Identification, Evolutionary Dynamics, and Gene Expression Patterns of the ACP Gene Family in Responding to Salt Stress in Brassica Genus
Abstract
:1. Introduction
2. Results
2.1. Identification and Characteristics of ACP Genes in the Brassica Genus
2.2. Subcellular Localization Analysis of ACP Family Proteins in B. napus
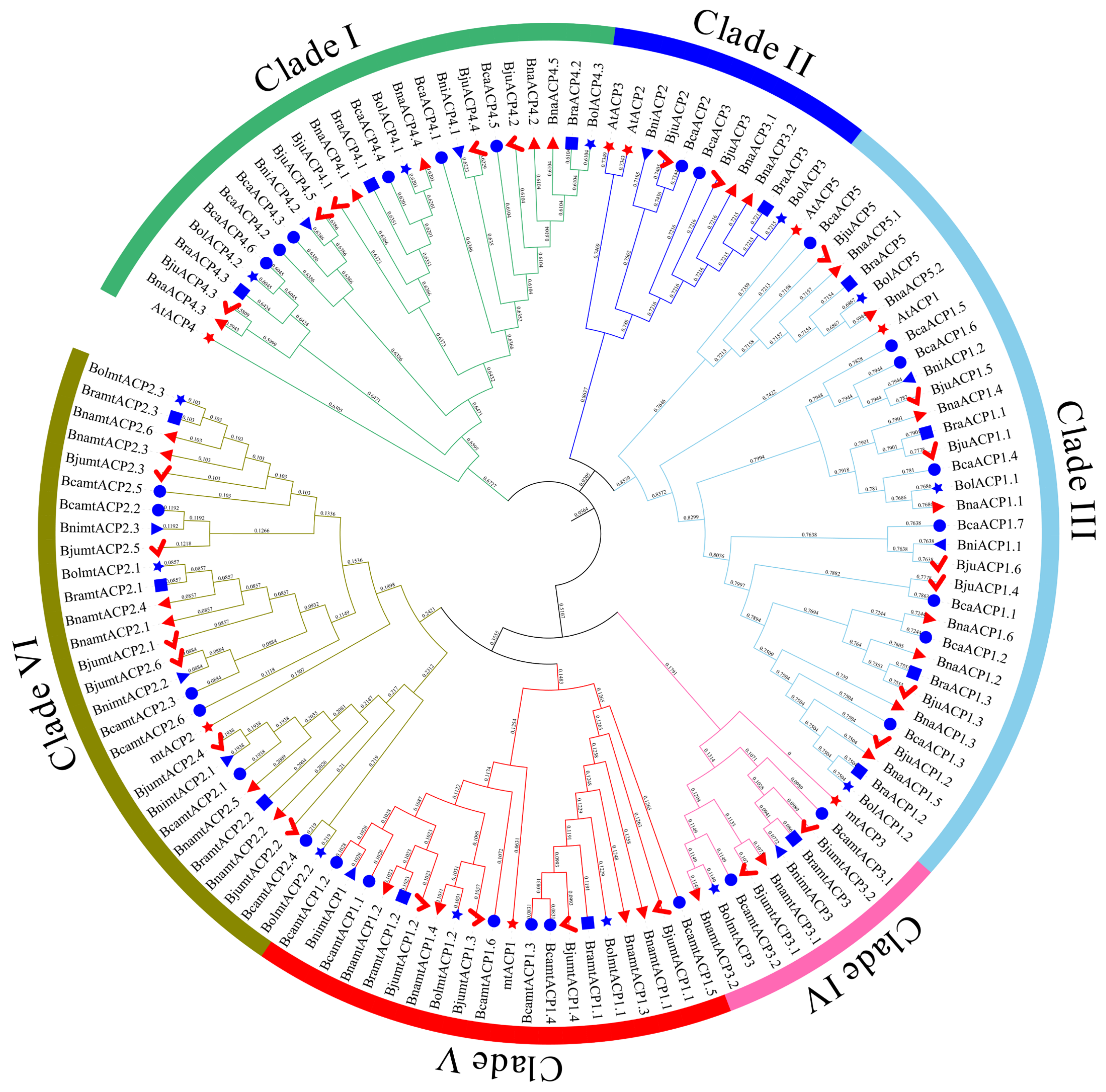
2.3. Phylogenetic Analysis of ACP Proteins
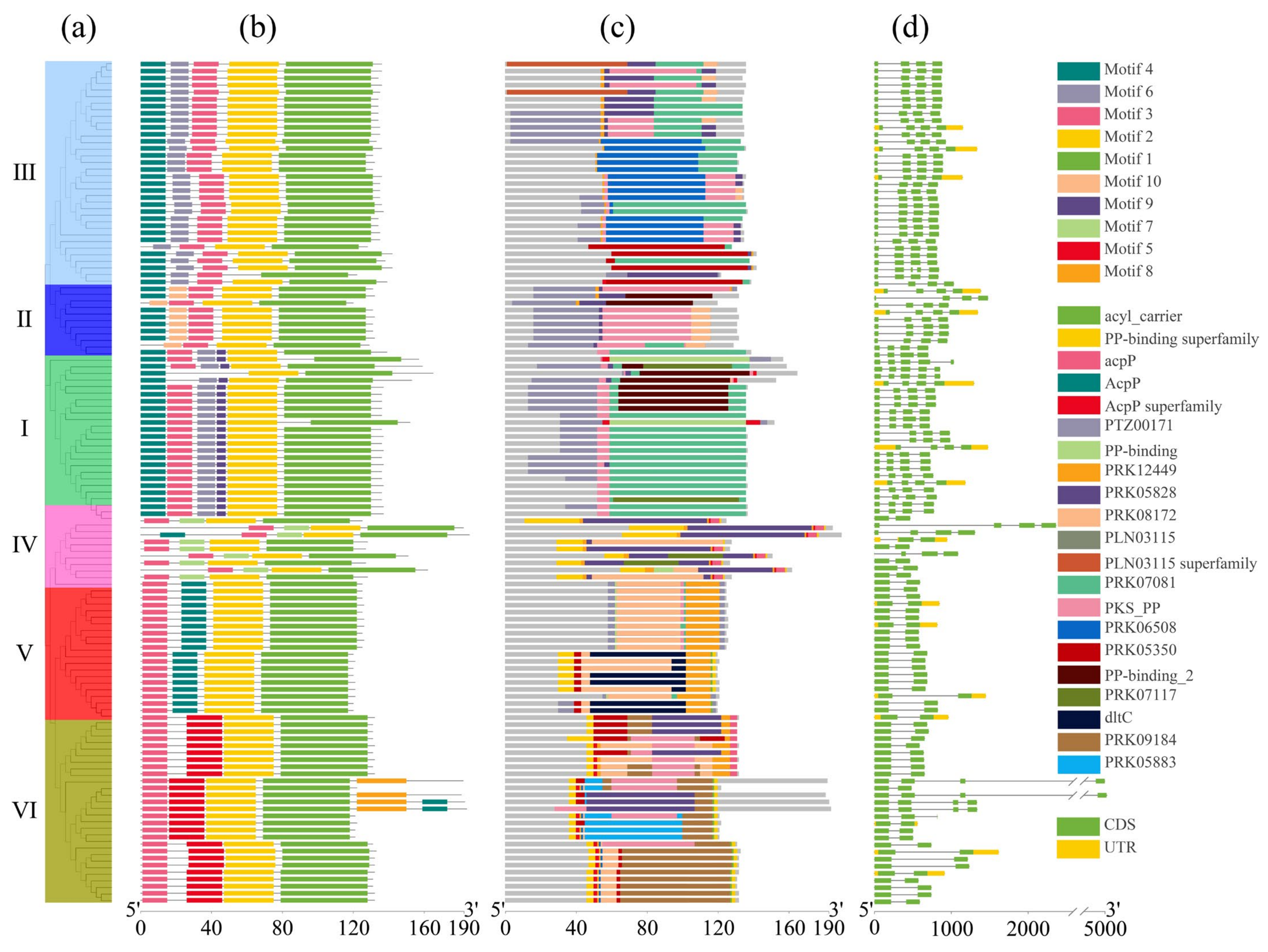
2.4. Conserved Motif, Conserved Domain, and Gene Structure Analysis of ACP Family Proteins
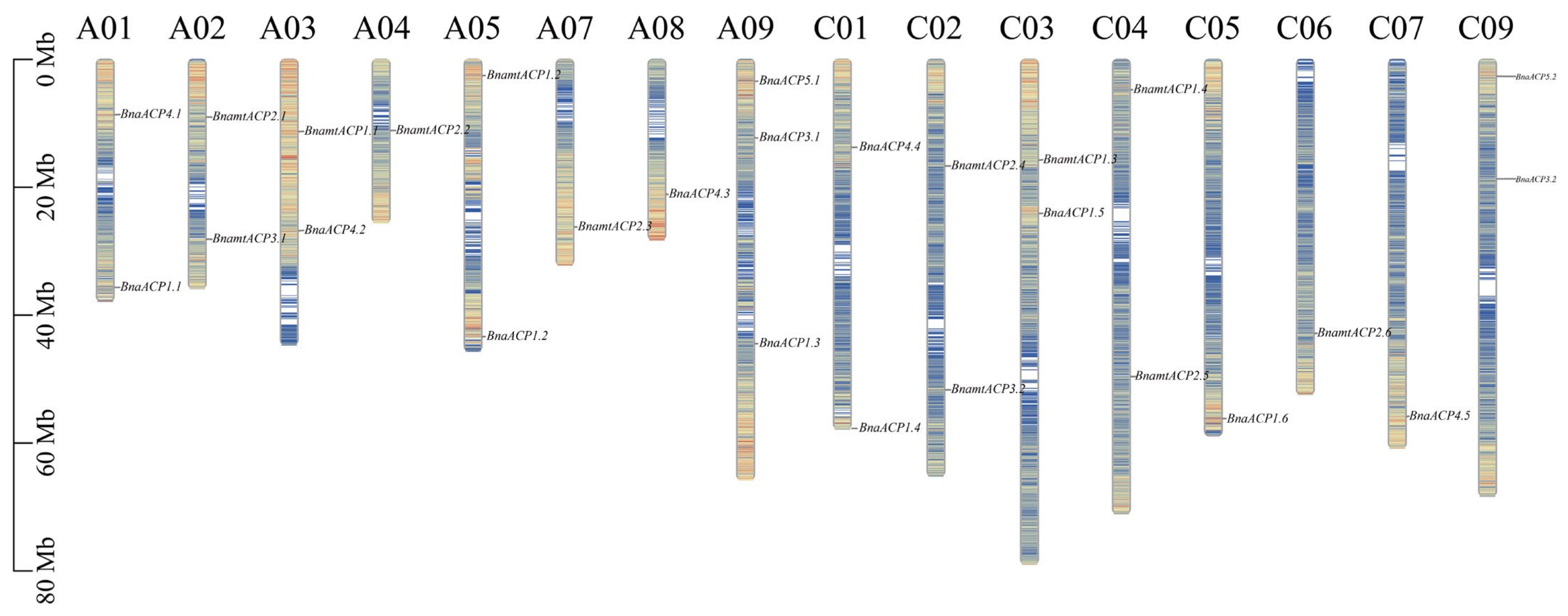
2.5. Chromosomal Localization Analysis of ACP Genes
2.6. Cis-Acting Elements of ACP Genes
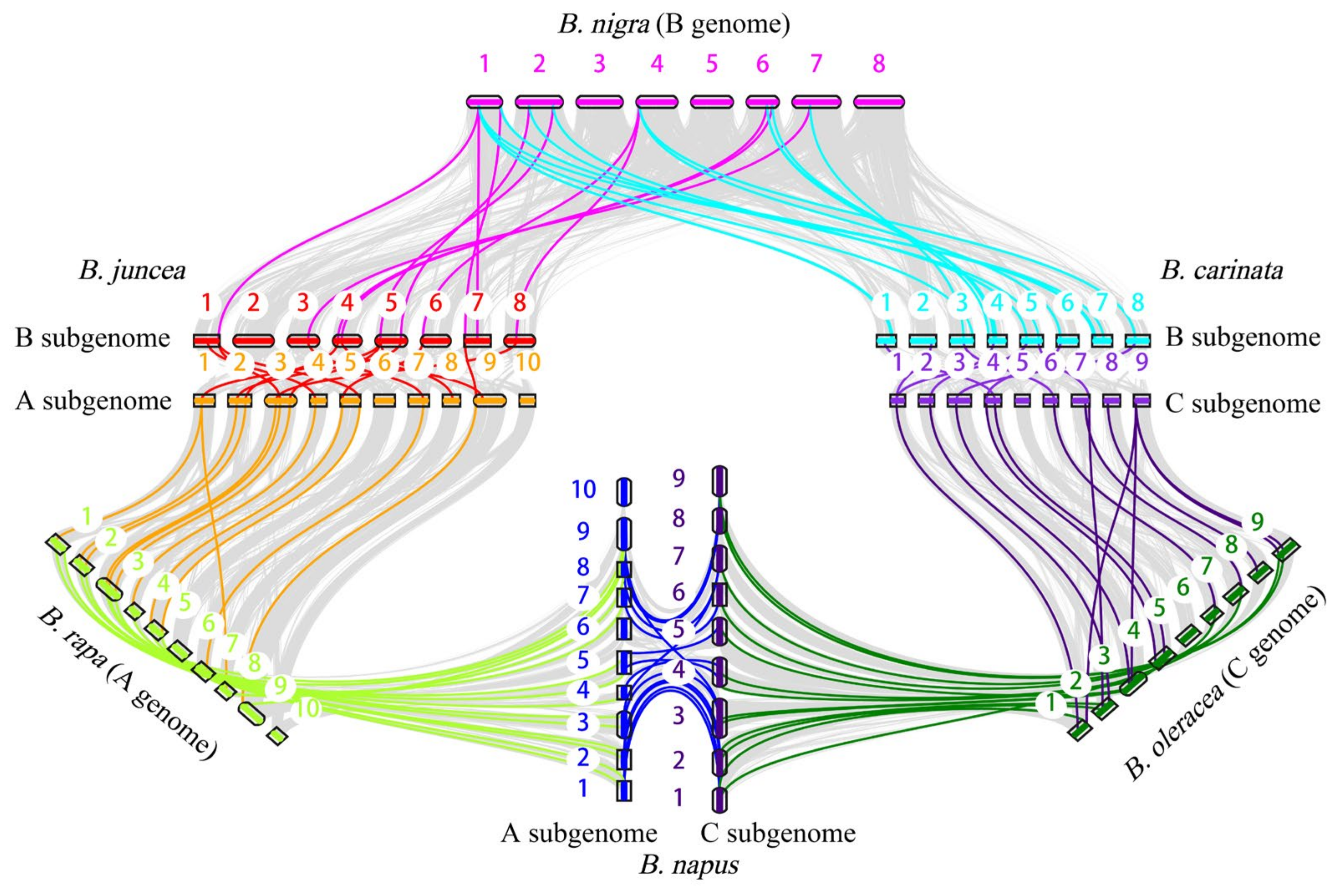
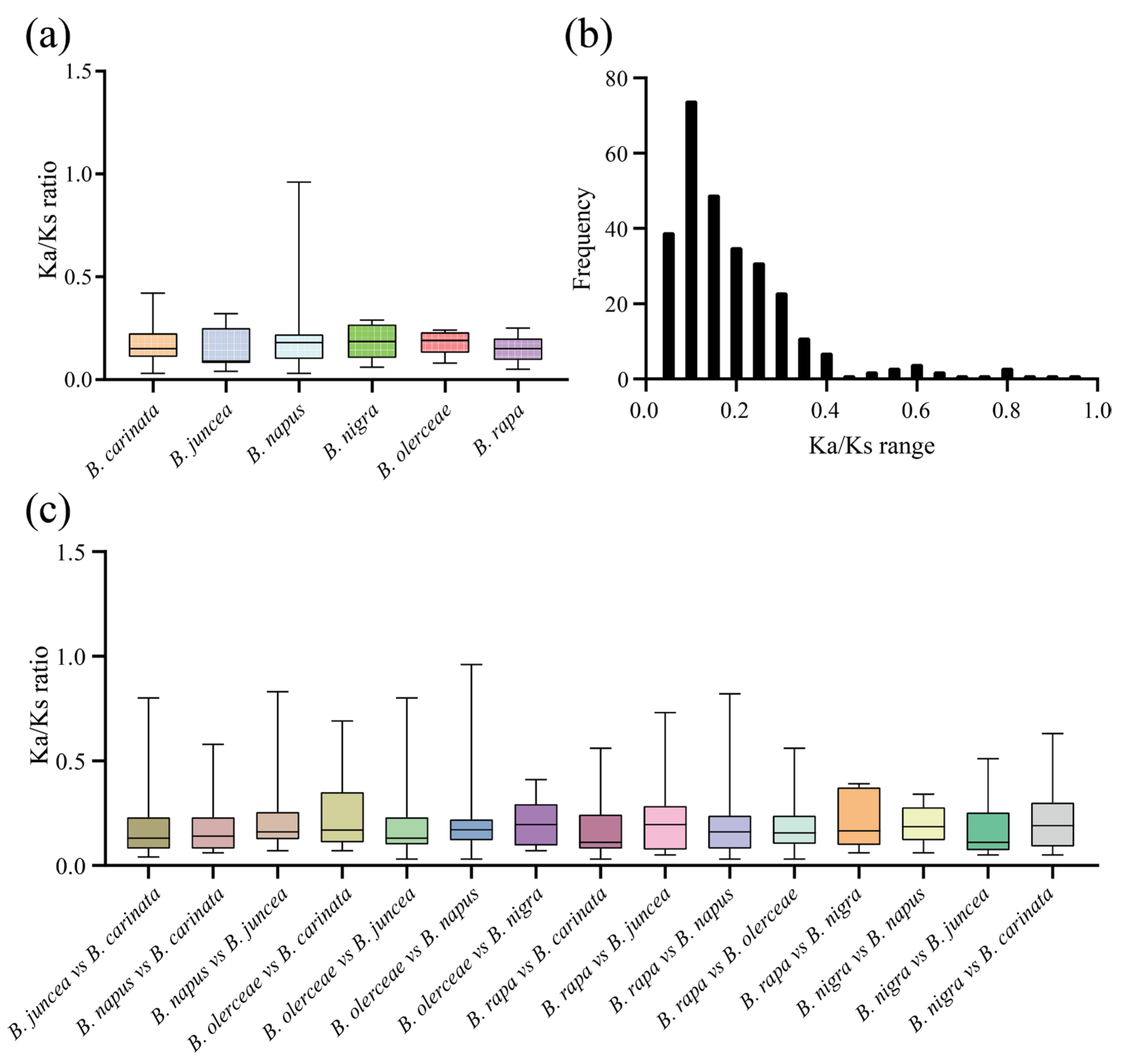
2.7. Gene Duplication, Synteny, and Evolution Analysis of ACP Genes in Six Brassica Species
2.8. Tissue-Specific Expression of ACP Genes
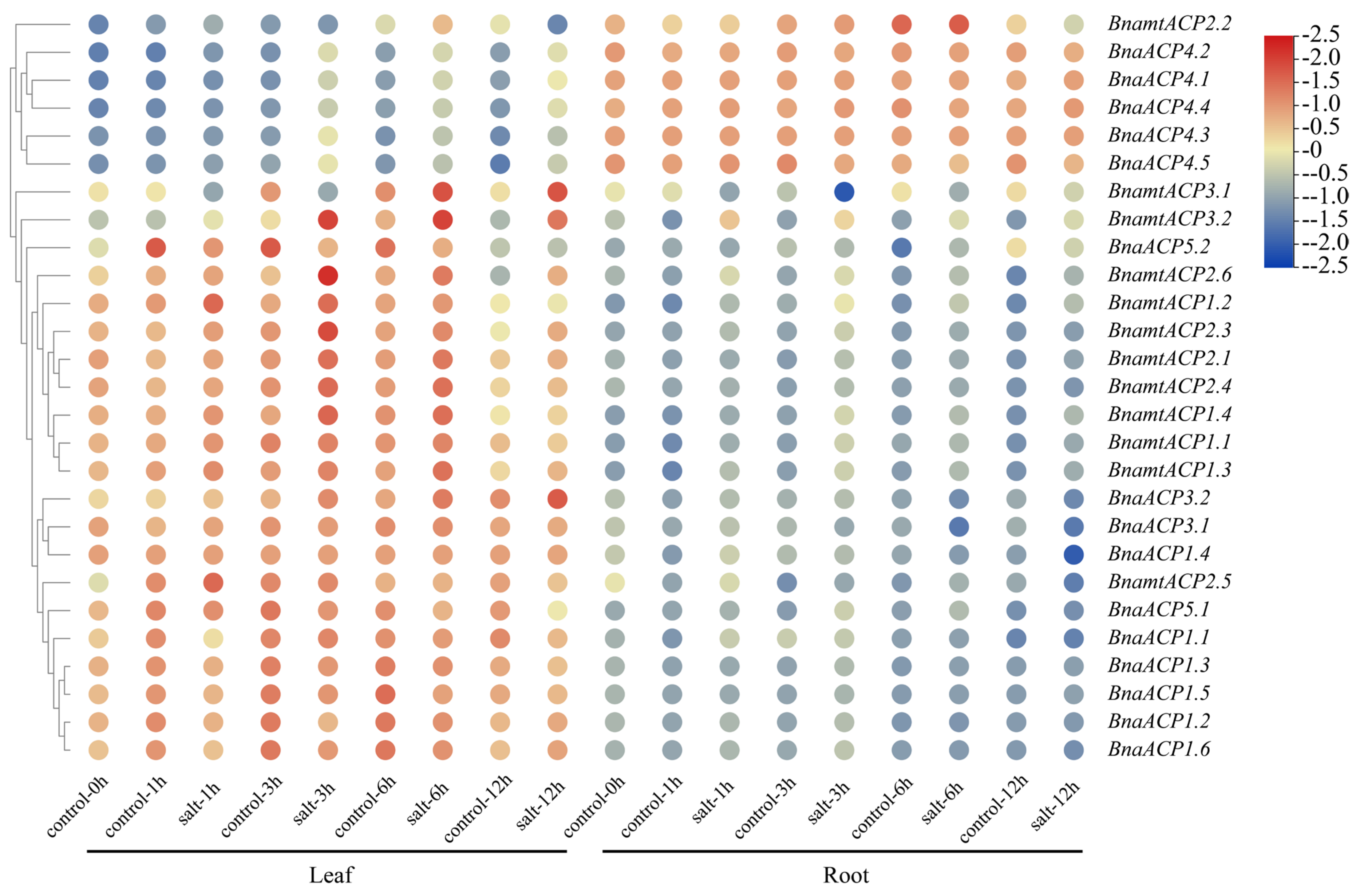
2.9. Expression Patterns of ACP Genes in B. napus under Salts
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Data Sources
4.2. Identification and Characterization of ACP Genes
4.3. Phylogenetic Relationship Analysis
4.4. Conserved Motif and Gene Structure Analysis
4.5. ACP Gene Promoter Sequences and Chromosomal Localization Analysis
4.6. Gene Duplication, Synteny, and Evolution Analysis of ACP Genes
4.7. Analysis of Tissue Expression Characteristics and qRT–PCR
4.8. Subcellular Localization
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hannapel, D.J.; Ohlrogge, J.B. Regulation of acyl carrier protein messenger RNA levels during seed and leaf development. Plant Physiol. 1988, 86, 1174–1178. [Google Scholar] [CrossRef]
- Li, M.J.; Wang, X.J.; Su, L.; Bi, Y.P.; Wan, S.B. Characterization of five putative acyl carrier protein (ACP) isoforms from developing seeds of Arachis hypogaea L. Plant Mol. Biol. Rep. 2010, 28, 365–372. [Google Scholar] [CrossRef]
- Misra, A.; Surolia, N.; Surolia, A. Catalysis and mechanism of malonyl transferase activity in type II fatty acid biosynthesis acyl carrier proteins. Mol. BioSyst. 2009, 5, 651–659. [Google Scholar] [CrossRef]
- Nguyen, C.; Haushalter, R.W.; Lee, D.J.; Markwick, P.R.; Bruegger, J.; Caldara-Festin, G. Trapping the dynamic acyl carrier protein in fatty acid biosynthesis. Nature 2014, 505, 427–431. [Google Scholar] [CrossRef]
- Branen, J.K.; Chiou, T.J.; Engeseth, N.J. Overexpression of acyl carrier protein-1 alters fatty acid composition of leaf tissue in Arabidopsis. Plant Physiol. 2001, 127, 222–229. [Google Scholar] [CrossRef]
- Liu, X.; Hicks, W.M.; Silver, P.A.; Way, J.C. Engineering acyl carrier protein to enhance production of shortened fatty acids. Biotechnol. Biofuels 2016, 9, 1–7. [Google Scholar] [CrossRef]
- Branen, J.K.; Shintani, D.K.; Engeseth, N.J. Expression of antisense acyl carrier protein-4 reduces lipid content in Arabidopsis leaf tissue. Plant Physiol. 2003, 132, 748–756. [Google Scholar] [CrossRef]
- Song, P.; Allen, R.D. Identification of a cotton fiber-specific acyl carrier protein cDNA by differential display. Biochim. Biophys. Acta (BBA)-Gene Struct. Expr. 1997, 1351, 305–312. [Google Scholar] [CrossRef]
- Aluru, M.R.; Mazourek, M.; Landry, L.G.; Curry, J.; Jahn, M.; O’Connell, M.A. Differential expression of fatty acid synthase genes, Acl, Fat and Kas, in Capsicum fruit. J. Exp. Bot. 2003, 54, 1655–1664. [Google Scholar] [CrossRef] [PubMed]
- Silva, J.; Loader, N.M.; Jarman, C.; Windust, J.H.; Hughes, S.G.; Safford, R. The isolation and sequence analysis of two seed-expressed acyl carrier protein genes from Brassica napus. Plant Mol. Biol. 1990, 14, 537–548. [Google Scholar] [CrossRef]
- Suh, M.C.; Schultz, D.J.; Ohlrogge, J.B. Isoforms of acyl carrier protein involved in seed-specific fatty acid synthesis. Plant J. 1999, 17, 679–688. [Google Scholar] [CrossRef] [PubMed]
- Finking, R.; Marahiel, M.A. Biosynthesis of nonribosomal peptides. Ann. Rev. Microbiol. 2004, 58, 453–488. [Google Scholar] [CrossRef]
- Zhang, D.; Song, H.; Cheng, H.; Hao, D.; Wang, H.; Kan, G. The acid phosphatase-encoding gene GmACP1 contributes to soybean tolerance to low-phosphorus stress. PLoS Genet. 2014, 10, e1004061. [Google Scholar] [CrossRef]
- Huang, J.; Xue, C.; Wang, H.; Wang, L.; Schmidt, W.; Shen, R.; Lan, P. Genes of ACYL CARRIER PROTEIN family show different expression profiles and overexpression of ACYL CARRIER PROTEIN 5 modulates fatty acid composition and enhances salt stress tolerance in Arabidopsis. Front. Plant Sci. 2017, 8, 987. [Google Scholar] [CrossRef] [PubMed]
- Nazari, M.; Moosavi, S.S.; Maleki, M.; Jamshidi Goharrizi, K. Chloroplastic acyl carrier protein synthase I and chloroplastic 20 kDa chaperonin proteins are involved in wheat (Triticum aestivum) in response to moisture stress. J. Plant Interact. 2020, 15, 180–187. [Google Scholar] [CrossRef]
- Ge, H.; Xu, J.; Hua, M.; An, W.; Wu, J.; Wang, B. Genome-wide identification and analysis of ACP gene family in Sorghum bicolor (L.) Moench. BMC Genom. 2022, 23, 538. [Google Scholar] [CrossRef]
- Al-Shehbaz, I.A.; Beilstein, M.A.; Kellogg, E.A. Systematics and phylogeny of the Brassicaceae (Cruciferae): An overview. Plant Syst. Evol. 2006, 259, 89–120. [Google Scholar] [CrossRef]
- Warwick, S.I.; Francis, A.; Al-Shehbaz, I.A. Brassicaceae: Species checklist and database on CD-Rom. Plant Syst. Evol. 2006, 259, 249–258. [Google Scholar] [CrossRef]
- Nagaharu, U.; Nagaharu, N. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn. J. Bot. 1935, 7, 389–452. [Google Scholar]
- Wang, X.; Wang, H.; Wang, J.; Sun, R.; Wu, J. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 2011, 43, 1035–1039. [Google Scholar] [CrossRef]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Liu, Y.; Yang, X.; Tong, C.; Edwards, D.; Parkin, I.A. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 2014, 5, 3930. [Google Scholar] [CrossRef]
- Yang, J.; Liu, D.; Wang, X.; Ji, C.; Cheng, F.; Liu, B. The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection. Nat. Genet. 2016, 48, 1225–1232. [Google Scholar] [CrossRef] [PubMed]
- Perumal, S.; Koh, C.S.; Jin, L.; Buchwaldt, M.; Higgins, E.E.; Zheng, C. A high-contiguity Brassica nigra genome localizes active centromeres and defines the ancestral Brassica genome. Nat. Plants 2020, 6, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Yim, W.C.; Swain, M.L.; Ma, D.; An, H.; Bird, K.A.; Curdie, D.D. The final piece of the Triangle of U: Evolution of the tetraploid Brassica carinata genome. Plant Cell 2022, 34, 4143–4172. [Google Scholar] [CrossRef]
- Qiao, X.; Li, Q.; Yin, H.; Qi, K.; Li, L.; Wang, R. Gene duplication and evolution in recurring polyploidization–diploidization cycles in plants. Genome Biol. 2019, 20, 1–23. [Google Scholar] [CrossRef]
- Hurst, L.D. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. 2002, 18, 486–487. [Google Scholar] [CrossRef]
- Lu, K.; Wei, L.; Li, X.; Wang, Y.; Wu, J.; Liu, M. Whole-genome resequencing reveals Brassica napus origin and genetic loci involved in its improvement. Nat. Commun. 2019, 10, 1154. [Google Scholar] [CrossRef]
- Bus, A.; Körber, N.; Snowdon, R.J.; Stich, B. Patterns of molecular variation in a species-wide germplasm set of Brassica Napus. Theor. Appl. Genet. 2001, 123, 1413–1423. [Google Scholar] [CrossRef] [PubMed]
- Chan, D.I.; Vogel, H.J. Current understanding of fatty acid biosynthesis and the acyl carrier protein. Biochem. J. 2010, 430, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Murugan, E.; Kong, R.; Sun, H.; Rao, F.; Liang, Z.X. Expression, purification and characterization of the acyl carrier protein phosphodiesterase from Pseudomonas Aeruginosa. Protein Expres. Purif. 2010, 71, 132–138. [Google Scholar] [CrossRef]
- Bonaventure, G.; Ohlrogge, J.B. Differential regulation of mRNA levels of acyl carrier protein isoforms in Arabidopsis. Plant Physiol. 2002, 128, 223–235. [Google Scholar] [CrossRef]
- Meyer, E.H.; Heazlewood, J.L.; Millar, A.H. Mitochondrial acyl carrier proteins in Arabidopsis thaliana are predominantly soluble matrix proteins and none can be confirmed as subunits of respiratory Complex I. Plant Mol. Biol. 2007, 64, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tóth, K.; Tanaka, K.; Nguyen, C.T.; Yan, Z.; Brechenmacher, L. A soybean acyl carrier protein, GmACP, is important for root nodule symbiosis. Mol. Plant-Microbe Interact. 2014, 27, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Town, C.D.; Cheung, F.; Maiti, R.; Crabtree, J.; Haas, B.J.; Wortman, J.R. Comparative genomics of Brassica oleracea and Arabidopsis thaliana reveal gene loss, fragmentation, and dispersal after polyploidy. Plant Cell 2006, 18, 1348–1359. [Google Scholar] [CrossRef]
- Yang, T.J.; Kim, J.S.; Kwon, S.J.; Lim, K.B.; Choi, B.S.; Kim, J.A. Sequence-level analysis of the diploidization process in the triplicated FLOWERING LOCUS C region of Brassica rapa. Plant Cell 2006, 18, 1339–1347. [Google Scholar] [CrossRef]
- Blanc, G.; Hokamp, K.; Wolfe, K.H. A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res. 2003, 13, 137–144. [Google Scholar] [CrossRef] [PubMed]
- Saha, G.; Park, J.I.; Jung, H.J.; Ahmed, N.U.; Kayum, M.A.; Chung, M.Y. Genome-wide identification and characterization of MADS-box family genes related to organ development and stress resistance in Brassica rapa. BMC Genom. 2015, 16, 178. [Google Scholar] [CrossRef]
- Tang, J.; Wang, F.; Hou, X.L.; Wang, Z.; Huang, Z.N. Genome-wide fractionation and identification of WRKY transcription factors in Chinese cabbage (Brassica rapa ssp. pekinensis) reveals collinearity and their expression patterns under abiotic and biotic stresses. Plant Mol. Biol. Rep. 2014, 32, 781–795. [Google Scholar] [CrossRef]
- Chen, D.; Chen, H.; Dai, G.; Zhang, H.; Liu, Y.; Shen, W. Genome-wide identification of R2R3-MYB gene family and association with anthocyanin biosynthesis in Brassica species. BMC Genom. 2022, 23, 441. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Q.; Zhao, Y.; Han, G.; Zhu, S. Systematic analysis of maize class III peroxidase gene family reveals a conserved subfamily involved in abiotic stress response. Gene 2015, 566, 95–108. [Google Scholar] [CrossRef]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jin, Q.; Lin, Y.; Cai, Y. Structural, evolutionary, and functional analysis of the class III peroxidase gene family in Chinese pear (Pyrus bretschneideri). Front. Plant Sci. 2016, 7, 1874. [Google Scholar] [CrossRef]
- Force, A.; Lynch, M.; Pickett, F.B.; Amores, A.; Yan, Y.L.; Postlethwait, J. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 1999, 151, 1531–1545. [Google Scholar] [CrossRef]
- Smith, S. Architectural options for a fatty acid synthase. Science 2006, 311, 1251–1252. [Google Scholar] [CrossRef] [PubMed]
- Byers, D.M.; Gong, H. Acyl carrier protein: Structure–function relationships in a conserved multifunctional protein family. Biochem. Cell Biol. 2007, 85, 649–662. [Google Scholar] [CrossRef] [PubMed]
- Jha, J.K.; Sinha, S.; Maiti, M.K.; Basu, A.; Mukhopadhyay, U.K.; Sen, S.K. Functional expression of an acyl carrier protein (ACP) from Azospirillum brasilense alters fatty acid profiles in Escherichia coli and Brassica juncea. Plant Physiol. Biochem. 2007, 45, 490–500. [Google Scholar] [CrossRef]
- Yang, X.; Liu, X.; Zhou, Y.; Zhang, F.; Huang, L.; Wang, J. New insights on the function of plant acyl carrier proteins from comparative and evolutionary analysis. Genomics 2021, 113, 1155–1165. [Google Scholar] [CrossRef] [PubMed]
- Zhao, K.; Chen, S.; Yao, W.; Cheng, Z.; Zhou, B.; Jiang, T. Genome-wide analysis and expression profile of the bZIP gene family in poplar. BMC Plant Biol. 2021, 21, 122. [Google Scholar] [CrossRef]
- Li, X.; Zhang, X.; Shi, T.; Chen, M.; Jia, C.; Wang, J. Identification of ARF family in blueberry and its potential involvement of fruit development and pH stress response. BMC Genom. 2022, 23, 329. [Google Scholar] [CrossRef]
- Lang, L.; Xu, A.; Ding, J.; Zhang, Y.; Zhao, N.; Tian, Z.; Liu, Y. Quantitative trait locus mapping of salt tolerance and identification of salt-tolerant genes in Brassica napus L. Front. Plant Sci. 2017, 8, 1000. [Google Scholar] [CrossRef]
- Raees, N.; Ullah, S.; Nafees, M. Interactive effect of tocopherol, salicylic acid and ascorbic acid on agronomic characters of two genotypes of Brassica napus L. under induced drought and salinity stresses. Gesunde Pflanz. 2022, 75, 1905–1923. [Google Scholar] [CrossRef]
- Chen, H.; Wang, T.; He, X.; Cai, X.; Lin, R.; Liang, J. BRAD V3.0: An upgraded Brassicaceae database. Nucleic Acids Res. 2022, 50, D1432–D1441. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Wang, S.; Wei, L.; Huang, Y.; Liu, D.; Jia, Y. BnIR: A multi-omics database with various tools for Brassica napus research and breeding. Mol. Plant 2023, 16, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 2013, 41, e121. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Shen, H.B. Cell-PLoc: A package of Web servers for predicting subcellular localization of proteins in various organisms. Nat. Protoc. 2008, 3, 153–162. [Google Scholar] [CrossRef]
- Edgar, R.C. Muscle5: High-accuracy alignment ensembles enable unbiased assessments of sequence homology and phylogeny. Nat. Commun. 2022, 13, 6968. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Goll, J.; Rusch, D.B.; Tanenbaum, D.M.; Thiagarajan, M.; Li, K.; Methé, B.A.; Yooseph, S. METAREP: JCVI metagenomics reports—An open source tool for high-performance comparative metagenomics. Bioinformatics 2010, 26, 2631–2632. [Google Scholar] [CrossRef] [PubMed]
- Yan, G.; Lv, X.; Gao, G.; Li, F.; Li, J.; Qiao, J. Identification and characterization of a glyoxalase I gene in a rapeseed cultivar with seed thermotolerance. Front. Plant Sci. 2016, 7, 150. [Google Scholar] [CrossRef]
- Gan, C.; Liu, Z.; Pang, B.; Zuo, D.; Hou, Y.; Zhou, L. Integrative physiological and transcriptome analyses provide insights into the Cadmium (Cd) tolerance of a Cd accumulator: Erigeron canadensis. BMC Genom. 2022, 23, 778. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qian, F.; Zuo, D.; Zeng, T.; Gu, L.; Wang, H.; Du, X.; Zhu, B.; Ou, J. Identification, Evolutionary Dynamics, and Gene Expression Patterns of the ACP Gene Family in Responding to Salt Stress in Brassica Genus. Plants 2024, 13, 950. https://doi.org/10.3390/plants13070950
Qian F, Zuo D, Zeng T, Gu L, Wang H, Du X, Zhu B, Ou J. Identification, Evolutionary Dynamics, and Gene Expression Patterns of the ACP Gene Family in Responding to Salt Stress in Brassica Genus. Plants. 2024; 13(7):950. https://doi.org/10.3390/plants13070950
Chicago/Turabian StyleQian, Fang, Dan Zuo, Tuo Zeng, Lei Gu, Hongcheng Wang, Xuye Du, Bin Zhu, and Jing Ou. 2024. "Identification, Evolutionary Dynamics, and Gene Expression Patterns of the ACP Gene Family in Responding to Salt Stress in Brassica Genus" Plants 13, no. 7: 950. https://doi.org/10.3390/plants13070950





