Abstract
Species of the genus Microdochium (Microdochiaceae, Xylariales) have been reported from the whole world and separated from multiple plant hosts. The primary aim of the present study is to describe and illustrate three new species isolated from the leaf spot of Bambusaceae sp. and saprophytic leaves in Hainan and Yunnan provinces, China. The proposed three species, viz., Microdochium bambusae, M. nannuoshanense and M. phyllosaprophyticum, are based on multi-locus phylogenies from a combined dataset of ITS rDNA, LSU, RPB2 and TUB2 in conjunction with morphological characteristics. Descriptions and illustrations of three new species in the genus are provided.
1. Introduction
Microdochium Syd. & P. Syd., belonging to Microdochiaceae Hern.-Restr., Crous & J.Z. Groenew., was introduced by Syd. & P. Syd. in 1924 [1]. Microdochium phragmitis Syd. & P. Syd., the genus type, was introduced by Sydow on leaves of Phragmites australis in Germany in 1919 [1]. Microdochium species had frequently been separated into endophytes, plant pathogens and saprophytes, and were commonly isolated from some diseased plant hosts [2,3,4,5,6,7]. At present, about 62 species of Microdochium are listed in the Index Fungorum (http://www.indexfungorum.org/, accessed on 30 October 2023). Species of Microdochium are characterized by coelomycetous asexual morphs producing polyblastic, sympodial or annellidic conidiogenous cells with hyaline conidia. Sexual morphs are Monographella-like with present or absent stromata, perithecial ascomata, eight-spored, oblong, clavate asci, apical ring, and hyaline to pale brown, fusoid ellipsoid or oblong ascospores [8].
Previous studies have shown that Microdochium belongs to Amphisphaeriaceae, based on its morphological resemblance [9,10,11,12]. Hernández-Restrepo et al. [8] suggested that Idriella and Microdochium could be congeneric. In their phylogenetic analysis, Idriella, Microdochium and Selenodriella clustered in Xylariales as a distinct monophyletic lineage. Therefore, Hernández-Restrepo et al. [8] introduced the new family Microdochiaceae to accommodate this clade. The hosts of Microdochium are diverse and widely distributed [8,13,14,15,16,17,18]. In recent years, Microdochium has included important plant pathogens; for example, Kwasna et al. [19] gave an up-to-date description of Microdochium triticicola, which was split from the roots of Triticum aestivum L. in the UK. Zhang et al. [20] identified Microdochium paspali, which caused leaf blight of Paspalum vaginatum Sw., a widely used lawn grass for tropical and subtropical golf courses. Liu et al. [21] described three species, Microdochium miscanthi, M. sinense and M. hainanense, isolated from Miscanthus sinensis Anderss. and Phragmites australis (Cav.) Trin. ex Steud in Hainan, China. Dissanayake et al. [22] described M. sichuanense isolated from a Poaceae host in Sichuan, China. Liang et al. [2] described new species Microdochium poae, which caused leaf blight of Creeping bentgrasses (Agrostis stolonifera L.) and Kentucky blue grass (Poa pratensis L. var. anceps Gaud.). In particular, Mandyam et al. [5] sporulated dark septate endophytes from roots of mixed tallgrass prairie plant communities, and sporulated species of Aspergillus, Fusarium, Microdochium and Periconia by ITS-RFLP and/or sequencing of the internal transcribed spacer of the ribosomal RNA gene (ITS) region. In addition, Microdochium produced abundant melanized inter- and intracellular chlamydospores.
Fungi associated with leaf spots were collected from Bambusaceae sp. and saprophytic leaves. Morphological characteristics were obtained by separation and purification. The sequences of four molecular markers, viz., the partial nuclear ribosomal large subunit (LSU), the ITS gene, the partial RNA polymerase II second-largest subunit (RPB2) and the partial β-tubulin gene (TUB2), were used in this study. We identify these fungi as three species of the genus Microdochium, proposed herein.
2. Materials and Methods
2.1. Sample Collection, Fungal Isolation and Morphology
Bambusaceae sp. and saprophytic specimens showing necrotic spots were collected during a series of field visits in Hainan and Yunnan provinces in China in 2022–2023. A specimen usually isolates multiple fungi, and we obtained a single colony through single spore isolation and tissue isolation methods [23]. We removed fragments (4 × 4 mm) from the damaged edges of the leaves and disinfected the surface by continuously soaking them in 75% ethanol solution and rinsing them in sterile distilled water and 10% sodium hypochlorite solution, and then we rinsed them three times in sterile deionized water for 60 s, 45 s, 45 s and 30 s, respectively. We placed the processed fragments on sterile filter paper to absorb moisture, and then placed them on Potato Dextrose Agar (PDA potato: 200 g, dextrose: 15 g, agar: 15 g, distilled water 1 L and natural pH) or Oat Meal Agar (OA: oats: 30 g, agar: 15 g, distilled water: 1 L and natural pH) and incubated them at 24 °C for 3–5 days. Then, the agar portion containing fungal hyphae from the periphery of the colony was transferred to a new PDA plate and was photographed using a Sony ZV-E10L digital camera (Sony Group Corporation, Tokyo, Japan) on days 5, 10 and 15. Using an Olympus SZ61 stereo microscope and an Olympus BX43 microscope (Olympus Corporation, Tokyo, Japan), respectively, in conjunction with an Olympus DP73 and OPTIKA SC2000 high-definition color digital camera, the microscopic morphological characteristics of the structures produced in the culture were observed to capture and record fungal structures. All fungal strains were stored in 15% sterilized glycerol at 4 °C, with each strain stored in three tubes (2.0 mL tubes), for further research. Digimizer software (v5.6.0) was used for structural measurements, with 25 or more measurements taken for each character (conidiophores, conidiogenous cells, conidia and so on) [23]. Specimens were deposited in the HSAUP and HMAS [23]. Living cultures were deposited in the SAUCC [23].
2.2. DNA Extraction, PCR and Sequencing
Fungal DNA was extracted from the fresh mycelia grown on PDA or OA using a CTAB (cetyltrimethylammonium bromide) method and a kit method (OGPLF-400, GeneOnBio Corporation, Changchun, China) [24,25]. Four molecular markers, including LSU, ITS, RPB2 and TUB2 gene, were amplified with the primer pairs listed in Table 1. An amplification reaction was carried out at 20 μL reaction volume, including 10 μL 2 × Hieff Canace® Plus PCR Master Mix (Shanghai, China) (with dye) (Yeasen Biotechnology, Shanghai, China, Cat No. 10154ES03), 0.5 μL each of forward and reverse primer, and 0.5 μL template genomic DNA, adjusted to a total volume of 20 μL using distilled deionized water. Some 1% agarose gel and GelRed (TsingKe, Qingdao, China) were used to separate and purify the PCR product, and ultraviolet light was used to observed whether the fragment was consistent. Then a Gel Extraction Kit (Cat: AE0101-C) (Shandong Sparkjade Biotechnology Co., Ltd., Jinan, China) was used for gel recovery. The PCR products were processed for purification and bidirectional sequencing by TsingKe Biological Technology, Qingdao, China. The raw data (trace data) were processed using MEGA v. 7.0 to obtained consistent sequences, including removing disordered peak sequences and complementary concatenation of forward and reverse sequences (ClustalW) [26]. All sequences generated in this study were deposited in GenBank under the accession numbers in Table 2.

Table 1.
The PCR primers, sequence and cycles used in this study.

Table 2.
GenBank accession numbers of the taxa used in phylogenetic reconstruction.
2.3. Phylogeny
The newly generated sequences in this study and closely associated sequences from Liu et al. [21] and Dissanayake et al. [22] were aligned using the MAFFT 7 online service with the default strategy and corrected manually using MEGA 7 [26,31]. To determine the identity of the isolates at species level, phylogenetic analysis was first conducted separately for each marker, followed by a combination (ITS-LSU-RPB2-TUB2) (See Supplementary File S1).
Phylogenetic analysis of multi-labeled data was based on Bayesian inference (BI) and maximum likelihood (ML) algorithms. Firstly, MrModeltest v. 2.3 [32] was used determine the best evolutionary model for each partition under the Akaike Information Criterion (AIC), which was used to identified the best nucleotide substitution model settings prior to the BI analysis. Secondly, ML and BI were run on the CIPRES Science Gateway portal (https://www.phylo.org/, accessed on 30 October 2023) or offline software (ML was operated in RaxML-HPC2 on XSEDE v8.2.12, and BI analysis was operated in MrBayes v3.2.7a with 64 threads on Linux) [33,34,35,36,37]. Thirdly, for ML analyses, the default parameters were used and 1000 rapid bootstrap replicates were run with the GTR+G+I model of nucleotide evolution; BI analysis was performed using a fast bootstrap algorithm with an automatic stop option. Finally, all resulted trees were plotted using FigTree v. 1.4.4 (http://tree.bio.ed.ac.uk/software/figtree, accessed on 20 October 2023) or ITOL: Interactive Tree of Life (https://itol.embl.de/, accessed on 20 October 2023) [38], and the layout of the trees was produced in Adobe Illustrator CC 2019.
3. Results
3.1. Phylogenetic Analyses
The comparison contained 58 isolates representing Microdochium and related taxa, and the used strains CBS 204.56 and CBS 177.57 of Idriella lunata were used as an outgroup. The final alignments consisting of 2945 characters were used for phylogenetic analyses, viz., 1–581 (ITS), 582–1418 (LSU), 1419–2258 (RPB2) and 2259–2945 (TUB2), including gaps. Of these characters, 2213 were constant, 68 were variable and parsimony-uninformative, and 664 were parsimony-informative. The topology of the ML tree was consistent with that of the Bayesian tree and was therefore considered to be representative of the evolutionary history of the genus Microdochium (Figure 1). The final ML optimization likelihood was −16,911.019760. The matrix had 681 distinct alignment patterns with 15.92% undetermined characters or gaps. Estimated base frequencies were as follows: A = 0.259370, C = 0.234527, G = 0.263997 and T = 0.242106; substitution rates were AC = 0.945501, AG = 4.719533, AT = 1.249092, CG = 0.946009, CT = 7.264699 and GT = 1.000000; and the gamma distribution shape parameter α = 0.124118. The Dirichlet base frequencies and the GTR+I+G evolutionary mode were used for ITS, K80+I+G was used for LSU, HKY+I+G was used for RPB2, and GTR+I was used for TUB2. MCMC analysis of these four tandem genes was performed over 245,000 generations in 4902 trees. The first 1224 trees representing the aging phase of the analysis were discarded, while the remaining trees were used to calculate the posterior probability in the majority-rule consensus tree (Figure 1; first value: PP > 0.80 shown). The alignment embodied a total of 848 unique site patterns (ITS: 237, LSU: 94, RPB2: 350 and TUB2: 167).
The 58 strains were assigned to 37 species clades on the phylogram (Figure 1). Among them, six strains conducted three new species lineages: Microdochium bambusae (SAUCC 1862-1, SAUCC 1866-1) was closely related to M. indocalami (SAUCC 1016) with good support (1.0 BIPP and 90% MLBV), M. nannuoshanense (SAUCC 2450-1 and SAUCC 2450-3) was closely related to M. sinense (SAUCC 211097 and SAUCC 211098) with full support (1.0 BIPP and 99% MLBV), and M. phyllosaprophyticum (SAUCC 3583-1 and SAUCC 3583-6) formed a separate single-species lineage. Based on the phylogenetic resolution and morphological analyses, three new species of the Microdochium species, viz., Microdochium bambusae sp. Nov., M. nannuoshanense sp. Nov. and M. phyllosaprophyticum sp. nov., were reported in the present study.

Figure 1.
A maximum likelihood tree based on combined dataset of analyzed ITS, LSU, RPB2 and TUB2 sequence. Left, BIPP ≥ 0.80 and right, MLBV ≥ 70% are shown as BIPP/ML above the nodes. The branches BIPP/ML with 1/100 are indicated in bold. Ex-type cultures are indicated in bold face and marked with “T”. Strains from the present study are in red. The tree was rooted in Idriela lunata (CBS 204.56* and CBS 177.57). The yellow and green areas are used to distinguish different species. The scale bar at the bottom middle indicates 0.05 substitutions per site.
Figure 1.
A maximum likelihood tree based on combined dataset of analyzed ITS, LSU, RPB2 and TUB2 sequence. Left, BIPP ≥ 0.80 and right, MLBV ≥ 70% are shown as BIPP/ML above the nodes. The branches BIPP/ML with 1/100 are indicated in bold. Ex-type cultures are indicated in bold face and marked with “T”. Strains from the present study are in red. The tree was rooted in Idriela lunata (CBS 204.56* and CBS 177.57). The yellow and green areas are used to distinguish different species. The scale bar at the bottom middle indicates 0.05 substitutions per site.

3.2. Taxonomy
3.2.1. Microdochium bambusae J. Zhang, Z.X. Zhang, & Z. Li, sp. Nov.; Figure 2
MycoBank—No. MB850598
Etymology—Referring to the generic name of the host plant Bambusaceae sp.
Type—China, Yunnan Province: Xishuangbanna Tropical Botanical Garden, Chinese Academy of Sciences; on diseased leaves of Bambusaceae sp.; 15 March 2023; Z. X. Zhang (HMAS 352651, holotype); ex-holotype living culture SAUCC 1862-1.
Description—Endogenic on diseased leaves of Bambusaceae sp. Sexual morphs are unknown. Mycelia are superficial and immersed, 2.3–3.6 µm wide, branched, membranous and hyaline. Conidiophores are inapparent and often reduced to conidiogenous cells. Conidiogenous cells are straight or slightly curved, 17.4–30.0 × 2.5–3.0 µm, mono- or polyblastic, terminal, hyaline, septate, cylindrical and smooth, and produced on aerial mycelia. Conidia are solitary, hyaline, oblong to ellipsoid, straight or curved, 13.0–17.0 × 2.5–3.5 µm, multi-guttulate and sometimes borne directly from hyphae. Chlamydospores were not observed; see Figure 2.
Culture characteristics—Cultures incubated on PDA at 25 °C in darkness, reaching 55–60 mm diam., had a growth rate of 3.9–4.3 mm/day after 14 days, with moderate aerial mycelia, were gray white with regular margins, and the reverses were dark brown in the center and light brown to white at the edge. Cultures incubated on PDA at 25 °C in darkness, reaching 82–87 mm diam., had a growth rate of 5.8–6.2 mm/day after 14 days, with moderate aerial mycelia on the surface, were gray white with regular margins, and the reverses were similar in color.
Additional specimen examined—China, Yunnan Province: Xishuangbanna Tropical Botanical Garden, Chinese Academy of Sciences; on diseased leaves of Bambusaceae sp.; 15 March 2023; J. Zhang (HSAUP 1866-1); living culture SAUCC 1866-1.
Notes—Phylogenetic analyses of four combined genes (ITS, LSU, RPB2 and TUB2) showed that Microdochium bambusae sp. nov. formed an independent clade closely related to M. indocalami. M. bambusae is distinguished from M. indocalami (SAUCC 1016) by 7/519, 3/836, 59/840 and 19/715 characters in ITS, LSU, RPB2 and TUB2 sequences, respectively. Morphologically, M. bambusae differs from M. indocalami in conidia (13.0–17.0 × 2.5–3.5 μm vs. 13.0–15.5 × 3.5–5.5 μm) and in colony texture (gray white with regular margins vs. white with regular margins on PDA), and M. indocalami has more aerial mycelia than M. bambusae [17]. For details, see Table 3.
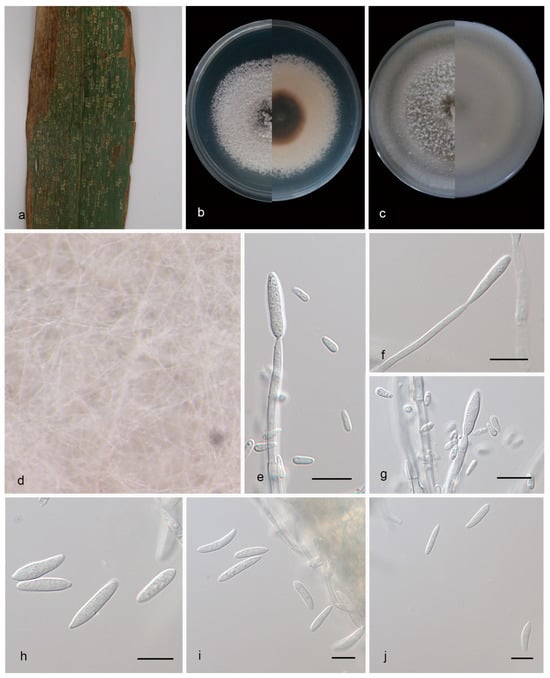
Figure 2.
Microdochium bambusae (HMAS 352651, holotype). (a) A leaf of Bambusaceae sp.; (b) colonies on PDA from above and below after 14 days; (c) colonies on OA from above and below after 14 days; (d) colony overview; (e–g) conidiogenous cells and conidia; (h–j) conidia. Scale bars: (e–j) 10 μm.
Figure 2.
Microdochium bambusae (HMAS 352651, holotype). (a) A leaf of Bambusaceae sp.; (b) colonies on PDA from above and below after 14 days; (c) colonies on OA from above and below after 14 days; (d) colony overview; (e–g) conidiogenous cells and conidia; (h–j) conidia. Scale bars: (e–j) 10 μm.
3.2.2. Microdochium nannuoshanense J. Zhang, Z.X. Zhang, & Z. Li, sp. Nov.; Figure 3
MycoBank—No. MB850597
Etymology—Referring to the location of the holotype, Nannuoshan, Yunnan Province, China.
Type—China, Yunnan Province, Nannuoshan; on diseased leaves of Bambusaceae sp.; 12 April 2023; J. Zhang (HMAS 352652, holotype); ex-holotype living culture SAUCC 2450-1.
Description—Endogenic on diseased leaves of Bambusaceae sp. Sexual morphs are unknown. Mycelia are superficial and immersed, 1.8–2.6 µm wide, branched, membranous and hyaline. Conidiophores are inapparent and often reduced to conidiogenous cells. Conidiogenous cells are straight or slightly curved, 19.0–27.0 × 2.0–3.0 µm, mono- or polyblastic, terminal, denticulate, transparent, smooth, cylindrical and septate, and produced on aerial mycelia. Conidia are solitary, hyaline, spindle to rod-shaped, straight or curved, oblong to ellipsoid, 7.0–12.0 × 2.5–4.0 µm, multi-guttulate and sometimes borne directly from the hyphae. Chlamydospores were not observed; see Figure 3.
Culture characteristics—Cultures incubated on PDA at 25 °C in darkness, reaching 68–73 mm diam., had a growth rate of 4.8–5.2 mm/day after 14 days, were creamy white to pale brown with regular margins, had moderate aerial mycelia, and the reverse was similar. Cultures incubated on OA at 25 °C in darkness, reaching 53–57 mm diam., with a growth rate of 3.7–4.1 mm/day, were creamy white with regular margins, had luxuriant aerial hyphae, and the reverse was similar.
Additional specimen examined—China, Yunnan, Nannuoshan; on diseased leaves of Bambusaceae sp.; 12 April 2023; J. Zhang; HSAUP2450-3; living culture SAUCC 2450-3.
Notes—Phylogenetic analyses of four combined genes (ITS, LSU, RPB2 and TUB2) showed that Microdochium nannuoshanense sp. nov. formed an independent clade closely related to M. sinense. M. nannuoshanense is distinguished from M. sinense (SAUCC 211097) by 4/535, 3/836, 16/909 and 4/717 characters in ITS, LSU, RPB2 and TUB2 sequences, respectively. Morphologically, M. nannuoshanense differs from M. sinense in conidia (7.0–12.0 × 2.5–4.0 μm vs. 11.5–19.34 × 2.8–5.4 μm) and in colony texture (creamy white to pale brown with regular margins vs. yellow-brown overall and fluffy at the edge on PDA) [21]. For details, see Table 3.
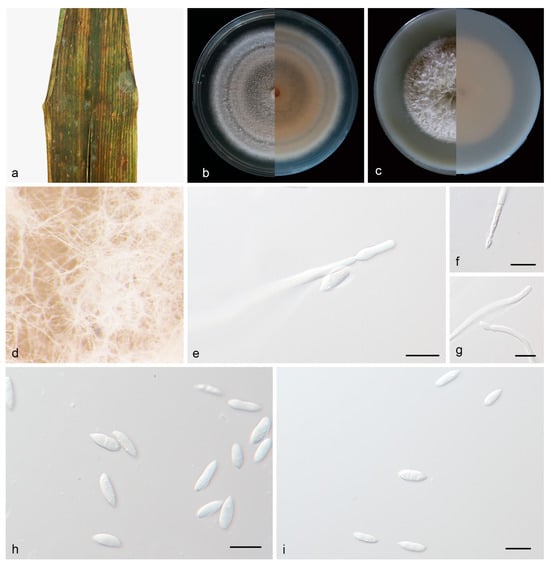
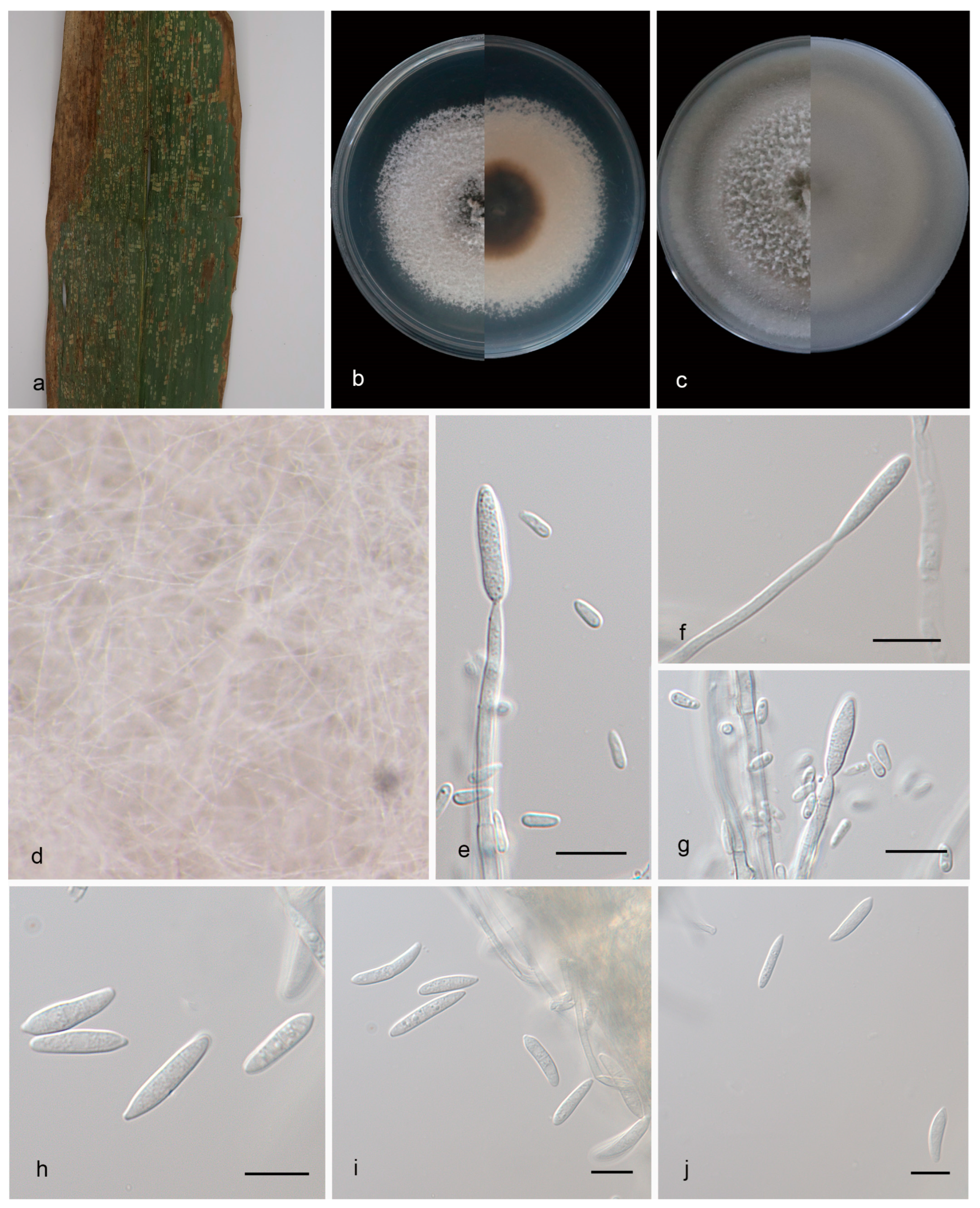
Figure 3.
Microdochium nannuoshanense (holotype, HMAS 352652). (a) A leaf of Bambusaceae sp; (b) colonies on PDA from above and below after 14 days; (c) colonies on OA from above and below after 14 days; (d) colony overview; (e–g) conidiogenous cells and conidia; (h,i) conidia. Scale bars: (e–i) 10 μm.
Figure 3.
Microdochium nannuoshanense (holotype, HMAS 352652). (a) A leaf of Bambusaceae sp; (b) colonies on PDA from above and below after 14 days; (c) colonies on OA from above and below after 14 days; (d) colony overview; (e–g) conidiogenous cells and conidia; (h,i) conidia. Scale bars: (e–i) 10 μm.

3.2.3. Microdochium phyllosaprophyticum J. Zhang, Z.X. Zhang, & Z. Li, sp. Nov.; Figure 4
MycoBank No. MB850599
Etymology—Referring to the generic name of the host plant’s saprophytic leaves (Greek “phyllum” and Latin “sapro[phyticum]”).
Type—China, Hainan Province: Bawangling National Forest Park; on saprophytic leaves; 11 April 2023; J. Zhang; holotype HMAS 352653; ex-holotype living culture SAUCC SAUCC 3583-1.
Description—Saprophytic on dead leaves. Sexual morphs are unknown. Mycelia are superficial and immersed, 2.4–3.3 µm wide, branched, membranous and hyaline. Conidiophores are inapparent and often reduced to conidiogenous cells. Conidiogenous cells are straight or slightly curved, 16.0–32.0 × 2.0–2.5 µm, mono- or polyblastic, terminal, denticulate, hyaline, smooth, cylindrical and membranous, and produced on aerial mycelia. Conidia are solitary, hyaline, cylindrical, oblong to ellipsoid, spindle to rod-shaped, straight or curved, 7.5–15.0 × 2.0–3.5 µm, multi-guttulate and sometimes borne directly from hyphae. Chlamydospores were not observed; see Figure 4.
Culture characteristics—Cultures incubated on PDA at 25 °C in darkness, reaching 60.0–63.0mm diam., had a growth rate of 4.3–4.5 mm diam/day after 14 days, were creamy white with regular margins, had luxuriant aerial hyphae, and the reverses were light brown in the center and white at the edge. Cultures incubated on PDA at 25 °C in darkness, reaching 62.0–68.0 mm diam., had a growth rate of 4.4–4.8 mm diam/day after 14 days, were creamy white with regular margins, had luxuriant aerial hyphae, and the reverses were similar.
Additional specimen examined—China, Hainan Province: Bawangling National Forest Park; on saprophytic leaves; 11 April 2023; J. Zhang; HSUAP3583-6; living culture SAUCC SAUCC 3583-6.
Notes—Phylogenetic analyses of four combined genes (ITS, LSU and RPB2) showed that Microdochium phyllosaprophyticum sp. nov. formed an independent clade closely related to M. bambusae and M. indocalami. M. phyllosaprophyticum is distinguished from M. bambusae (SAUCC 1862-1) by 6/535, 15/836, 81/857 and 15/714 characters and from M. indocalami (SAUCC 1016) by 8/535, 14/836, 63/840 and 17/714 characters in ITS, LSU, RPB2 and TUB2 sequences, respectively. Morphologically, M. phyllosaprophyticum differs from M. bambusae and M. indocalami in conidia (7.5–15.0 × 2.0–3.5 μm vs. 13.0–17.0 × 2.5–3.5 μm vs. 13.0–15.5 × 3.5–5.5 μm) and in colony texture (light brown to creamy white with regular margins vs. gray white with regular margins vs. white with regular margins on PDA) [21]. For details, see Table 3.
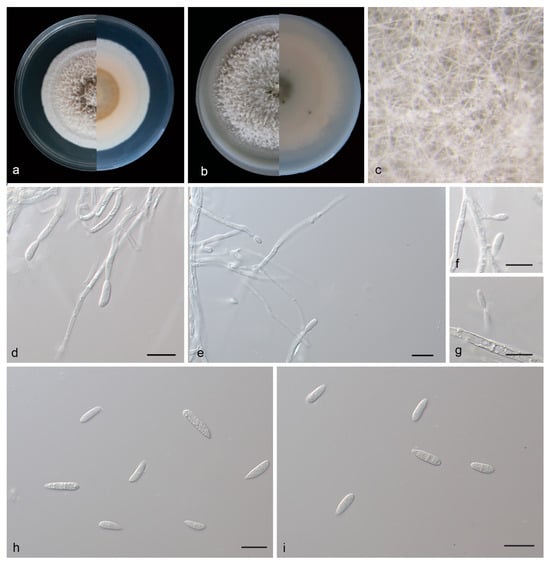
Figure 4.
Microdochium phyllosaprophyticum (holotype, HMAS 352653). (a) Colonies on PDA from above and below after 14 days; (b) colonies on OA from above and below after 14 days; (c) colony overview; (d–g) conidiogenous cells and conidia; (h,i) conidia. Scale bars: (e–i) 10 μm.
Figure 4.
Microdochium phyllosaprophyticum (holotype, HMAS 352653). (a) Colonies on PDA from above and below after 14 days; (b) colonies on OA from above and below after 14 days; (c) colony overview; (d–g) conidiogenous cells and conidia; (h,i) conidia. Scale bars: (e–i) 10 μm.
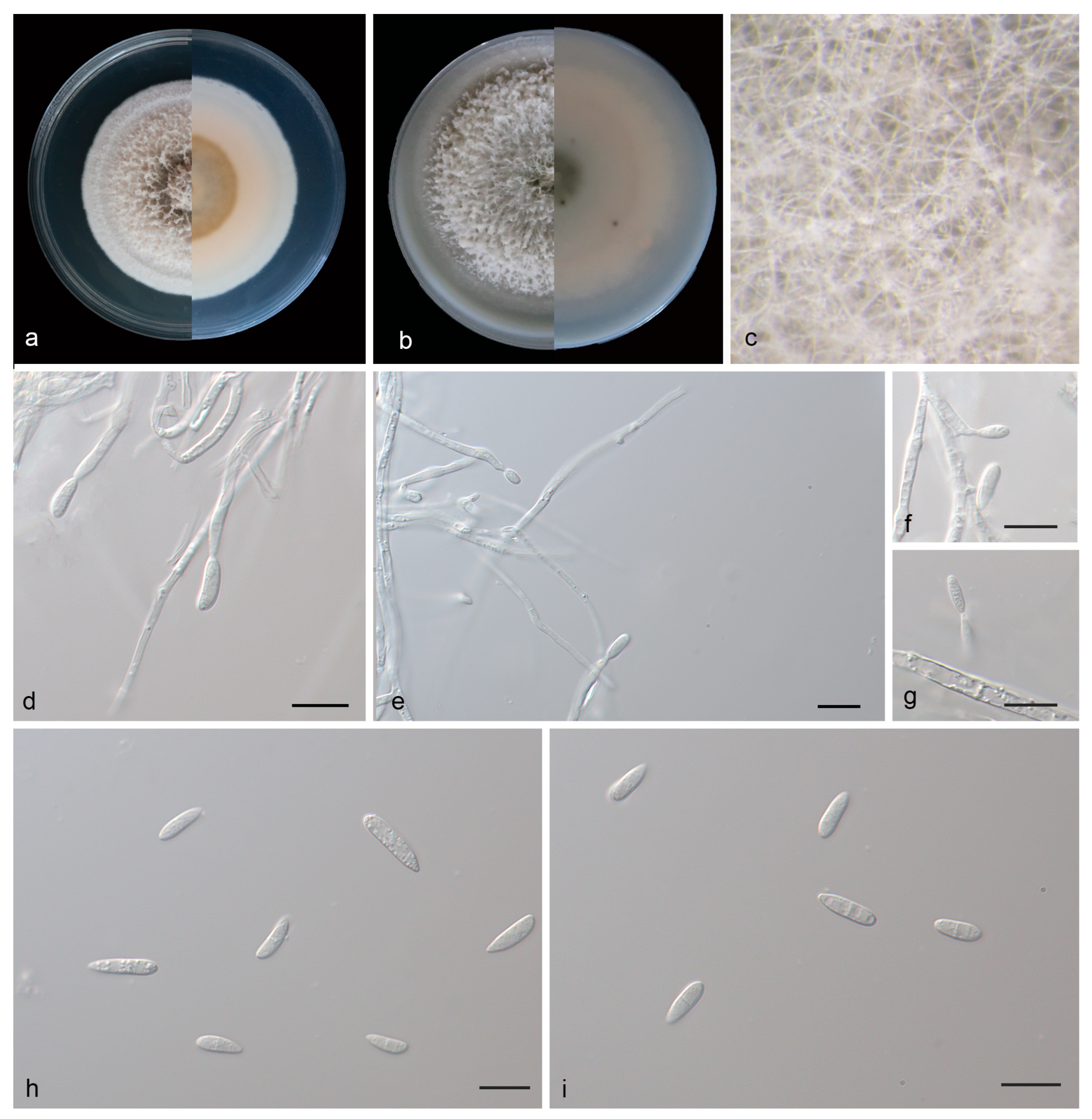

Table 3.
The asexual morphological characters of some Microdochium species.
Table 3.
The asexual morphological characters of some Microdochium species.
| Species | Conidiogenous Cells | Size of Conidiogenous Cells (μm) | Conidia | Size of Conidia (μm) | References |
|---|---|---|---|---|---|
| Microdochium albescens | Doliiform to obpyriform | 6–15 × 1.5–4 | Falcate, apex pointed | 11–16 × 3.5–4.5 | [13] |
| M.bambusae | Cylindrical and smooth | 17.4–30.0 × 2.5–3.0 | Oblong to ellipsoid | 13.0–17.0 × 2.5–3.5 | This study |
| M. bolleyi | Ampullate or cylindrical | 3.1–6.4 × 2.5–3.8 | Crescent, lunate | 5.0–8.7 × 1.6–2.3 | [13] |
| M. chrysanthemoides | Cylindrical to ellipsoidal | 5–12 × 3.0–4.5 | Ellipsoid or allantoid | 4.5–7 × 2–3 | [13] |
| M. chuxiongense | Solitary, clavate | 27–74 × 2–3 | Shuttle or sickle | 4–12 × 2–5 | [39] |
| M. citrinidiscum | Denticulate, cylindrical | 11–29 × 1.5–2 | clavate to obovoid | 7–31 × 2–3 | [13] |
| M. colombiense | Ampulliform, polyblastic | 5–11.5 × 2.5–3.5 | Fusiform, allantoid | 5–8 × 1.5–2.5 | [13] |
| M. dawsoniorum | Cylindrical to irregular | 20–30 × 1–2 | Flexuous to falcate | 25–75 × 1–2 | [13] |
| M. fisheri | Denticulate, cylindrical | 19–60 × 1.5–2 | Obovoid, subpyriform | 7–12 × 3–4 | [13] |
| M. hainanense | Ampulliform and lageniform | 4.8–8.2 × 2.0–2.5 | Spindle to rod shaped | 7.0–16.1 × 2.5–4.7 | [21] |
| M. indocalami | Cylindrical, straight or bent | 11.0–28.3 × 1.5–2.9 | Clavate to obovoid | 13.0–15.5 × 3.5–5.5 | [17] |
| M. lycopodinum | Ampulliform to lageniform | 4–12 × 2.5–3.5 | Fusiform or with one side | 8–15 × 2.5–3.5 | [13] |
| M. maculosum | Denticulate, raduliform | 7–39 × 1–3 | Fusiform, straight or curved | 6–15 × 2–4 | [18] |
| M. miscanthi | Smooth and cylindrical | 9.7–14.5 × 3.6–4.1 | Spindle to rod shaped | 7.0–16.1 × 2.5–4.7 | [21] |
| M. musae | |||||
| M. nannuoshanense | Cylindrical and smooth | 19.0–27.0 × 2.0–3.0 | Oblong to ellipsoid | 7.0–12.0 × 2.5–4.0 | This study |
| M. neoqueenslandicum | Lageniform to subcylindrical | 4.5–10 × 2–3.5 | Lunate, allantoid | 4–9 × 1.5–3 | [13] |
| M. nivale | Doliiform to obpyriform | 6–15 × 2.2–4 | Falcate, apex pointed | 5–36 × 2–4.5 | [13] |
| M. paspali | Lageniform to cylindrical | 6.5–15.5 × 2.5–4 | Falcate, apex pointed | 7–20.5 × 2.5–4.5 | [13] |
| M. phyllosaprophyticum | Cylindrical and smooth | 16.0–32.0 × 2.0–2.5 | Oblong to ellipsoid | 7.5–15.0 × 2.0–3.5 | This study |
| M. phragmitis | Ampulliform to lageniform | 5–12(–30) × 2.5–3 | Ellipsoid-fusiform | 10–16 × 2–3.5 | [13] |
| M. poae | Cylindrical or subcylindrical | 1.5–6.5 ×1–2 | Fusiform, ovoid, pyriform | 3.5–8.5 ×2–3 | [13] |
| M. ratticaudae | – | – | Fusoid, falcat | 7-11 × 1.5-2.5 | [13] |
| M. rhopalostylidis | Ampulliform | 4–10 × 3–3.5 | Fusoid, curved | 16–20(–23) × 2.5–3 | [15] |
| M. seminicola | Ampulliform to lageniform | 7–9.5 × 3–4 | Cylindrical to fusiform | 19–54 × 3–4.5 | [13] |
| M. sinense | Smooth and cylindrical | 16.3–22.4 × 4.1–5.7 | Spindle shaped or cylindrical | 11.5–19.34 × 2.8–5.4 | [21] |
| M. sorghi | Ampulliform to obclavate | 5–13 × 3–4 | Filiform, obclavate | 20–90 × 1.5–4.5 | [13] |
| M. tainanense | Cylindrical or ampulliform | 3–10 × 1–3 | Lunate | 10–15 × 2–3 | [13] |
| M. trichocladiopsis | Cylindrical to clavate | 4–37 × 2–3 | Oblong, fusiform to obovoid | 6–18 × 2–3.5 | [13] |
| M. yunnanense | Ampulliform, lageniform | 6.5–10.0 × 2.5–3.4 | Ellipsoid and cylindrical | 6.8–10.0 × 2.4–3.5 | [17] |
4. Discussion
Microdochium was established by Syd. & P. Syd. in 1924; it has belonged to Microdochiaceae since 2016 (Microdochiacea was introduced by Hernández-Restrepo in 2016) [1,9,10,11,12]. Monographella Petr. was considered a sexual morph of the genus; “One Fungus = One Name” was proposed in The Amsterdam Declaration on Fungal Nomenclature in 2011, but the genus name of Microdochium was retained as the correct genus name [40]. With the development of molecular techniques, Microdochium was divided into a new family called Microdochiaceae by Hernández-Restrepo et al. [8]. Microdochiaceae is characterized by asexual morphs of polyblastic, sympodial or annellidic conidiogenous cells with hyaline conidia; the conidia come in a variety of shapes, i.e., cylindrical, fusiform, oval, rod-shaped, vertical or curved, truncated at the base and mostly rounded at the apex, and Monographella-like sexual morphs [8,21].
In the present study, six strains from one host plant (Bambusaceae sp.) and saprophytic leaves were split into three new species (Microdochium bambusae, M. nannuoshanense and M. phyllosaprophyticum). Currently, the Global Biodiversity Information Facility (GBIF, https://www.gbif.org/, accessed on 30 October 2023) contains 1693 georeferenced records of Microdochium species reported around the world. America, Asia, Europe and Oceania were the main distribution locations for this species; the United States has an especially great distribution, followed by Poland [2,8,13,14,15,19,20,41]. Since the 21st century, 12 species of this genus have been reported from five provinces (Fujian, Guizhou, Hainan, Sichuan and Yunnan) and Beijing in China (including the present study), viz, Microdochium bambusae sp. nov. (Bambusaceae sp.), M. chrysanthemoides (air), M. chuxiongense (Bondarzewia sp.), M. graminearum (decaying herbaceous grass stem), M. hainanense (Phragmites australis), M. indocalami (Indocalamus longiauritus), M. miscanthi (Miscanthus sinensis), M. nannuoshanense sp. nov. (Bambusaceae sp.), M. paspali (Paspalum vaginatum), M. phyllosaprophyticum sp. nov. (saprophytic leaves), M. poae (Poa annua), M. shilinense (decaying herbaceous stem of grass), M. sinense (Miscanthus sinensis) and M. yunnanense (Indocalamus longiauritus) [2,13,17,20,21,39,42]. Previous studies have shown that Microdochium species had been introduced on a range of host families, such as Arecaceae, Asteraceae, Cactaceae, Euphorbiaceae, Iridaceae, Lycopodiaceae, Musaceae, Myrtaceae, Passifloraceae and Phyllanthaceae, and more than half of Microdochium fungi were associated with Poaceae plants [42]. In particular, the study of Bambusaceae, as a subfamily of Poaceae plants, has become increasingly popular in recently years; an especially large number of Apiospora fungi have been isolated from Bambusaceae [43], whereas only a few Microdochium fungi have been isolated from Bambusaceae.
Microdochium species were reported as endophytes, plant pathogens and saprophytes. M. bolleyi was reported as a root endophyte and was proved to act against Gaeumannomyces graminis var. tritici, and it may be further used as a biocontrol agent [44]. Most species of Microdochium were reported as plant pathogens, especially against economical cereal crops: for example, Microdochium albescens, M. oryzae isolated from Oryza sativa; M. majus, M. nivale, M. trichocladiopsis and M. triticicola isolated from Triticum aestivum; and so on [42]. In reality, some species have not been regarded as plant pathogens; they are only isolated from leaf spot. However, the pathogenicity of this fungi has not been proven by Koch’s Postulates. Generally, we believe that the type of fungus is an endophytic fungus associated with leaf spot. In addition, we should focus more on disease detection and biological control.
5. Conclusions
In this study, we isolated six strains associated with Microdochium on Bambusaceae sp. and saprophytic leaves by tissue isolation and single spore isolation from Hainan and Yunnan provinces in China. Based on morphology and phylogeny, six strains were identified as three new species, viz., Microdochium bambusae sp. nov., M. nannuoshanense sp. nov. and M. phyllosaprophyticum sp. nov. In the future, we firmly believe that Microdochium species will be isolated from more plants around the world.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/jof9121176/s1, Supplementary File S1: The combined ITS, LSU, RPB2 and TUB2 sequences.
Author Contributions
Conceptualization, J.Z.; methodology, J.Z.; software, Z.Z.; validation, Z.Z.; formal analysis, Z.Z.; investigation, D.L.; resources, D.L.; data curation, J.X.; writing—original draft preparation, J.X.; writing—review and editing, Z.L.; visualization, Z.L.; supervision, Z.L.; project administration, Z.L.; funding acquisition, Z.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research was supported by National Natural Science Foundation of China (nos. 32270024, U2002203 and 32370001).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The sequences from the present study were submitted to the NCBI database (https://www.ncbi.nlm.nih.gov/, accessed on 20 October 2023), and the accession numbers are listed in Table 2.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Sydow, H. Mycotheca germanic. Fasc. XLII–XLV (No. 2051–2250). Annls Mycol. 1924, 22, 257–268. [Google Scholar]
- Liang, J.M.; Li, G.S.; Zhao, M.Q.; Cai, L. A new leaf blight disease of turfgrasses caused by Microdochium poae, sp. nov. Mycologia 2019, 111, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Von Arx, J.A. Plant pathogenic fungi. Nova Hedwig. 1987, 87, 1–288. [Google Scholar]
- Hiruma, K.; Kobae, Y.; Toju, H. Beneficial associations between Brassicaceae plants and fungal endophytes under nutrientlimiting conditions: Evolutionary origins and host–symbiont molecular mechanisms. Curr. Opin. Plant Biol. 2018, 44, 145–154. [Google Scholar] [CrossRef] [PubMed]
- Mandyam, K.G.; Roe, J.; Jumpponen, A. Arabidopsis thaliana model system reveals a continuum of responses to root endophyte colonization. Fungal Biol. 2013, 117, 250–260. [Google Scholar] [CrossRef] [PubMed]
- Glynn, N.C.; Hare, M.C.; Parry, D.W.; Edwards, S.G. Phylogenetic analysis of EF-1 alpha gene sequences from isolates of Microdochium nivale leads to elevation of varieties Majus and Nivale to species status. Fungal Biol. 2005, 109, 872–880. [Google Scholar] [CrossRef] [PubMed]
- Jewell, L.E.; Hsiang, T. Multigene differences between Microdochium nivale and Microdochium majus. Botany 2013, 91, 99–106. [Google Scholar] [CrossRef]
- Hernández-Restrepo, M.; Groenewald, J.Z.; Crous, P.W. Taxonomic and phylogenetic re-evaluation of Microdochium, Monographella and Idriella. Pers. Mol. Phylogeny Evol. Fungi. 2016, 36, 57–82. [Google Scholar] [CrossRef]
- Parkinson, V.O.; Sivanesan, A.; Booth, C. The perfect state of the rice leaf-scald fungus and the taxonomy of both the perfect and imperfect states. Trans. Br. Mycol. Soc. 1981, 76, 59–69. [Google Scholar] [CrossRef]
- Samuels, G.J.; Hallett, I.C. Microdochium stoveri and Monographella stoveri, new combinations for Fusarium stoveri and Micronectriella stoveri. Trans. Br. Mycol. Soc. 1983, 81, 473–483. [Google Scholar] [CrossRef]
- Von Arx, J.A. Notes on Monographella and Microdochium. Trans. Br. Mycol. Soc. 1984, 83, 373–374. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Voglmayr, H. Phylogenetic relationships of five genera of Xylariales and Rosasphaeria gen. nov. (Hypocreales). Fungal Divers. 2012, 52, 75–98. [Google Scholar] [CrossRef]
- Zhang, Z.F.; Liu, F.; Zhou, X.; Liu, X.Z.; Liu, S.J.; Cai, L. Culturable Mycobiota from karst caves in China, with descriptions of 20 new species. Pers. Mol. Phylogeny Evol. Fungi. 2017, 39, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Schumacher, R.K.; Wingfield, M.J.; Akulov, A.; Denman, S.; Roux, J.; Braun, U.; Burgess, T.I.; Carnegie, A.J.; Váczy, K.Z. New and interesting fungi 1. Fuse 2018, 1, 169–215. [Google Scholar] [CrossRef]
- Crous, P.W.; Schumacher, R.K.; Akulov, A.; Thangavel, R.; Hernández-Restrepo, M.; Carnegie, A.J.; Cheewangkoon, R.; Wingfield, M.J.; Summerell, B.A.; Quaedvlieg, W. New and interesting fungi 2. Fuse 2019, 3, 57–134. [Google Scholar] [CrossRef]
- Marin-Felix, Y.; Hernández-Restrepo, M.; Wingfield, M.J.; Akulov, A.; Carnegie, A.J.; Cheewangkoon, R.; Gramaje, D.; Groenewald, J.Z.; Guarnaccia, V.; Halleen, F. Genera of phytopathogenic fungi: GOPHY 2. Stud. Mycol. 2019, 92, 47–133. [Google Scholar] [CrossRef]
- Huang, S.T.; Xia, J.W.; Zhang, X.G.; Sun, W.X.; Li, Z. Two new species of Microdochium from Indocalamus longiauritus in south-western China. MycoKeys 2020, 72, 93–108. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Cowan, D.A.; Maggs-Kölling, G.; Yilmaz, N.; Thangavel, R.; Wingfield, M.J.; Noordeloos, M.E.; Dima, B.; Brandrud, T.E.; Jansen, G.M.; et al. Fungal Planet description sheets: 1182–1283. Pers. Mol. Phylogeny Evol. Fungi. 2021, 46, 313–528. [Google Scholar] [CrossRef]
- Kwasna, H.; Bateman, G.L. Microdochium Triticicola sp. nov. from roots of Triticum aestivum in the United Kingdom. Mycologia 2007, 99, 765–776. [Google Scholar] [CrossRef]
- Zhang, W.; Nan, Z.B.; Tian, P.; Hu, M.J.; Gao, Z.Y.; Li, M.; Liu, G.D. Microdochium paspali, a new species causing seashore paspalum disease in southern China. Mycologia 2015, 107, 80–89. [Google Scholar] [CrossRef]
- Liu, S.B.; Liu, X.Y.; Zhang, Z.X.; Xia, J.W.; Zhang, X.G.; Meng, Z. Three new species of Microdochium (Sordariomycetes, Amphisphaeriales) on Miscanthus sinensis and Phragmites australis from Hainan, China. J. Fungi 2022, 8, 577. [Google Scholar] [CrossRef] [PubMed]
- Dissanayake, L.S.; Kang, J.C.; Maharachchikumbura, S.S.N. Microdochium sichuanense sp. nov. (Microdochiaceae, Xylariales), from a Poaceae host in Sichuan, China. Phytotaxa 2023, 600, 206–216. [Google Scholar] [CrossRef]
- Jiang, N.; Voglmayr, H.; Ma, C.Y.; Xue, H.; Piao, C.G.; Li, Y. A new Arthrinium-like genus of Amphisphaeriales in China. MycoKeys 2022, 92, 27–43. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.X.; Liu, X.Y.; Tao, M.F.; Liu, X.Y.; Xia, J.W.; Zhang, X.G.; Meng, Z. Taxonomy, Phylogeny, Divergence Time Estimation, and Biogeography of the Family Pseudoplagiostomataceae (Ascomycota, Diaporthales). J. Fungi 2023, 9, 82. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.D.; Hyde, K.D.; Liew, E.C.Y. Identification of endophytic fungi from Livistona chinensis based on morphology and rDNA sequences. New Phytol. 2000, 147, 617–630. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, F.J.R.M.; Lee, S.H.; Taylor, L.; Shawe-Taylor, J. Amplification and direct sequencing of fungal ribosomal rna genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., Eds.; Academic Press Inc.: New York, NY, USA, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic Relationships among Ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Sung, G.H.; Sung, J.M.; Hywel-Jones, N.L.; Spatafora, J.W. A multi-gene phylogeny of Clavicipitaceae (Ascomycota, Fungi): Identification of localized incongruence using a combinational bootstrap approach. Mol. Phylogenet. Evol. 2007, 44, 1204–1223. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef]
- Nylander, J.A.A. MrModelTest v. 2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. The CIPRES science gateway: Enabling high-impact science for phylogenetics researchers with limited resources. In Proceedings of the 1st Conference of the Extreme Science and Engineering Discovery Environment. Bridging from the Extreme to the Campus and Beyond, Chicago, IL, USA, 16–20 July 2012; Association for Computing Machinery: San Diego, CA, USA, 2012; p. 8. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogeny. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian Phylogenetic Inference under Mixed Models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, 293–296. [Google Scholar] [CrossRef] [PubMed]
- Tang, D.; Zou, W.; Wang, Y.; Huang, O.U.; Yu, H. Morphology and phylogeny of Microdochium chuxiongense sp. nov., a fungus from Southwest China. Phytotaxa 2022, 555, 147–158. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Crous, P.W.; Redhead, S.A.; Reynolds, D.R.; Samson, R.A.; Seifert, K.A.; Taylor, J.W.; Wingfield, M.J.; Abaci, O.; Aime, C. The Amsterdam declaration on fungal nomenclature. IMA Fungus 2011, 2, 105–111. [Google Scholar] [CrossRef]
- Abdelhalim, M.; Brurberg, M.B.; Hofgaard, I.S.; Rognli, O.A.; Tronsmo, A.M. Pathogenicity, host specificity and genetic diversity in Norwegian isolates of Microdochium nivale and Microdochium majus. Eur. J. Plant Pathol. 2020, 156, 885–895. [Google Scholar] [CrossRef]
- Gao, Y.; Ren, G.; Wanasinghe, D.N.; Xu, J.; Gomes De Farias, A.R.; Gui, H. Two New Species and a New Record of Microdochium from Grasses in Yunnan Province, South-West China. J. Fungi 2022, 8, 1297. [Google Scholar] [CrossRef]
- Monkai, J.; Phookamsak, R.; Tennakoon, D.S.; Bhat, D.J.; Xu, S.; Li, Q.; Lumyong, S. Insight into the Taxonomic Resolution of Apiospora: Introducing Novel Species and Records from Bamboo in China and Thailand. Diversity 2022, 14, 918. [Google Scholar] [CrossRef]
- Shadmani, L.; Jamali, S.; Fatemi, A. Biocontrol activity of endophytic fungus of barley, Microdochium bolleyi, against Gaeumannomyces graminis var. tritici. Mycol. Iran. 2018, 5, 7–14. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
