Agriculture Organization (FAO) of the United Nations. Potato genome sequencing has been completed. The development of potato genome provides facilitation for exploring the function of related genes in potato growth and development [46,47,48,49]. Previous studies have shown that BR is involved in regulating tuber sprouting [33]. However, studies on the function and regulatory mechanism of BZR in potato, involved in tuber development, are still limited. There is a lack of systematic genome-wide identification and functional analysis of potato BZR TF gene family members.
In this study, from the whole potato genome, bioinformatic methods were used to identify BZR TF gene family members, carried out chromosome mapping, analyzed systematic evolution, characterized structural characteristics of genes and proteins, investigated contraction and expansion of the gene family, and identified cis acting elements in promoter regions. In addition, the expression profiles of potato BZR TF genes in different tissues and in response to abiotic stresses and hormones were analyzed. Our findings will lay the foundation for the further functional analysis of BZR TF genes in tuber formation and development in potato, especially in response to abiotic stresses.
2. Results
2.1. Identification and Chromosome Distribution of BZR Gene Family Members
A total of eight BZR genes were identified in the potato genome following a genome-wide analysis conducted according to chromosomal location (Table 1, Figure 1). The eight StBZR proteins encoded contained one conserved domain (Figure S1). The amino acid sequence characteristics of StBZR family proteins revealed the StBZR proteins were composed of 315–695 amino acids. The StBZR6 amino acid sequence was the shortest at 315 aa, while that of StBZR1 was the longest at 695 aa. The proteins molecular weights ranged from 33, 905.1 to 78, 032.6 Da, with StBZR1 exhibiting the highest molecular weight (78,032.6 Da) and StBZR6 the lowest (33,905.1 Da). The theoretical isoelectric points ranged from 5.41 to 9.11. The prediction of protein localization results revealed that all BZR proteins are essentially localized in the nucleus (Table 1), which is consistent with the function of TFs.

Table 1.
Profiles of the BZR gene family members identified in S. tuberosum.
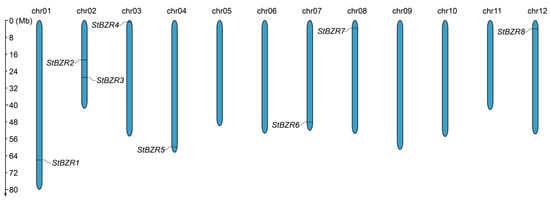
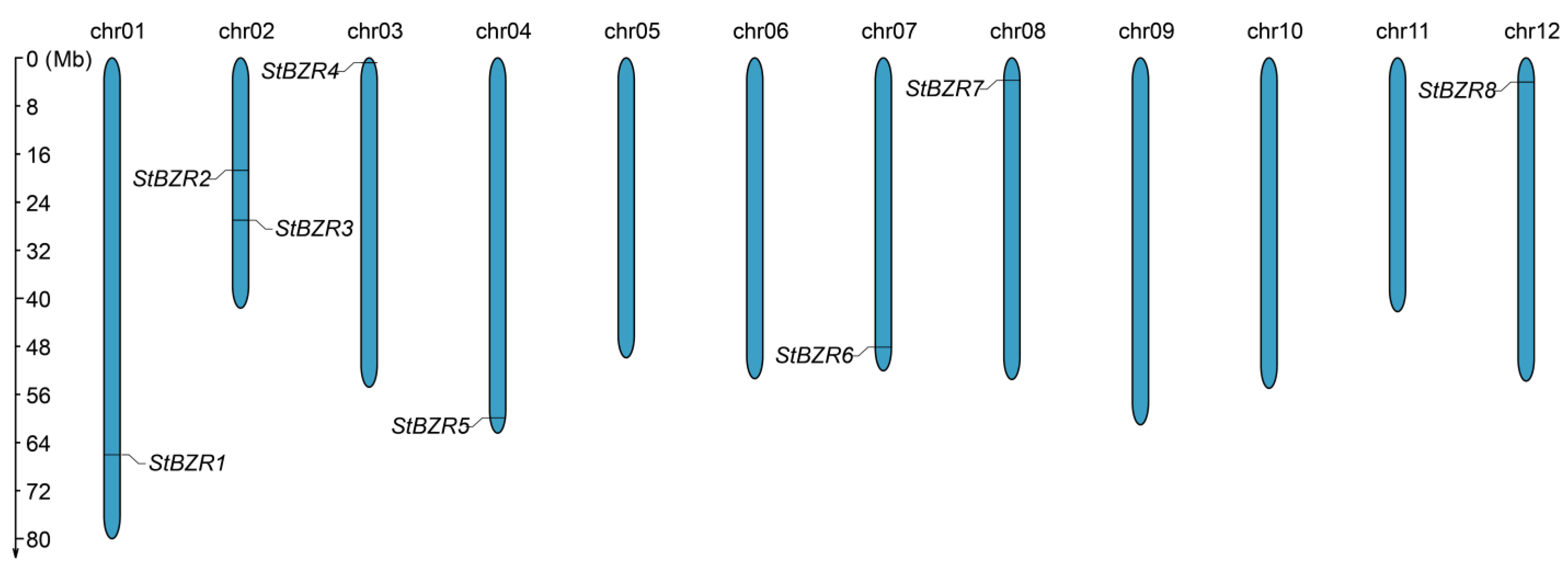
Figure 1.
Distribution of the StBZR genes on chromosomes. The left scale indicates chromosome length.
Chromosomal mapping results showed the eight StBZR genes to be unevenly distributed on seven chromosomes (Figure 1). Chr01, chr03, chr04, chr07, chr08 and chr12 each harbored one StBZR gene, while two were located on chr02.
2.2. Phylogenetic Classification and Analysis of BZR Genes
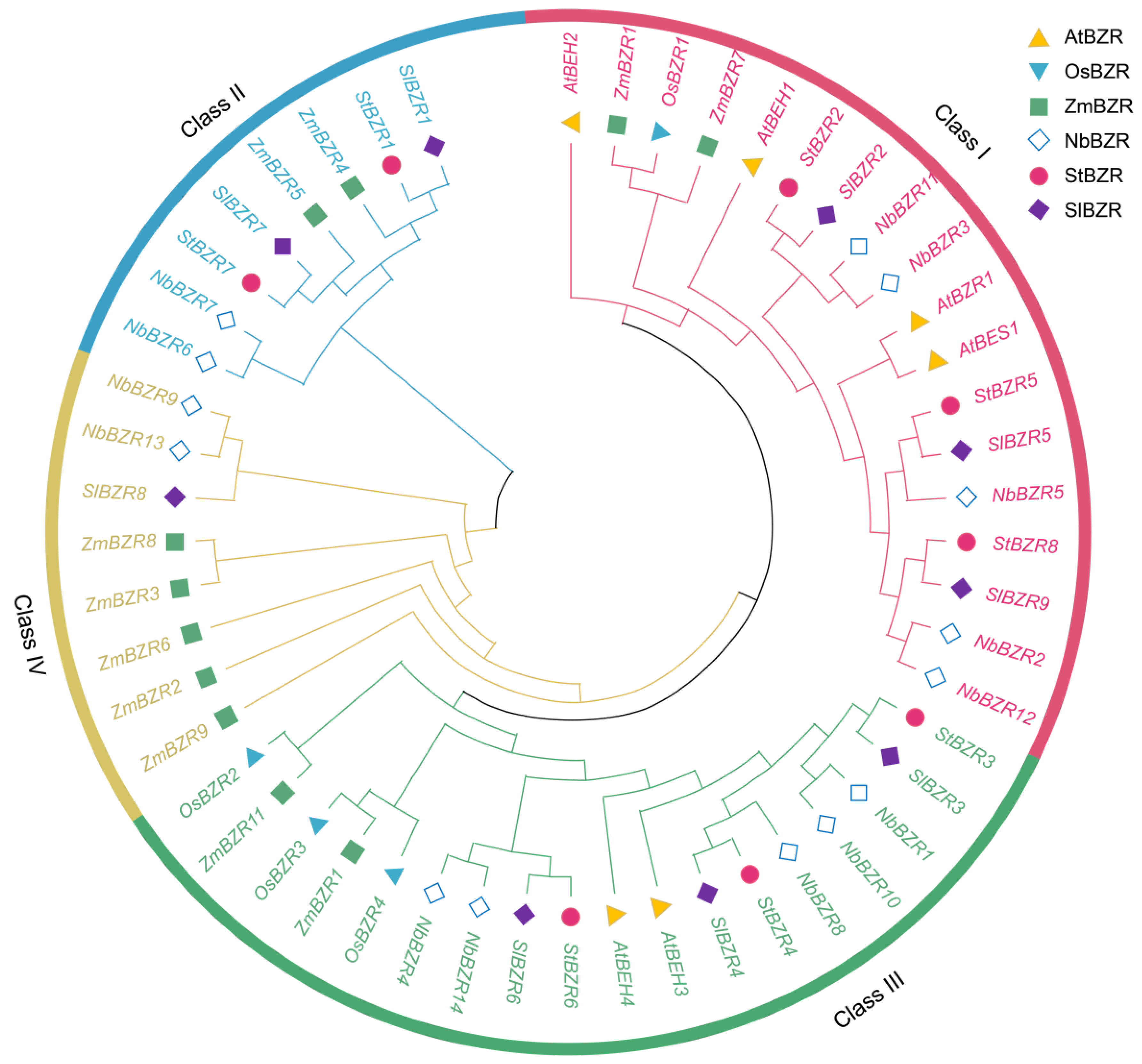
To investigate the phylogenetic relationships between BZR family genes, the amino acid sequences of BZR proteins from A. thaliana, S. tuberosum, S. lycopersicum, Nicotiana tabacum, O. sativa and Zea mays databases were retrieved and phylogenetic trees using multiple sequence alignment were constructed (Table S1). According to the topological structure of the phylogenetic tree, potato BZR proteins were divided into subgroups I, II and III (Figure 2). There were three BZR proteins in subgroup I, two BZR proteins in subgroup II and three BZR proteins in subgroup III. The BZR proteins from S. tuberosum and S. lycopersicum were found to be closely related, with high similarity.
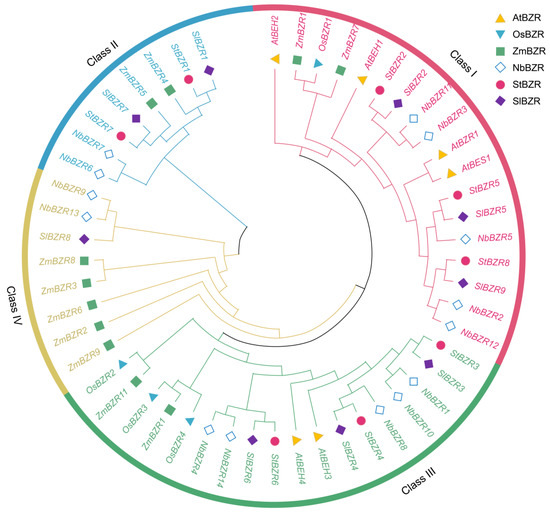
Figure 2.
A phylogenetic tree of BZR family genes from S. tuberosum (St), A. thaliana (At), S. lycopersicum (Sl), N. tabacum (Nb), O. sativa (Os) and Z. mays (Zm).
2.3. Gene Structure and Protein Motifs of BZR Genes
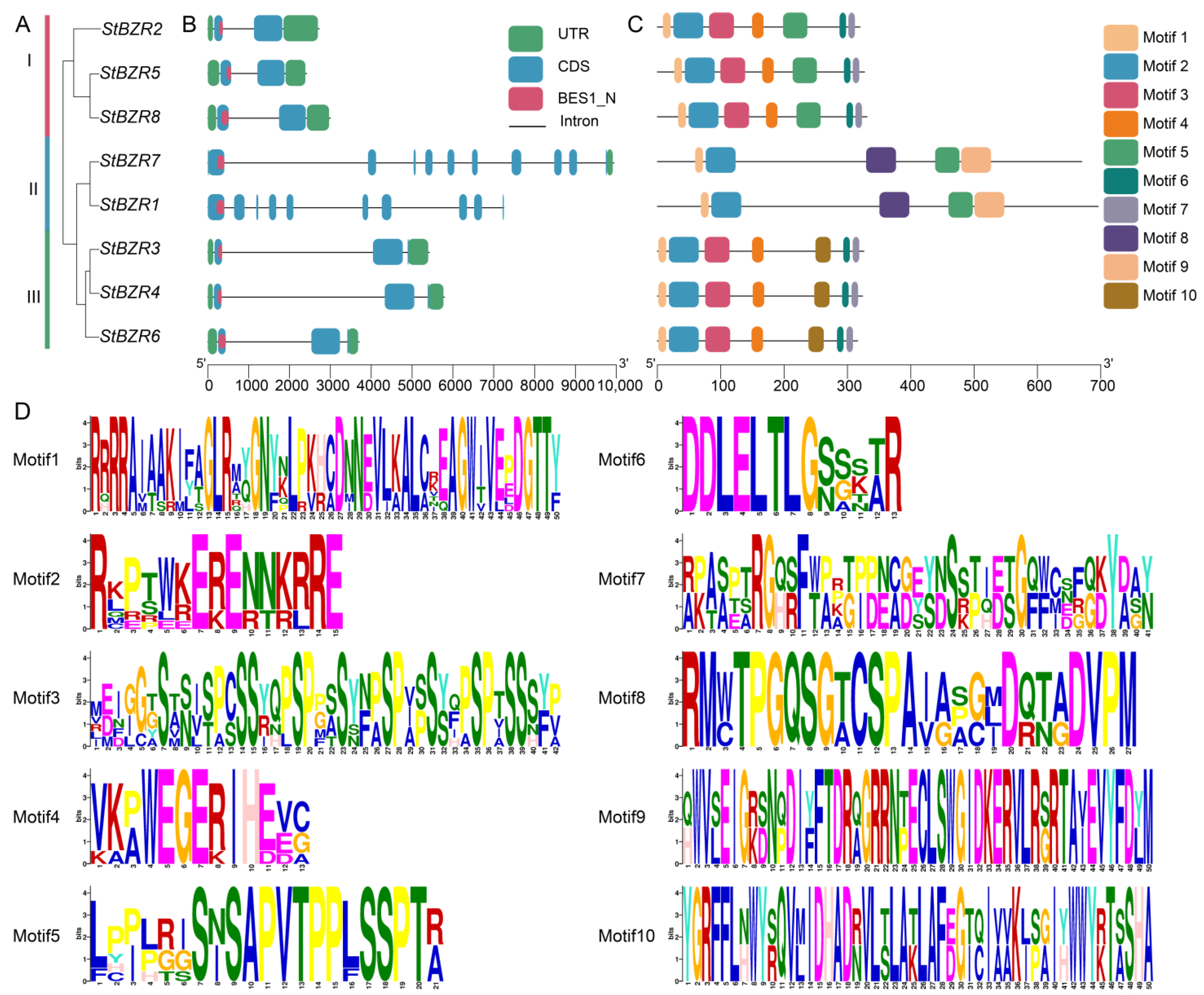
Eight BZR proteins evolutionary tree revealed that these proteins cluster into three subgroups in potato (Figure 3A). A gene structure analysis showed that the StBZR genes contained 2–10 exons and 4–7 introns. Subgroups I, II and III contained the conserved BES1_N functional domain (Figure 3B). StBZR1 and StBZR7 in subgroup II were longer, and StBZR2, StBZR5 and StBZR8 in subgroup I were shorter. Although genes lengths differed in the same subgroup, the exon–intron lengths and gene structures were similar within each subgroup. The motif distributions in BZR proteins were analyzed, and 10 conserved motifs were predicted (Figure 3C,D). The high similarity in the motif structure of the BZR proteins in the same subgroup indicated that the BZR gene members is highly conserved. Six members contained motifs 1, 2 and 3, while StBZR1 and StBZR7 did not contain motif 3.
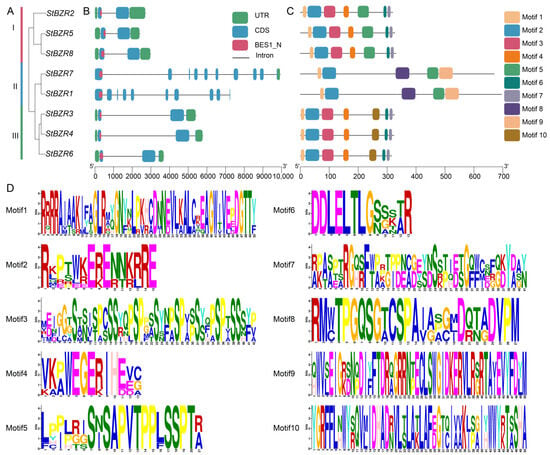
Figure 3.
The distributions of motif and exon-intron structures for BZR family members in S. tuberosum. (A) phylogenetic tree constructed using the StBZR protein sequences; (B) gene structure of StBZR members; UTR, untranslated region, represented by green boxes; CDS, coding sequence, represented by blue boxes. (C) ten types of conserved motifs were predicted in the StBZR protein sequences. (C) represent protein length with specific regions of conserved domains. (D) The sequence logo conserved motif of the potato BZR proteins.
2.4. Contraction versus Expansion
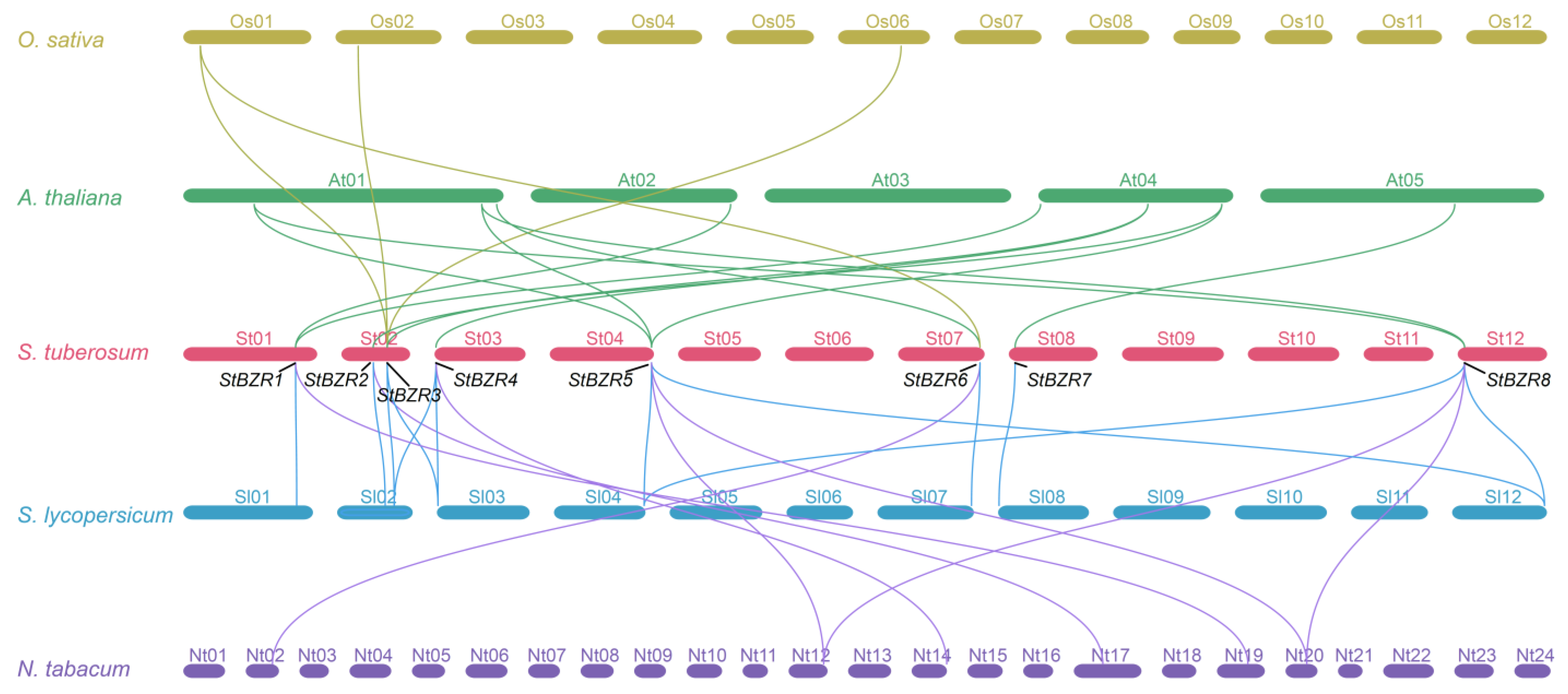
To study the contraction and expansion of the BZR gene family during evolution, the collinearity of orthologous BZR genes from A. thaliana, S. tuberosum, S. lycopersicum and N. tabacum were analyzed. There were eight, eight, and six putative orthologous StBZR genes identified in A. thaliana, S. lycopersicum, and N. tabacum, respectively, while 12, 12, and eight pairs of collinear genes were identified in the comparison between S. tuberosum and A. thaliana, S. lycopersicum, and N. tabacum, respectively (Figure 4, Table S3). These finding revealed that StBZR family genes number has expanded. The paralogous BZR gene pairs analysis showed that StBZR has two paralogous gene pairs in the potato genome, namely StBZR5/8 and StBZR3/4, with each gene located on a different chromosome (Figure S2).
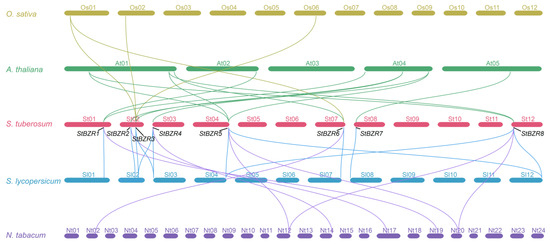
Figure 4.
Syntenic relationships analysis between BZR genes in O. sativa, A. thaliana, S. tuberosum, S. lycopersicum and N. tabacum. Yellow, green, blue and purple lines highlight syntenic BZR gene pairs.
2.5. Expression Patterns of StBZR Genes in Different Tissues by qRT-PCR
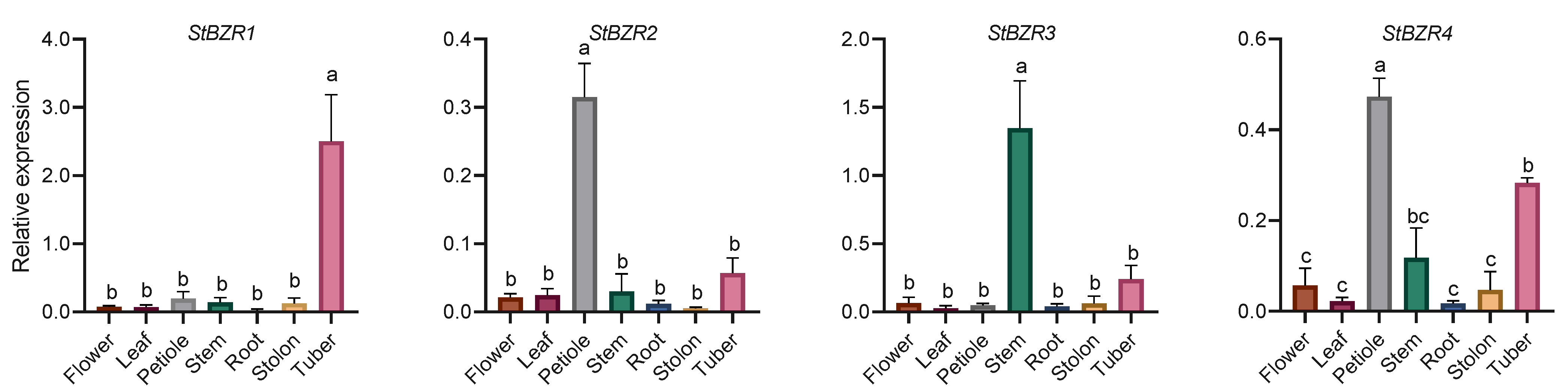
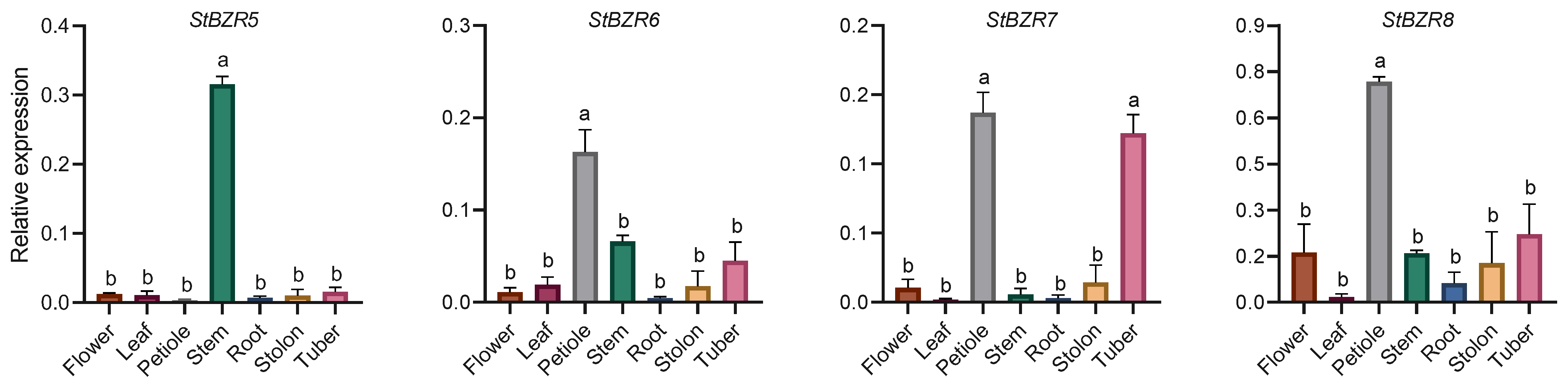
The expression characteristics of eight StBZR genes in seven tissues were analyzed. The expression levels of eight StBZR genes were significantly different among the various tissues (Figure 5). StBZR3 and StBZR5 expression patterns were similar, with high expression levels in stem and low expression in other tissues. Moreover, the expression patterns of StBZR2, StBZR6, and StBZR8 were similar, with the highest expression levels observed in petiole, followed by stem and tuber. The expression patterns of StBZR4 and StBZR7 were similar, with the highest expression levels seen in petiole and tuber. The expression of StBZR1 in tuber was higher than that in other tissues. In summary, StBZR genes were highly expressed in petiole, stem, and tuber, while StBZR1 was significantly specifically expressed only in tuber.
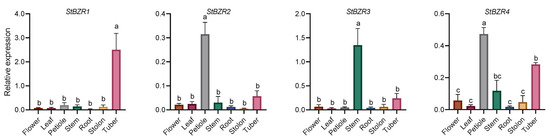
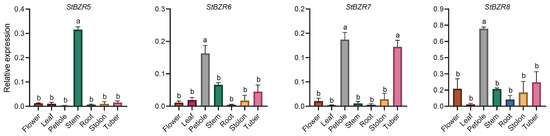
Figure 5.
Expression patterns of the eight StBZR genes in seven different tissues analyzed using qRT-PCR. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD.
2.6. Distribution of Cis-Reactive Elements on the Promoter of StBZR Genes
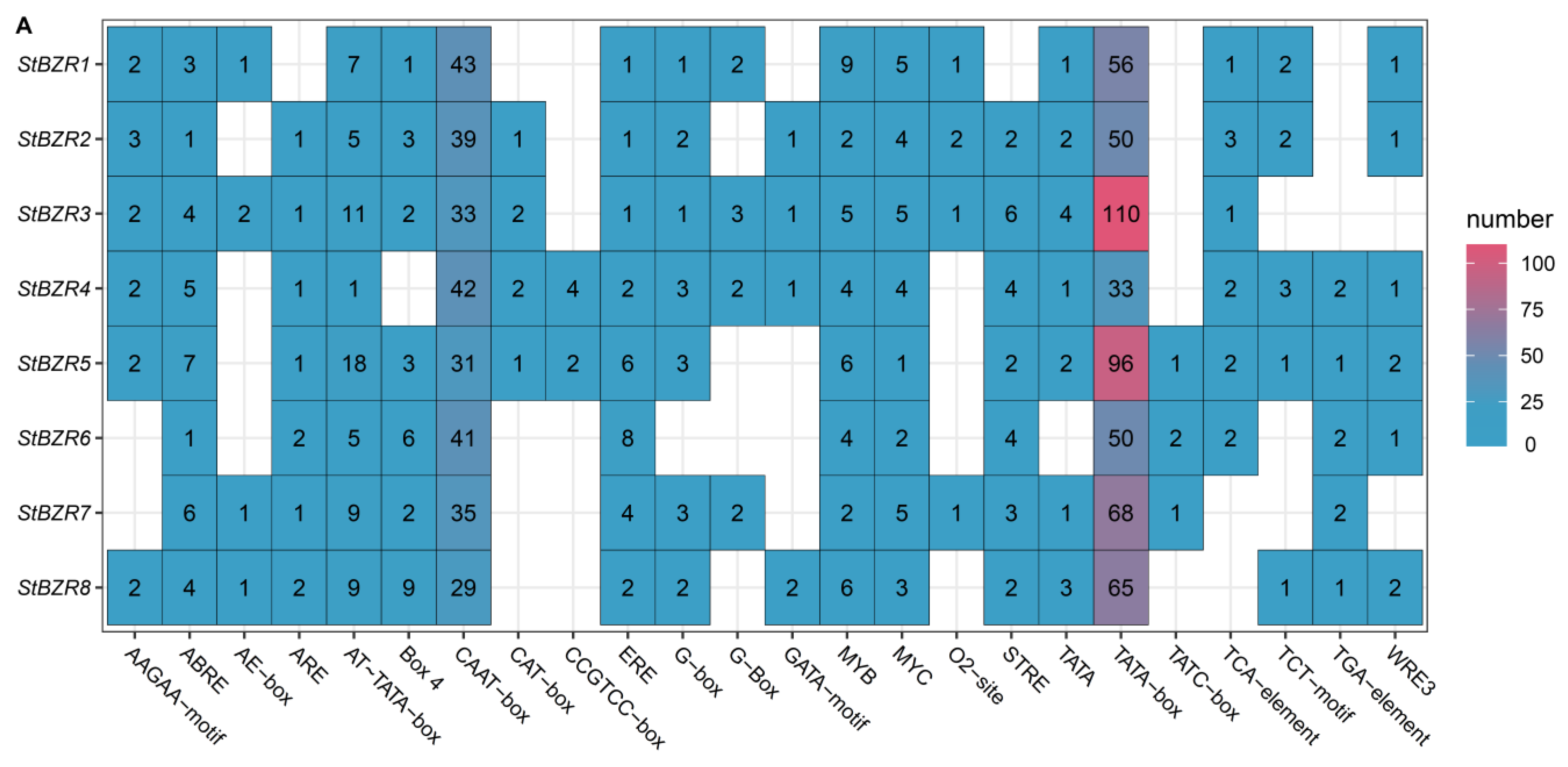
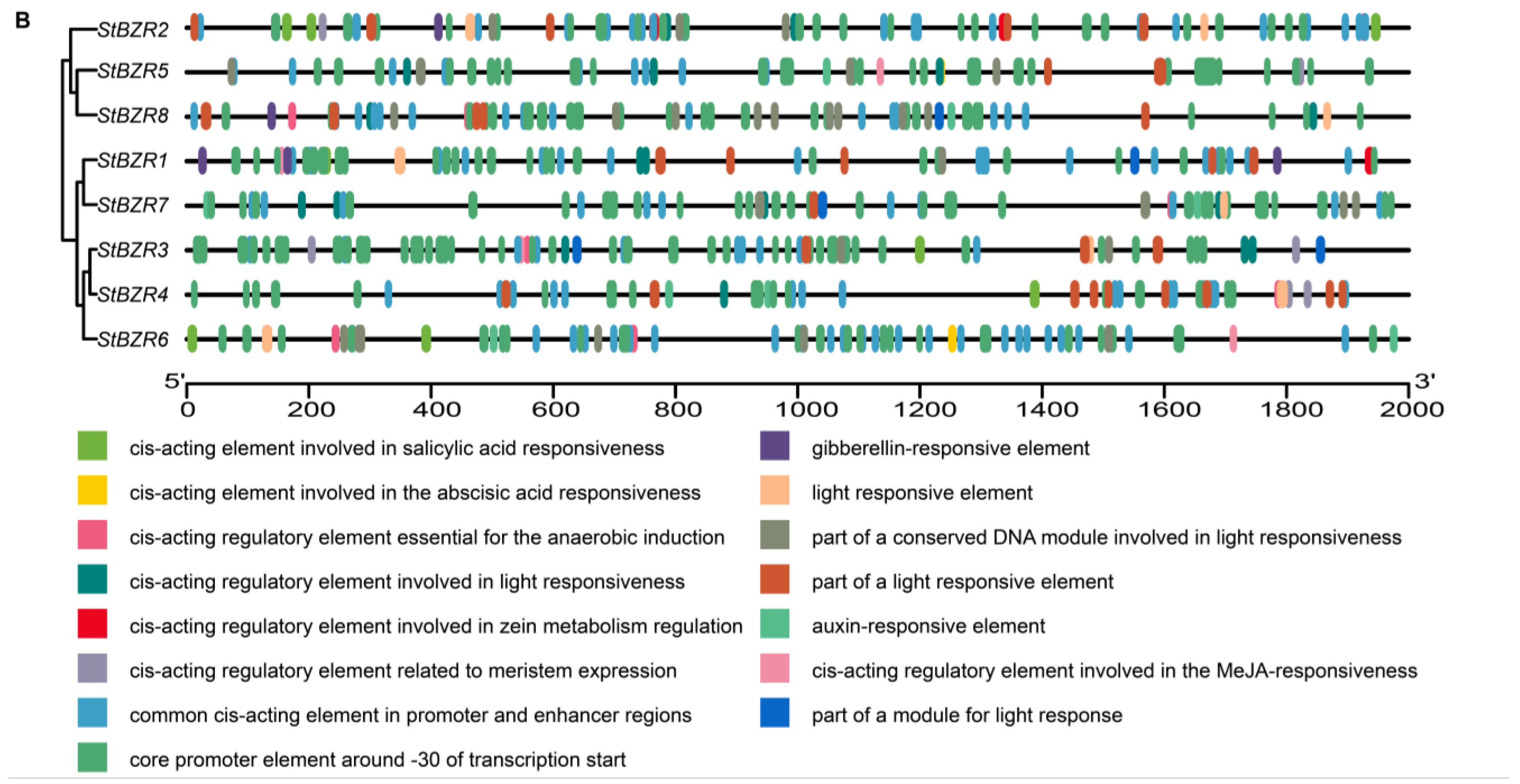
To investigate the cis-acting elements in the promoter regions of StBZR genes, approximately 2000 bp of sequence upstream of the translation initiation site were analyzed. The analysis revealed several regulatory elements related to the induction of phytohormones such as ABRE involved in abscisic acid (ABA), TGA element involved in auxin and the P-box and GARE-motif involved in gibberellin (GA) responsive elements. The cis-acting elements also included Sp1, G-box, and Box 4 involved in light responsiveness, the CGTCA-motif involved in MeJA-responsiveness, CAT-box related to meristem expression, ARE element essential to anaerobic induction, and the TCA-element involved in salicylic acid responsiveness (Figure 6A). In addition, the cis-acting elements related to drought and salt responses (Figure 6B). The results showed that all of the StBZR gene promoter regions contained various cis-regulatory elements involved in light, stress and hormone-responsiveness, which implied that the StBZR family may participate in many growth and development processes. Therefore, we analyzed the gene expression patterns under light, drought, salt, ABA, and BR treatments to check the StBZR genes in response to abiotic stresses and hormone.
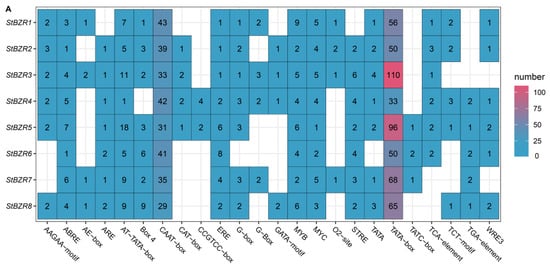
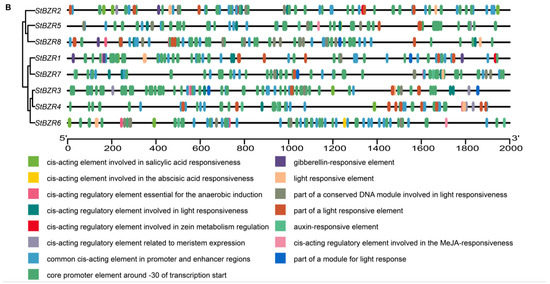
Figure 6.
Distribution of cis-elements in the promoter regions of StBZR genes. (A) Heat map representing cis-acting components. Colors represent different types of elements; and (B) distribution of cis-reactive elements in the promoter. The ruler at the bottom indicates the direction and length of the sequence.
2.7. Expression Profiles of StBZR Genes in Response to Abiotic Stresses and Hormones by qRT-PCR
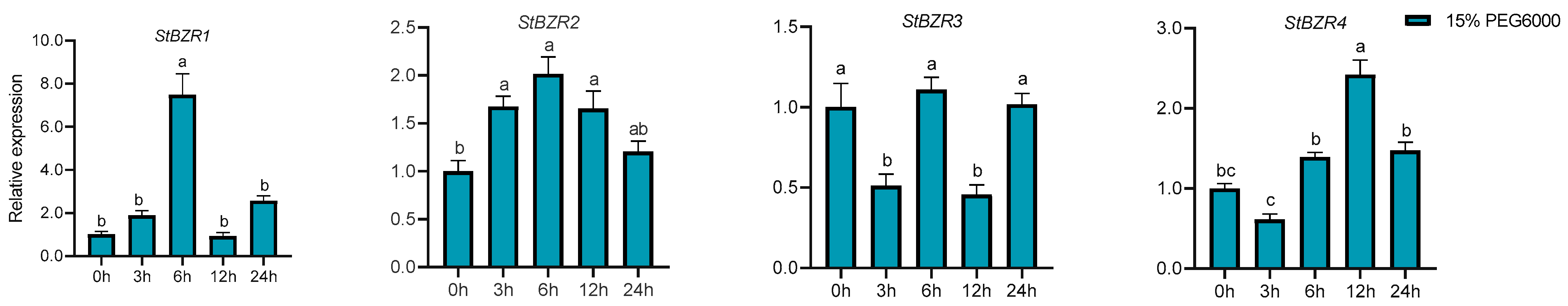
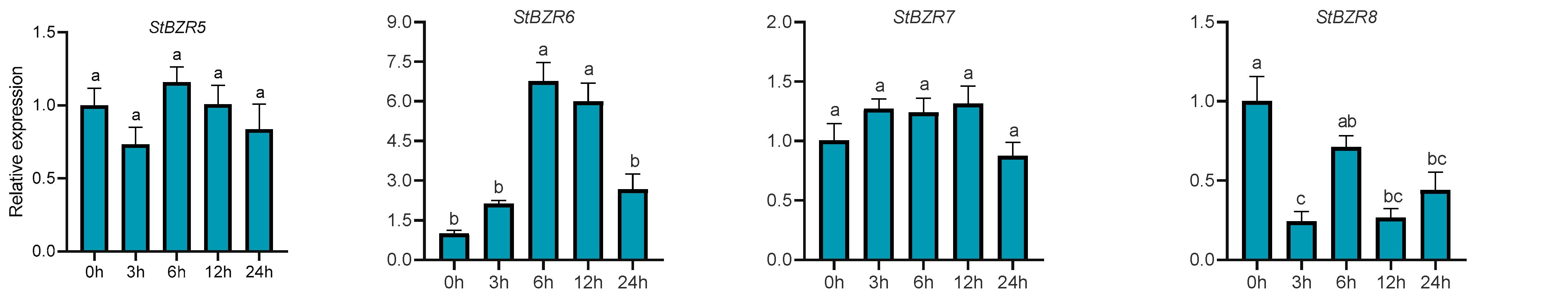
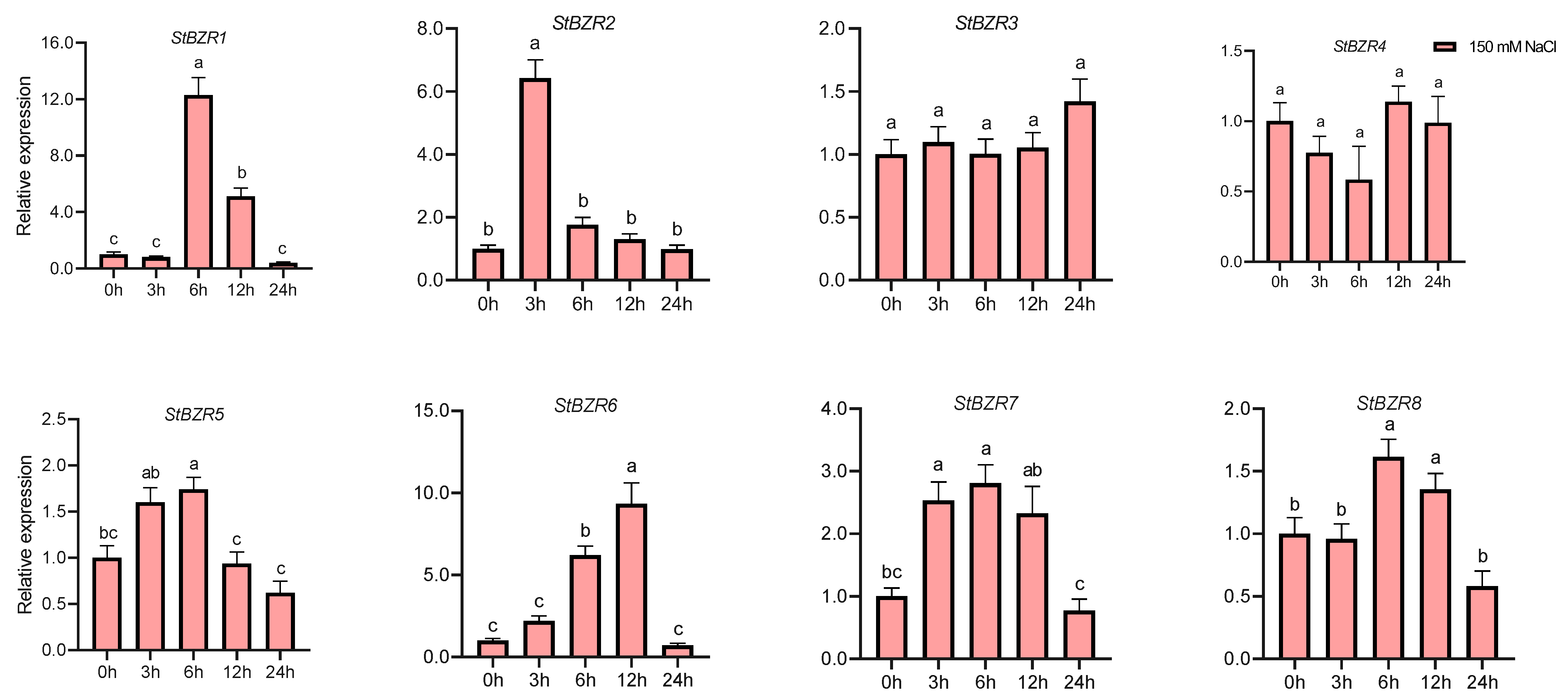
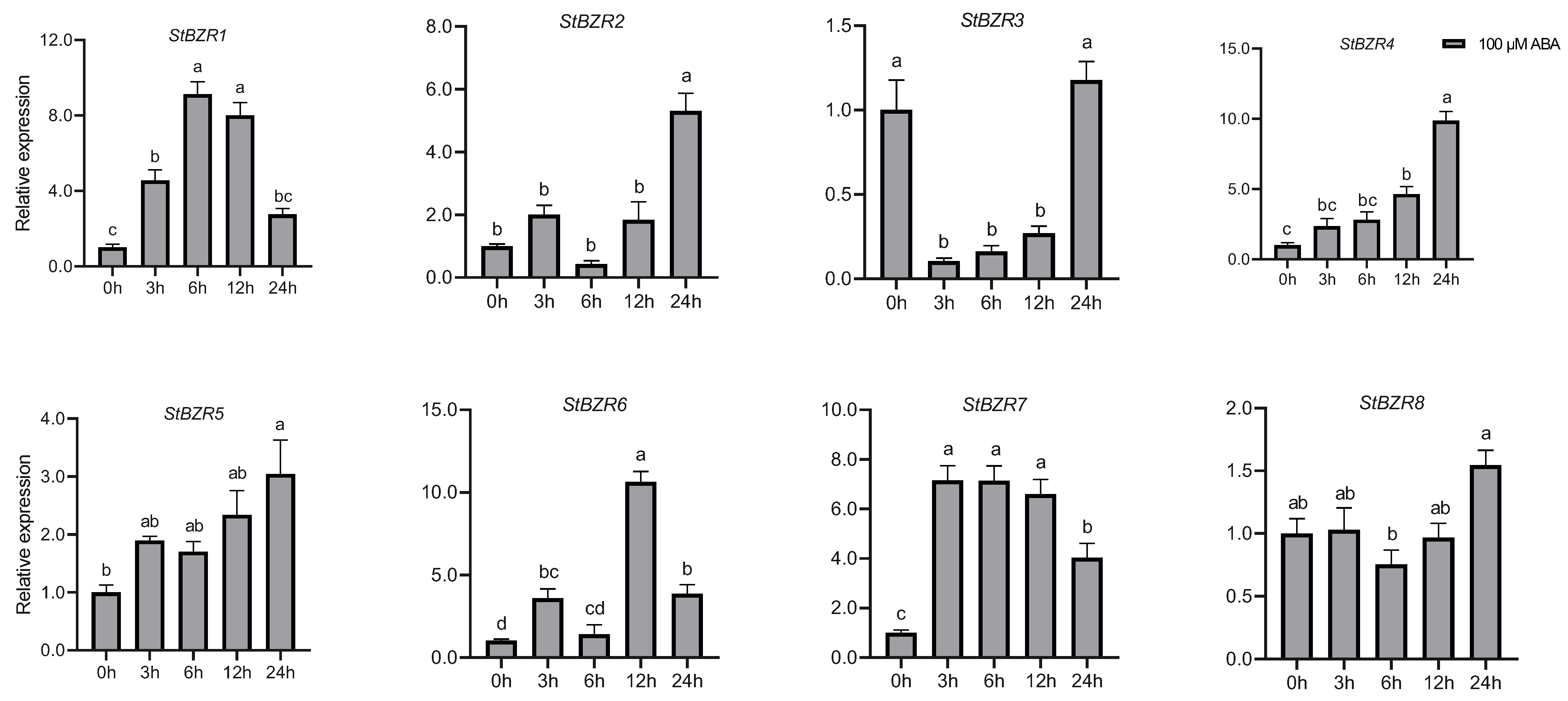
We examined the expression patterns of the StBZR genes under abiotic stresses in the presence of phytohormones. Plants were treated with various wavelengths of light, drought, salt, ABA, and BR. The leaf development of the potato plantlets was severely inhibited under monochromatic red light, there was no leaf sample under red light. The expression levels of StBZR1, StBZR3, StBZR4, StBZR7, and StBZR8 under blue light were higher than under white light in leaf, and in microtubers, the expression levels of StBZR4, StBZR7, and StBZR8 under white light were higher than under red and blue light (Figure S3). Under drought (15% PEG6000) treatment, StBZR1 showed progressive up-regulation from 0 to 6 h and progressive down-regulation from 6 to 24 h. StBZR8 was down-regulated under drought treatment (Figure 7). StBZR1, StBZR2, StBZR5, StBZR6, StBZR7, and StBZR8 were first up-regulated and then down-regulated under salt treatment (150 mM NaCl) (Figure 8). Under treatment with 100 μM ABA, StBZR4 was strongly induced, StBZR3 was strongly and rapidly inhibited from an early stage, and then induced after 12 h (Figure 9). Under 50 μM BR treatment, StBZR1, StBZR2, StBZR4, StBZR5, StBZR6, StBZR7, and StBZR8 were significantly induced from 0 to 3 h and then inhibited. In particular, the expression of StBZR2, StBZR5, and StBZR8 was increased about 10-fold after 3 h. StBZR3 was first down-regulated and then up-regulated (Figure S4).
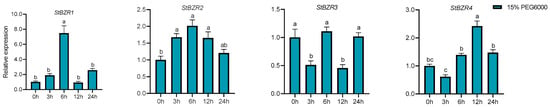
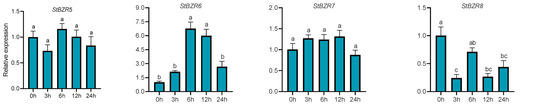
Figure 7.
Expression patterns of the eight StBZR genes under drought (15% PEG6000) stress. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD.
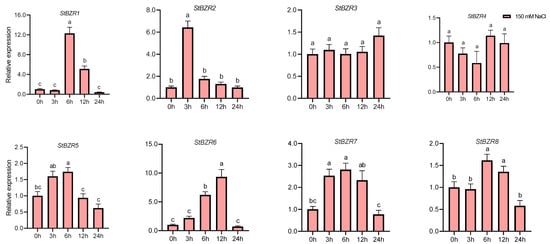
Figure 8.
Expression patterns of the eight StBZR genes under salt (150 mM, NaCl) stress. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD.
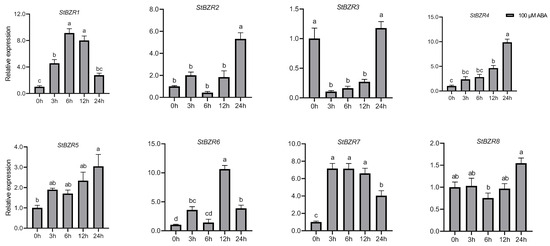
Figure 9.
Expression patterns of the eight StBZR genes after 100 μM ABA treatment. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD.
3. Discussion
The BZR gene family is an important TF family that regulates plant growth, development, and the BZR-mediated abiotic stress response. However, the BZR TF family members in S. tuberosum, an important and widespread food and vegetable crop, have not been thoroughly investigated to date. In this study, we characterized the BZR genes in S. tuberosum and discovered discrepancies and variations in gene sequences, structures, and conserved motifs. We also identified instances of both the conservation and divergence of gene and protein expression in this family. Expression data analysis and cis-regulation prediction further revealed that potato BZR genes may be intrinsically involved in regulating pathways associated with plant development and stress resistance. The genome-wide identification and characterization of BZR TF family members in S. tuberosum is an essential starting point for in-depth exploration of the function of this gene family. The accumulation of genomic and transcriptomic data from S. tuberosum will provide insights into the functional and molecular characteristics of the BZR gene family.
BZR signaling components are highly conserved between primitive to modern plants [14]. Although plant genomes differ in size and numbers of genes, many BZR gene family members have been identified. For instance, six BZR genes have been characterized in A. thaliana [10], 11 in Z. mays [50], six in Beta vulgaris [51], nine in S. lycopersicum [52], five and two in wheat and foxtail millet, respectively [5], and six in Cucumis sativus [11]. In our study, a total of eight BZR genes were identified in S. tuberosum (Table 1), and these were unevenly distributed on seven chromosomes (Figure 1). The number of gene family members has shrunk or expanded in different species [5,10,11,50,51,52], due to environmental adaptations that have occurred during evolution.
Plant regulatory genes evolve quickly, but the rates at which different domains of BZR family proteins have evolved differ substantially. Subgroup I, II, and III members have no similarities in gene structure, and may have evolved from different origins. In our study, it was found that the StBZR gene does not belong to the intron-enriched gene family, so it is inferred that the continuous diversification of the StBZR gene family may lead to intron loss [53]. Group I of the BZR genes in potato contains three StBZR genes, group II contains two StBZR genes, and group III contains three StBZR genes. The gene structure in groups I and III is relatively conserved, while that in group II is considerably different (Figure 3), indicating that expansion of the BZR gene family has mainly occurred in subgroup III. To investigate the evolutionary relationship between BZR family members among species, we included BZR proteins from S. tuberosum, S. lycopersicum, N. tabacum, O. sativa, and Z. mays in a phylogenetic analysis and identified the conserved motifs. Cucumber and tomato were found to be closely related [11], while the phylogenetic trees of potato and tomato (S. lycopersicum) were shown to be similar (Figure 2). This result is consistent with the taxonomic relationships between potato and tomato within the Solanaceae.
BZR family genes are specifically expressed in different tissues and organs of many plants [34]. In Arabidopsis, the transcriptional expression of BZR genes is higher in roots and buds but lower in stems, fruits, and flowers [15,54], while in maize, these genes are highly expressed in seedlings and endosperm [50]. In tomato, SlBZR2 and SlBZR9 are generally expressed at high levels in most tissues, while other SlBZR genes are highly expressed only in certain tissues [55]. In our study, there were significant differences in the expression levels of the eight StBZR genes among various tissues in potato (Figure 5). The expression patterns of StBZR3 and StBZR5 were similar, with high and low expression levels seen in the stem and other tissues, respectively. StBZR2, StBZR6, and StBZR8 were highly expressed in the petiole, followed by the stem and tuber (Figure 5). The differential expression of StBZR family members in different tissues indicates that these genes play a regulatory role in the growth and development of potato, and that there is likely to be functional redundancy. BRs are not only involved in starch utilization but also in the regulation of sucrose transport [56]. In sugar beet, BZR genes are involved in accumulating sugars in the taproot, eventually leading to increases in root diameter and weight [51]. The expression of StBZR1 was demonstrated to be higher in the tuber than other tissues (Figure 5), indicating that this gene is could be involved in promoting sugar accumulation and thereby tuber expansion.
The differential pattern of StBZR gene expression in various tissues (Figure 5) suggested that these genes may act as growth regulators. An analysis of the promoter regions of the StBZR genes identified in this study revealed the existence of a variety of cis-acting elements that are involved in regulating the temporal and spatial expression levels of the genes (Figure 6). The StBZR gene promoter regions contained various cis-regulatory elements involved in light, stress and hormone-responsiveness, which implied that the StBZR family may participate in growth and development processes by responding to different signals (Figure 6). Differential expression patterns among the StBZRs were also observed following light spectrum, drought, salt, ABA and BR treatments. Thus, StBZR genes may participate in the regulation of plant development and abiotic stress resistance pathways. Some reports have confirmed that the BZR family is involved in a variety of hormone signaling pathways [57,58,59], stress responses [7,25,34], and plant growth regulation [60,61]. The analysis of gene expression suggested that some StBZR members participate in multiple stress responses. For example, the expression levels of StBZR4, StBZR7, and StBZR8 under white light were higher than those under red and blue light, in microtubers (Figure S3). Under drought and salt stress, StBZR1 was up-regulated first and then down-regulated, showing progressive up-regulation from 0 to 6 h and progressive down-regulation from 6 to 24 h after treatment (Figure 7 and Figure 8). StBZR4 was strongly induced under ABA treatment (Figure 9), whereas, under BR treatment, StBZR1, StBZR2, StBZR4, StBZR5, StBZR6, StBZR7, and StBZR8 were significantly induced from 0 to 3 h and then inhibited (Figure S4). This implied that these StBZR genes are involved in phytohormone and stress response signaling pathways. These data provide valuable insights into the functional mechanisms of the StBZR gene family in response to phytohormones and different biotic and abiotic stresses during plant development.
4. Materials and Methods
4.1. Plant Materials
Plantlets of potato (cv. Shepody) were provided by Anhui Science and Technology University. Plantlets of potato were cut into 1–1.5 cm segments with leaves. Then placed them into medium in tissue culture bottles (63 mm of inner diameter; 85 mm of height). Four stem segments were placed in each tissue culture bottle and were placed in a tissue culture room. The medium used for propagation of plantlets in vitro was solid Murashige and Skoog (MS) medium (4%, w/v, sucrose, 0.9%, w/v, agar). The medium used for the induction of microtubers was solid MS medium (8%, w/v, sucrose, 0.9%, w/v, agar). The relative humidity was 65 ± 5%, day-time temperature was 22 ± 2 °C, night temperature was 18 ± 2 °C, photoperiod uniformly was set to 8-h light/16-h dark and photosynthetic photon flux density was 65 μmol m–2 s–1 in tissue culture room. Potato plantlets grown for 30 d were acclimated and transplanted to pots containing vegetative soil and vermiculite (1:1) and placed in a plant growth room where the relative humidity was 65 ± 5%, the temperature was 22 ± 2 °C, the light intensity was 200 μmol m–2 s–1 and the photoperiod was set at 12-h light/12-h dark.
4.2. Identification of BZR Transcription Factor Gene Family Members in S. tuberosum
We downloaded the genome sequence, proteins and corresponding coding sequences of S. tuberosum (Version 6.1) from the Phytozome v13 website (https://phytozome.jgi.doe.gov/pz/portal.html, accessed on 1 September 2023). The BES1_N domain file (PF05687) from the InterPro website (www.ebi.ac.uk/interpro/entry/pfam/PF05687/, accessed on 1 September 2023) were downloaded and uploaded it to the HMMERv3.3.2 (https://www.ebi.ac.uk/Tools/hmmer/, accessed on 1 September 2023) to search for potential genes in the potato genome containing this conserved domain, with an E < 1 × 10−5 [62]. The sequences of A. thaliana and O. sativa BZR family members were downloaded from uniport (https://www.uniprot.org/, accessed on 1 September 2023) and compared with potato protein sequences using blast-2.11.0, with an E < 1 × 10−5. Based on the above method, putative BZR candidates were selected. After removing redundant results, the remaining sequences were further verified for the existence of BES1_N domains using other databases: Simple Modular 132 Architecture Research Tool (SMART, http://smart.emblheidelberg.de/, accessed on 1 September 2023), NCBI Batch CD-Search Tool (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi, accessed on 1 September 2023) and the sequences with lacking BES1_N domains were removed. Finally, the protein sequences with BES1_N domains were taken and named sequentially according to their locations on the chromosomes. We used the website Cell—Ploc 2.0 (http://www.csbo.sjtu.edu.cn/bioinf/Cell-PLoc-2, accessed on 1 September 2023) for predicting the subcellular localization of candidate genes encoding BZR proteins [63]. We used the online software ExPASy (Version 3.0, https://web.expasy.org/protparam/, accessed on 1 September 2023) to analyze the physicochemical properties of the amino acids encoded by the candidate genes.
4.3. Chromosomal Location
We used TBtools v1.09876 software [64] to calculate the location of each StBZR gene and length information of chromosome. We used MG2C (http://mg2c.iask.in/mg2c_v2.0/, accessed on 6 September 2023) to construct the physical location of StBZR genes on the chromosomes.
4.4. Phylogenetic Tree Reconstruction, Gene Structure and Protein Motifs Analysis
Multiple sequence alignments were performed on the potato BZR protein sequences using ClustalW [65] with default parameters. BZR protein sequences of O. sativa, A. thaliana, Z. mays, N. tabacum, and S. lycopersicum were downloaded from NCBI (https://www.ncbi.nlm.nih.gov/, accessed on 3 September 2023), and the BZR protein sequences of potato and these species were used to reconstruct a phylogenetic tree. We used MEGAX 10.1.8 software [66] and the neighbor-joining method, with the bootstrap value set to 1000 cycles to reconstruct the phylogenetic tree of the BZR protein family members. The BZR member gff3 file was submitted to GSDS (http:/gsds.cbi.pku.edu.cn, accessed on 3 September 2023) for gene structure analysis [67]. We used the MEME (http://meme-suite.org/, accessed on 4 September 2023) website for the motif prediction, with the number set to 10. Batch CD-search (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi, accessed on 6 September 2023) was used to analyze conserved domain structure.
4.5. Analyses of Duplication Type and Synteny
We used Blast-2.13.0 + for comparison, and MCScanX [68] to calculate collinearity of homologous BZR genes within and between species, with a threshold of E < 10−5. Using MEGAX [66] analyzed multiple sequence alignment of StBZR members. Using TBtools v1.09876 software [64] showed the collinearity of BZR among A. thaliana, O. sativa, N. tabacum, S. lycopersicum and S. tuberosum to judge collinearity of the BZR gene families.
4.6. Tissue Expression Characteristics of the StBZR Genes in S. tuberosum
The leaves, petioles, stem sections grown for 30 dflower tissues grown for 45 days and stolons, roots, tubers grown for 60 d were collected under plant growth room, performing three biological replicates. These tissues were immediately flash-frozen in liquid nitrogen and stored at −80 °C for RNA extraction and gene expression analysis.
4.7. Cis-Regulatory Element Analysis of the StBZR Genes
We extracted sequences in the 2000 base pairs (bp) upstream of the start codon (ATG) in the BZR genes from the genome of S. tuberosum. For promoter cis-acting regulatory element screening, we analyzed the sequences extracted above using the PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/, accessed on 15 September 2023). Using TBtools v1.09876 software [64] drew the cis-elements in the promoter region.
4.8. Light Spectrum, Abiotic Stress and Hormone Treatments
Robust and uniform potato seedlings, which had been grown in test tubes for 30 d were selected for this study. The stems were cut into 1–1.5 cm segments and each section was placed in a separate glass test tube under aseptic conditions. The tubes were cultivated under dark conditions for 2 days and then grown under blue light at a wavelength of 460 nm, red light at a wavelength of 620 nm, and white light. Leaves and stolons of potato plantlets grown for 35 d and microtubers grown for 80 d were collected. Drought and salt were set as abiotic stress treatments. All reagents are purchased from Shanghai MACKLIN Co., Ltd, China. PEG 6000 and NaCl were added to 1 L of nutrient solution to prepare 15% PEG 6000 and 150 mM NaCl solutions, respectively. ABA and BR were dissolved in 100 mL distilled water to produce 1 mM solutions, which were then diluted accordingly. Hormone treatments involved the application of ABA (100 μM) and BR (50 μM). We collected samples at 0, 3, 6, 12, and 24 h post-treatment, and untreated seedlings from the same batch were used as controls. Three biological replicates were collected at each time point, with each replicate consisting of three independent seedlings.
4.9. qRT-PCR
We performed RNA extraction and reverse transcription as described by Huang et al. (2017) [69]. We performed qRT-PCR according to Jin et al. [70]. Using an ABIViiA7 real-time PCR instrument (Life Technologies, Carlsbad, CA, USA) performed qRT-PCR. Using Primer 3.0 tool (https://bioinfo.ut.ee/primer30.4.0/, accessed on 10 September 2023) designed the StBZR gene-specific amplification primers. We used elongation factor-1alpha (EF1α) as a reference gene [71] (Table S2). We performed three independent biological replicates of qPCR, and each PCR reaction was performed in triplicate. The 2–∆CT method was used to calculate the relative expression levels of genes in different tissues. The 2–∆∆CT method was used to calculate the relative expression levels of genes under light spectrum, abiotic stress and hormone treatments [72]. SPSS software (Version 20) was used for the statistical analysis. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). GraphPad Prism 9.5.0 software was used for drawing of data.
5. Conclusions
Eight BZR genes were identified from the genome of potato. These classify into three subgroups and are distributed on seven chromosomes unevenly. StBZR and SlBZR proteins are closely related phylogenetically and display high sequence similarity. 12 genes were orthologous to the BZR genes in S. tuberosum and S. lycopersicum. Tissue specific expression characteristics suggest functional differentiation of StBZR genes during evolution. StBZR gene promoters contain many regulatory elements that are involved in phytohormone, light and stress signaling. This study offers a basis to predict the functions of BZR genes in potato, and lay a foundation for further research of the biological functions of BZR genes in potato. In order to further deepen our understanding of the function of BZR genes, our future direction will focus on the study of the specific regulatory mechanisms of BZR genes in regulating potato tuber formation and development.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/plants13030407/s1, Table S1: Profiles of BZR gene family members of different species; Table S2: Information of primer sequence; Table S3: Gene ID of syntenic genes in other species; Figure S1: BZR amino acid sequence alignment of S. tuberosum. The BES1_N domain is boxed with light red color; Figure S2: Syntenic relationships of the BZR genes within S. tuberosum. The black lines represent gene pairs; Figure S3: Expression patterns of the eight StBZR genes under different spectrum. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD. The leaf development of the potato plantlets was severely inhibited under monochromatic red light, there was no leaf sample under red light; Figure S4: Expression patterns of the eight StBZR genes after BR (50 μM, 0–24 h) treatment. Different letters indicate significant differences between different tissues. Significant differences among the groups were compared based on Tukey’s test (p < 0.05). The data points represent mean ± SD.
Author Contributions
Conceptualization, R.L. and X.H.; Methodology, X.Y.; Software, R.L. and H.A.; Validation, X.Y. and T.F.; Formal Analysis, R.L. and T.L.; Investigation, R.L. and X.H.; Resources, R.L.; Data Curation, B.Z. and T.L.; Writing—Original Draft Preparation, R.L.; Writing—Review & Editing, R.L. and X.H.; Visualization, H.A. and T.F.; Supervision, R.L. and X.H.; Project Administration, R.L. and X.H.; Funding Acquisition, R.L. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Natural Science Foundation of Anhui (No. 2108085QC125), the Key Project of Anhui Provincial Education Department (No. 2022AH051627, No. 2023AH051886), the Talent Introduction Start up Fund Project of Anhui Science and Technology University (No. NXYJ202102, No. NXYJ202001), and the National Innovation and Entrepreneurship Training Program for College Students (No. 202310879078).
Data Availability Statement
All data analyzed during this study are included in the Supplementary Information Files.
Conflicts of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- Ma, B.; Zhu, J.; Huang, X. Diversification of plant SUPPRESSOR OF MAX2 1 (SMAX1)-like genes and genome-wide identification and characterization of cotton SMXL gene family. BMC Plant Biol. 2023, 23, 419. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wu, X.; Qiu, S.; Zheng, H.; Lu, Y.; Peng, J.; Wu, G.; Chen, J.; Rao, S.; Yan, F. Genome-wide identification and expression profiling of the BZR transcription factor gene family in Nicotiana benthamiana. Int. J. Mol. Sci. 2021, 22, 10379. [Google Scholar] [CrossRef]
- Vukašinović, N.; Wang, Y.; Vanhoutte, I.; Fendrych, M.; Guo, B.; Kvasnica, M.; Jiroutová, P.; Oklestkova, J.; Strnad, M.; Russinova, E. Local brassinosteroid biosynthesis enables optimal root growth. Nat. Plants 2021, 7, 619–632. [Google Scholar] [CrossRef] [PubMed]
- Nolan, T.M.; Vukašinović, N.; Liu, D.; Russinova, E.; Yin, Y. Brassinosteroids: Multidimensional regulators of plant growth, development, and stress responses. Plant Cell 2020, 32, 295–318. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Cui, Y.; Zhao, Z.; Li, S.; Liang, D.; Wang, C.; Feng, G.; Wang, J.; Liu, Z. Genome-wide identification and characterization of the BES/BZR gene family in wheat and foxtail millet. BMC Genom. 2021, 22, 682. [Google Scholar] [CrossRef]
- Li, L.; Deng, X.W. It runs in the family: Regulation of brassinosteroid signaling by the BZR1-BES1 class of transcription factors. Trends Plant Sci. 2005, 10, 266–268. [Google Scholar] [CrossRef]
- Li, Y.; He, L.; Li, J.; Chen, J.; Liu, C. Genome-wide identification, characterization, and expression profiling of the legume BZR transcription factor gene family. Front. Plant Sci. 2018, 9, 1332. [Google Scholar] [CrossRef]
- Yu, H.Q.; Sun, F.A.; Feng, W.Q.; Lu, F.Z.; Li, W.C.; Fu, F.L. The BES1/BZR1 transcription factors regulate growth, development and stress resistance in plants. Yi Chuan 2019, 41, 206–214. [Google Scholar] [CrossRef]
- Bai, Q.; Wang, L.; Huang, S.; Ali, K.; Li, G.; Ren, H.; Zheng, B. The receptor-like kinase EMS1 and BRI1 coordinately regulate stamen elongation via the transcription factors BES1/BZR1 in Arabidopsis. Plant Sci. 2023, 331, 111673. [Google Scholar] [CrossRef]
- Yin, Y.; Vafeados, D.; Tao, Y.; Yoshida, S.; Asami, T.; Chory, J. A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell 2005, 120, 249–259. [Google Scholar] [CrossRef]
- Luo, S.; Zhang, G.; Zhang, Z.; Wan, Z.; Liu, Z.; Lv, J.; Yu, J. Genome-wide identification and expression analysis of BZR gene family and associated responses to abiotic stresses in cucumber (Cucumis sativus L.). BMC Plant Biol. 2023, 23, 214. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Yuan, M.; Wang, R.; Yang, Y.; Wang, C.; Oses-Prieto, J.A.; Kim, T.W.; Zhou, H.W.; Deng, Z.; Gampala, S.S.; et al. PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat. Cell Biol. 2011, 13, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Li, L.; Zola, J.; Aluru, M.; Ye, H.; Foudree, A.; Guo, H.; Anderson, S.; Aluru, S.; Liu, P.; et al. A brassinosteroid transcriptional network revealed by genome-wide identification of BESI target genes in Arabidopsis thaliana. Plant J. 2011, 65, 634–646. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Y.; Nakano, T.; Gendron, J.; He, J.; Chen, M.; Vafeados, D.; Yang, Y.; Fujioka, S.; Yoshida, S.; Asami, T.; et al. Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev. Cell 2002, 2, 505–513. [Google Scholar] [CrossRef] [PubMed]
- Yin, Y.; Wang, Z.Y.; Mora-Garcia, S.; Li, J.; Yoshida, S.; Asami, T.; Chory, J. BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 2002, 109, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Park, C.H.; Kim, T.W.; Son, S.H.; Hwang, J.Y.; Lee, S.C.; Chang, S.C.; Kim, S.H.; Kim, S.W.; Kim, S.K. Brassinosteroids control AtEXPA5 gene expression in Arabidopsis thaliana. Phytochemistry 2010, 71, 380–387. [Google Scholar] [CrossRef] [PubMed]
- Chaiwanon, J.; Wang, Z.Y. Spatiotemporal brassinosteroid signaling and antagonism. with auxin pattern stem cell dynamics in Arabidopsis roots. Curr. Biol. 2015, 25, 1031–1042. [Google Scholar] [CrossRef]
- Li, Q.F.; He, J.X. BZR1 interacts with HY5 to mediate brassinosteroid- and light-regulated cotyledon opening in Arabidopsis in darkness. Mol. Plant 2016, 9, 113–125. [Google Scholar] [CrossRef]
- Tong, H.; Jin, Y.; Liu, W.; Li, F.; Fang, J.; Yin, Y.; Qian, Q.; Zhu, L.; Chu, C. DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J. 2009, 58, 803–816. [Google Scholar] [CrossRef]
- Fang, Z.; Ji, Y.; Hu, J.; Guo, R.; Sun, S.; Wang, X. Strigolactones and brassinosteroids antagonistically regulate the stability of the D53-OsBZR1 complex to determine FC1 expression in rice tillering. Mol. Plant 2020, 13, 586–597. [Google Scholar] [CrossRef]
- Domagalska, M.A.; Schomburg, F.M.; Amasino, R.M.; Vierstra, R.D.; Nagy, F.; Davis, S.J. Attenuation of brassinosteroid signaling enhances FLC expression and delays flowering. Development 2007, 134, 2841–2850. [Google Scholar] [CrossRef] [PubMed]
- Ye, Q.; Zhu, W.; Li, L.; Zhang, S.; Yin, Y.; Ma, H.; Wang, X. Brassinosteroids control male fertility by regulating the expression of key genes involved in Arabidopsis anther and pollen development. Proc. Natl. Acad. Sci. USA 2010, 107, 6100–6105. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Jia, C.; Zhang, M.; Chen, D.; Chen, S.; Guo, R.; Guo, D.; Wang, Q. Ectopic expression of a BZR1-1D transcription factor in brassinosteroid signalling enhances carotenoid accumulation and fruit quality attributes in tomato. Plant Biotechnol. J. 2014, 12, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.F.; Shan, W.; Liang, S.M.; Wu, C.J.; Wei, W.; Chen, J.Y.; Lu, W.J.; Kuang, J.F. MaBZR1/2 act as transcriptional repressors of ethylene biosynthetic genes in banana fruit. Physiol. Plant. 2019, 165, 555–568. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Ye, K.; Shi, Y.; Cheng, J.; Zhang, X.; Yang, S. BZR1 positively regulates freezing tolerance via CBF-dependent and CBF-independent pathways in Arabidopsis. Mol. Plant 2017, 10, 545–559. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Yin, Y. WRKY transcription factors are involved in brassinosteroid signaling and mediate the crosstalk between plant growth and drought tolerance. Plant Signal. Behav. 2017, 12, e1365212. [Google Scholar] [CrossRef] [PubMed]
- Tong, H.; Xiao, Y.; Liu, D.; Gao, S.; Liu, L.; Yin, Y.; Jin, Y.; Qian, Q.; Chu, C. Brassinosteroid regulates cell elongation by modulating gibberellin metabolism in rice. Plant Cell 2014, 26, 4376–4393. [Google Scholar] [CrossRef]
- Unterholzner, S.J.; Rozhon, W.; Papacek, M.; Ciomas, J.; Lange, T.; Kugler, K.G.; Mayer, K.F.; Sieberer, T.; Poppenberger, B. Brassinosteroids are master regulators of gibberellin biosynthesis in Arabidopsis. Plant Cell 2015, 27, 2261–2272. [Google Scholar] [CrossRef]
- Bai, X.; Zhan, G.; Tian, S.; Peng, H.; Cui, X.; Islam, M.A.; Goher, F.; Ma, Y.; Kang, Z.; Xu, Z.S.; et al. Transcription factor BZR2 activates chitinase Cht20.2 transcription to confer resistance to wheat stripe rust. Plant Physiol. 2021, 187, 2749–2762. [Google Scholar] [CrossRef]
- Fang, P.; Wang, Y.; Wang, M.; Wang, F.; Chi, C.; Zhou, Y.; Zhou, J.; Shi, K.; Xia, X.; Foyer, C.H.; et al. Crosstalk between brassinosteroid and redox signaling contributes to the activation of CBF expression during cold responses in tomato. Antioxidants 2021, 10, 509. [Google Scholar] [CrossRef]
- Li, J.; Nam, K.H.; Vafeados, D.; Chory, J. BIN2, a new brassinosteroid-insensitive locus in Arabidopsis. Plant Physiol. 2001, 127, 14–22. [Google Scholar] [CrossRef]
- Sun, Y.; Fan, X.Y.; Cao, D.M.; Tang, W.; He, K.; Zhu, J.Y.; He, J.X.; Bai, M.Y.; Zhu, S.; Oh, E.; et al. Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev. Cell 2010, 19, 765–777. [Google Scholar] [CrossRef] [PubMed]
- Li, L.Q.; Zou, X.; Deng, M.S.; Peng, J.; Huang, X.L.; Lu, X.; Fang, C.C.; Wang, X.Y. Comparative morphology, transcription, and proteomics study revealing the key molecular mechanism of camphor on the potato tuber sprouting effect. Int. J. Mol. Sci. 2017, 18, 2280. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.Y.; Gao, Y.; Guo, J.; Yu, T.F.; Zheng, W.J.; Liu, Y.W.; Chen, J.; Xu, Z.S.; Ma, Y.Z. BES/BZR transcription factor TaBZR2 positively regulates drought responses by activation of TaGST1. Plant Physiol. 2019, 180, 605–620. [Google Scholar] [CrossRef] [PubMed]
- Jia, C.; Zhao, S.; Bao, T.; Zhao, P.; Peng, K.; Guo, Q.; Gao, X.; Qin, J. Tomato BZR/BES transcription factor SlBZR1 positively regulates BR signaling and salt stress tolerance in tomato and Arabidopsis. Plant Sci. 2021, 302, 110719. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Meng, D.; Li, X.; Wang, L.; Cai, Y.; Jiang, L. A chinese white pear (Pyrus bretschneideri) BZR gene PbBZR1 Act as a transcriptional repressor of lignin biosynthetic genes in fruits. Front. Plant Sci. 2020, 11, 1087. [Google Scholar] [CrossRef] [PubMed]
- Lv, J.; Li, Y.; Liu, Z.; Li, X.; Lei, X.; Gao, C. Response of BpBZR genes to abiotic stress and hormone treatment in Betula platyphylla. Plant Physiol. Biochem. 2020, 151, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Chen, X.; Xue, H.; Jia, T.; Meng, F.; Liu, Y.; Luo, X.; Xiao, G.; Zhu, S. GhBZR3 suppresses cotton fiber elongation by inhibiting very-long-chain fatty acid biosynthesis. Plant J. 2022, 111, 785–799. [Google Scholar] [CrossRef]
- Oh, E.; Zhu, J.Y.; Ryu, H.; Hwang, I.; Wang, Z.Y. TOPLESS mediates brassinosteroid-induced transcriptional repression through interaction with BZR1. Nat. Commun. 2014, 5, 4140. [Google Scholar] [CrossRef]
- Zhu, W.J.; Jiao, D.L.; Zhang, J.; Xue, C.M.; Chen, M.; Yang, Q. Genome-wide identification and analysis of BES1/BZR1 transcription factor family in potato (Solanum tuberosum. L). Plant Growth Regul. 2020, 92, 375–387. [Google Scholar] [CrossRef]
- Zhu, W.J.; Chen, F.; Li, P.P.; Chen, Y.M.; Chen, M.; Yang, Q. Identification and characterization of brassinosteroid biosynthesis and signaling pathway genes in Solanum tuberosum. Russ. J. Plant Physiol. 2019, 66, 628–636. [Google Scholar] [CrossRef]
- Huang, S.H.; Zheng, C.Y.; Zhao, Y.; Li, Q.; Liu, J.W.; Deng, R.; Lei, T.T.; Wang, S.F.; Wang, X.F. RNA interference knockdown of the brassinosteroid receptor BRI1 in potato (Solanum tuberosum L.) reveals novel functions for brassinosteroid signaling in controlling tuberization. Sci. Hortic. 2021, 290, 110516. [Google Scholar] [CrossRef]
- Han, Y.; Yang, R.R.; Zhang, X.J.; Wang, Q.H.; Wang, B.; Zheng, X.Y.; Li, Y.C.; Prusky, D.; Bi, Y. Brassinosteroid accelerates wound healing of potato tubers by activation of reactive oxygen metabolism and phenylpropanoid metabolism. Foods 2022, 11, 906. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Deng, M.; Lyu, C.; Zhang, J.; Peng, J.; Cai, C.; Yang, S.; Lu, L.; Ni, S.; Liu, F.; et al. Quantitative phosphoproteomics analysis reveals that protein modification and sugar metabolism contribute to sprouting in potato after BR treatment. Food Chem. 2020, 325, 126875. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, D.; Carlson-Nilsson, U.; Ortíz, R.; Andreasson, E. Overview and breeding strategies of table potato production in Sweden and the Fennoscandian region. Potato Res. 2016, 59, 279–294. [Google Scholar] [CrossRef]
- Pham, G.M.; Hamilton, J.P.; Wood, J.C.; Burke, J.T.; Zhao, H.; Vaillancourt, B.; Ou, S.; Jiang, J.; Buell, C.R. Construction of a chromosome-scale long-read reference genome assembly for potato. Giga Sci. 2020, 9, giaa100. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Tang, D.; Huang, W.; Yang, Z.; Zhang, Y.; Hamilton, J.P.; Visser, R.G.F.; Bachem, C.W.B.; Robin, B.C.; Zhang, Z.; et al. Haplotype-resolved genome analyses of a heterozygous diploid potato. Nat. Genet. 2020, 52, 1018–1023. [Google Scholar] [CrossRef]
- Zhang, C.; Yang, Z.; Tang, D.; Zhu, Y.; Wang, P.; Li, D.; Zhu, G.; Xiong, X.; Shang, Y.; Li, C.; et al. Genome design of hybrid potato. Cell 2021, 184, 3873–3883. [Google Scholar] [CrossRef]
- Tang, D.; Jia, Y.; Zhang, J.; Li, H.; Cheng, L.; Wang, P.; Bao, Z.; Liu, Z.; Feng, S.; Zhu, X.; et al. Genome evolution and diversity of wild and cultivated potatoes. Nature 2022, 606, 535–541. [Google Scholar] [CrossRef]
- Manoli, A.; Trevisan, S.; Quaggiotti, S.; Varotto, S. Identification and characterization of the BZR transcription factor family and its expression in response to abiotic stresses in Zea mays L. Plant Growth Regul. 2018, 84, 423–436. [Google Scholar] [CrossRef]
- Wang, W.; Sun, Y.Q.; Li, G.L.; Zhang, S.Y. Genome-wide identification, characterization, and expression patterns of the BZR transcription factor family in sugar beet (Beta vulgaris L.). BMC Plant Biol. 2019, 19, 191. [Google Scholar] [CrossRef] [PubMed]
- Su, D.; Xiang, W.; Wen, L.; Lu, W.; Shi, Y.; Liu, Y.; Li, Z. Genome-wide identification, characterization and expression analysis of BES1 gene family in tomato. BMC Plant Biol. 2021, 21, 161. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, J.; Hu, Z.; Guo, X.; Tian, S.; Chen, G. Genome-wide analysis of the MADS-Box transcription factor family in Solanum lycopersicum. Int. J. Mol. Sci. 2019, 20, 2961. [Google Scholar] [CrossRef] [PubMed]
- He, J.X.; Gendron, J.M.; Sun, Y.; Gampala, S.S.; Gendron, N.; Sun, C.Q.; Wang, Z.Y. BZR1 is a transcriptional repressor with dual roles in brassinosteroid homeostasis and growth responses. Science 2005, 307, 1634–1638. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Zhu, X.; Wei, X. Genome-wide identification, structural analysis, and expression profiles of the BZR gene family in tomato. J. Plant Biochem. Biotechnol. 2022, 31, 739–750. [Google Scholar] [CrossRef]
- Bitterlich, M.; Krügel, U.; Boldt-Burisch, K.; Franken, P.; Kühn, C. Interaction of brassinosteroid functions and sucrose transporter SlSUT2 regulate the formation of arbuscular mycorrhiza. Plant Signal. Behav. 2014, 9, e970426. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Dou, L.; Gong, Z.; Wang, X.; Mao, T. BES1 hinders ABSCISIC ACID INSENSITIVE5 and promotes seed germination in Arabidopsis. New Phytol. 2019, 221, 908–918. [Google Scholar] [CrossRef]
- Li, Q.; Xu, F.; Chen, Z.; Teng, Z.; Sun, K.; Li, X.; Yu, J.; Zhang, G.; Liang, Y.; Huang, X.; et al. Synergistic interplay of ABA and BR signal in regulating plant growth and adaptation. Nat. Plants 2021, 7, 1108–1118. [Google Scholar] [CrossRef]
- Zhong, C.; Patra, B.; Tang, Y.; Li, X.; Yuan, L.; Wang, X. A transcriptional hub integrating gibberellin-brassinosteroid signals to promote seed germination in Arabidopsis. J. Exp. Bot. 2021, 72, 4708–4720. [Google Scholar] [CrossRef]
- Lachowiec, J.; Mason, G.A.; Schultz, K.; Queitsch, C. Redundancy, feedback, and robustness in the Arabidopsis thaliana BZR/BEH gene family. Front. Genet. 2018, 9, 523. [Google Scholar] [CrossRef]
- Xia, X.; Dong, H.; Yin, Y.; Song, X.; Gu, X.; Sang, K.; Zhou, J.; Shi, K.; Zhou, Y.; Foyer, C.H.; et al. Brassinosteroid signaling integrates multiple pathways to release apical dominance in tomato. Proc. Natl. Acad. Sci. USA 2021, 118, e2004384118. [Google Scholar] [CrossRef] [PubMed]
- Potter, S.C.; Luciani, A.; Eddy, S.R.; Park, Y.; Lopez, R.; Finn, R.D. HMMER web server: 2018 update. Nucleic Acids Res. 2018, 46, W200–W204. [Google Scholar] [CrossRef] [PubMed]
- Kesawat, M.S.; Kherawa, B.S.; Singh, A.; Dey, P.; Kabi, M.; Debnath, D.; Saha, D.; Khandual, A.; Rout, S.; Manorama, A.A.; et al. Genome-wide identification and characterization of the brassinazole-resistant (BZR) gene family and its expression in the various developmental stage and stress conditions in Wheat (Triticum aestivum L). Int. J. Mol. Sci. 2021, 22, 8743. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Higgins, D.G. Multiple sequence alignment using ClustalW and ClustalX. Curr. Protoc. Bioinform. 2002, 2.3.1–2.3.22. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yang, L.; Jin, Y.; Lin, J.; Liu, F. Generation, annotation, and analysis of a large-scale expressed sequence tag library from Arabidopsis pumila to explore salt-responsive genes. Front. Plant Sci. 2017, 8, 955. [Google Scholar] [CrossRef]
- Jin, Y.; Liu, F.; Huang, W.; Sun, Q.; Huang, X. Identification of reliable reference genes for qRT-PCR in the ephemeral plant Arabidopsis pumila based on full-length transcriptome data. Sci. Rep. 2019, 9, 8408. [Google Scholar] [CrossRef]
- Tang, X.; Zhang, N.; Si, H.; Calderón-Urrea, A. Selection and validation of reference genes for RT-qPCR analysis in potato under abiotic stress. Plant Methods 2017, 13, 85. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).