1. Introduction
Acute hepatopancreatic necrosis disease (AHPND) is a bacterial disease caused by
Vibrio spp. carrying toxin genes (
pirA and
pirB) located in a large plasmid (69 kb). AHPND affects the digestive tract of shrimp and the tubular cells of the hepatopancreas, disturbing digestion and resulting in mass mortality.
V.
parahaemolyticus is primarily associated with AHPND (
VpAHPND), but other
Vibrio species that carry binary toxin genes, including
V. campbellii (
VcAHPND),
V. owensii (
VoAHPND), and
V. harveyi (
VhAHPND), have been reported recently [
1,
2,
3,
4]. AHPND was first reported in China (2009), and it spread to several countries, including Vietnam (2010), Malaysia (2011), Thailand (2012), Mexico (2013), the Philippines (2015), the USA (2019), and South Korea (2020) [
1,
5,
6,
7,
8,
9,
10]. This disease is known to cause tremendous economic losses in the shrimp aquaculture industry. Shrimp production has considerably decreased following the outbreak of AHPND, and the economic damage is estimated to exceed 1 billion dollars per year in Asia [
11].
Although antibiotics have been extensively used to treat bacterial diseases in aquaculture for decades [
12], their utilization particularly in the form of overuse or misuse has resulted in antibiotic resistance [
13,
14,
15,
16,
17,
18]. As antibiotic alternatives, probiotics have been frequently used in aquaculture to control bacterial diseases, especially against pathogenic
Vibrio infections and AHPND. In a previous report, shrimp treated with
Bacillus probiotics in the form of dietary supplements showed a higher survival rate following challenge with
VpAHPND [
19,
20]. In addition to their antimicrobial activity, probiotics have various advantages in aquaculture such as promoting growth, strengthening immunity, and restoring water quality [
21,
22]. Meanwhile, spore-forming
Bacillus spp. are resistant to heat and pressure and are widely used as feed additives [
23].
Besides the use of probiotics, various methods have been utilized in shrimp aquaculture such as immunostimulant therapy, quorum sensing control of bacterial virulence, phage therapy, and herbal medicine [
18,
24,
25,
26]. Paopradit et al. [
27] reported the reduced virulence and decreased mortality of
VpAHPND following treatment with quorum sensing inhibitors such as indole or indole-containing compounds. In addition, previous studies [
28,
29] have confirmed the control of both growth and infection of
VpAHPND using bacteriophages in double-layer agar culture and a series of bioassays, respectively.
Although
V. parahaemolyticus is the cause of most cases of AHPND, other
Vibrio spp., such as
V. campbellii,
V. harveyi, and
V. owensii, are also known to cause this disease in fields, thereby resulting in substantial economic losses on farms. However, preventative methods and studies on AHPND have mainly focused on
VpAHPND, and the antimicrobial activity against
VcAHPND,
VhAHPND, and
VoAHPND has been poorly studied [
25,
30]. In this study, we isolated five
Bacillus strains and evaluated their antimicrobial activity against ten AHPND-causing
Vibrio strains and two non-AHPND
Vibrio strains using a dot-spot test (in vitro). In addition, a challenge test against
VcAHPND was performed using two
Bacillus strains (B1 and B3) that strongly inhibited
VcAHPND among five
Bacillus strains in the dot-spot test. The findings revealed that B1 and B3 treatment groups significantly suppressed
VcAHPND growth compared with the effect of the non-
Bacillus treatment group. Finally, the genomic sequences of these two
Bacillus strains were completely analyzed, and both strains were classified as
B. velezensis. Their 16S rRNA sequences and complete genome sequences were deposited in GenBank. The results of this study provide useful and practical strategies that can be applied in the shrimp culture industry, which is currently experiencing declines in shrimp production because of harmful shrimp diseases, including AHPND caused by
VpAHPND, and
VcAHPND.
2. Materials and Methods
2.1. Bacillus and Vibrio Candidate Isolation and Polymerase Chain Reaction (PCR)
For the isolation of
Bacillus spp., seawater samples were collected from six different sites in Jeju Island, South Korea. These seawater samples were subjected to a serial dilution process, and dilutions were spread onto tryptic soy agar (TSA; Difco, Becton Dickinson, Franklin Lakes, NJ, USA) plates supplemented with 2% NaCl (TSA+). The plates were incubated at 28 °C for 24–48 h. Subsequently, we picked the bacterial colonies displaying sporulated shapes on the agar plates based on morphology, and the colonies were sub-cultured for pure culture isolation. Isolates were preserved in tryptic soy broth (TSB; Difco, Becton Dickinson, Franklin Lakes, NJ, USA) supplemented with 2% NaCl (TSB+) containing 25% glycerol at −80 °C until further analysis. Each isolate was grown in TSB+ (28 °C, 200 rpm, 24–48 h) and DNA was extracted using the protocol of the modified DNeasy
® Blood & Tissue Kit (Qiagen, Germany). For
Bacillus identification, PCR was performed using the extracted DNA, and the BacF/R primers, as described by Solichova et al. [
31] (
Table 1).
For the isolation of
Vibrio spp., seawater and hepatopancreas samples were collected from Mexico, Vietnam, Thailand, South Korea, and the USA. These samples were serially diluted and spread on thiosulfate citrate bile salts sucrose (TCBS) (MB Cell, South Korea) agar plates, which were incubated at 28 °C for 24–48 h. Next, we picked green and yellow colonies from the TCBS plates, and the colonies were sub-cultured for pure culture isolation. Isolates were preserved in TSB+ containing 25% glycerol at −80 °C until further analysis. Each isolate was grown in TSB+ (28 °C, 200 rpm, 24–48 h) and used for DNA extraction using the boiling method described by Dashti et al. [
32]. For
Vibrio identification, PCR was conducted using the extracted DNA and the primer sets (Tox R-F/R, Vc.fts.z-F/R, and Vh.topA-F/R) described by Kim et al. [
33] and Cano-Gomez et al. [
34] (
Table 1). To identify AHPND virulence genes, PCR targeting AHPND toxin genes (
pirA and
pirB) was conducted using the method described by Han et al. [
35] (
Table 1).
Table 1.
Primers for Bacillus, AHPND toxin genes (pirA and pirB), and Vibrio species.
Table 1.
Primers for Bacillus, AHPND toxin genes (pirA and pirB), and Vibrio species.
| Target | Primers | Sequence (5′–3′) | Amplicon Size (bp) | Reference |
|---|
| Bacillus | BacF | GCTGGTTAGAGCGCACGCCTGATA | 263 | [31] |
| BacR | CATCCACCGTGCGCCCTTTCTAAC |
| AHPND toxin | VpPirA-284F | TGACTATTCTCACGATTGGACTG | 284 | [35] |
| VpPirA-284R | CACGACTAGCGCCATTGTTA |
| VpPirA-392F | TGATGAAGTGATGGGTGCTC | 392 |
| VpPirA-392R | TGTAAGCGCCGTTTAACTCA |
| V. parahaemolyticus | Tox R-F | GTCTTCTGACGCAATCGTTG | 368 | [33] |
| Tox R-R | ATACGAGTGGTTGCTGTCATG |
| V. campbellii | Vc.fts.z-F | AAGACAGAGATAGACTTAAAGAT | 294 | [34] |
| Vc.fts.z-R | CTTCTAGCAGCGTTACAC |
| V. harveyi | Vh.topA-F | TGGCGCAGCGTCTATACG | 121 |
| Vh.topA-R | TATTTGTCACCGAACTCAGAACC |
2.2. Antimicrobial Activity Test (In Vitro)
For antimicrobial activity testing, the
Bacillus strains that were obtained were further tested for their ability to inhibit the growth of
Vibrio strains using the dot-spot method described by Spelhaug and Harlander [
36].
Vibrio strains were grown in TSB+ with shaking at 200 rpm and 28 °C for 24 h, and bacterial suspensions of each strain were normalized with 2.5% NaCl to obtain a final concentration of approximately 5 × 10
6 colony forming units (CFU) mL
−1.
Bacillus strains were grown in TSB+ with shaking at 200 rpm and 28 °C for 24–48 h to obtain a final concentration of approximately 5 × 10
8 CFU mL
−1. Then, 100 μL of each
Vibrio strain suspension was inoculated into 5 mL of soft agar and poured onto TSA+ plates. Ten-microliter aliquots of
Bacillus strain suspensions were dot-spotted on the surface of
Vibrio-overlaid agar. The plates were incubated at 28 °C for 12–24 h, and the clear zones around each
Bacillus colony were recorded.
B. velezensis CR-502T (=NRRL B-41580T) was obtained from the Korean Collection for Type Cultures (KCTC) and set as the reference strain in this experiment. The experiments were also conducted using the same methods.
2.3. Antimicrobial Activity Test (Challenge Test)
Bacillus strains that showed the strongest inhibitory effects in the dot-spot test were further subjected to the challenge test. As experimental shrimp, the Pacific white leg shrimp (Penaeus vannamei) at the post-larval stage (stages PL15–PL16) were purchased from a local shrimp farm (Jeju Province, South Korea) and transported to the Laboratory of Aquatic Biomedicine, College of Veterinary Medicine, Kyungpook National University in South Korea. Shrimp were fed a commercial diet twice daily in a 700 L acrylic tank for 35 days to be acclimated to the experimental conditions and facilities. Then, the shrimp (average weight of 0.2 ± 0.05 g) were randomly distributed into 22 L acrylic tanks with 18 L of aerated artificial seawater. For the antimicrobial activity test (challenge test), experimental shrimp (N = 56) were divided into four groups with duplicates.
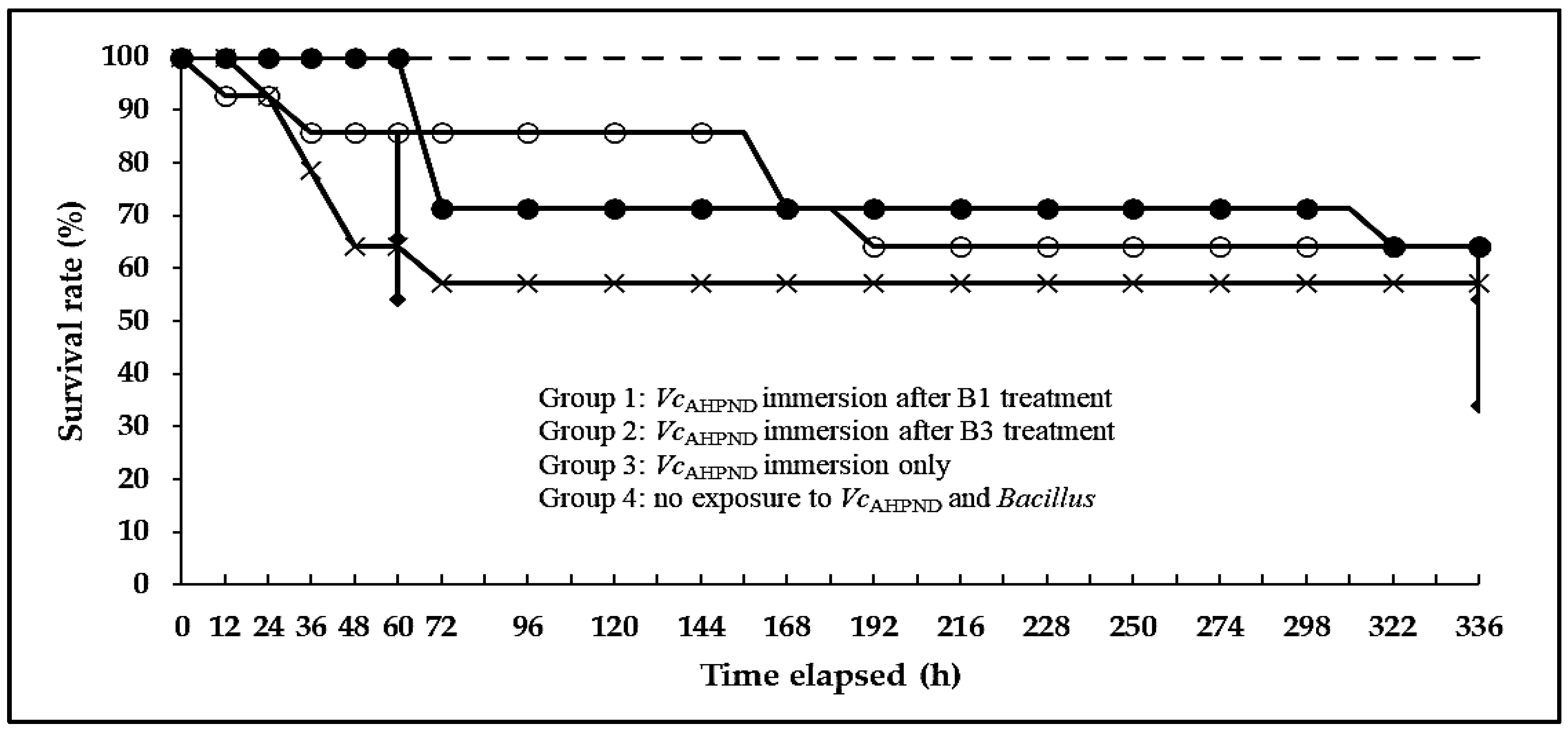
In group 1, the experimental shrimp (N = 14) were exposed to a suspension of Bacillus (B1) for 14 days via immersion at a concentration of 1.0 × 106 CFU mL−1 water. Then, the shrimp were challenged with a VcAHPND (16-904/1) suspension via immersion at a concentration of 2.0 × 106 CFU mL−1 water. In group 2, the experimental shrimp (N = 14) were exposed to Bacillus (B3) suspension for 14 days via immersion at a concentration of 1.0 × 106 CFU mL−1 water. Then, the shrimp were challenged with a VcAHPND (16-904/1) suspension via immersion at a concentration of 2.0 × 106 CFU mL−1 water. In group 3, the experimental shrimp (N = 14) were exposed to the same amount of fresh broth (TSB+) without Bacillus strains (B1 and B3) for 14 days via immersion. Then, they were challenged with a VcAHPND (16-904/1) suspension via immersion at a concentration of 2.0 × 106 CFU mL−1 water. In group 4, the experimental shrimp (N = 14) were exposed to the same amount of fresh broth (TSB+) without Bacillus for 14 days, and then they were not challenged VcAHPND (16-904/1). The experiment was started at the same time and under the same conditions for all groups. The tanks were filled with aerated artificial seawater and maintained without water change for 28 days. The water temperature, dissolved oxygen level, pH, and salinity were maintained at 25–28 °C, 6.39–7.21 ppm, 6.48–7.10, and 23–25 ppt, respectively. Shrimp were fed shrimp feed three times a day at 5% of their body weight and monitored for 28 days.
To confirm the presence of AHPND, dead shrimp were collected and tested using the PCR method previously described by Han et al. [
35]. To quantify AHPND, surviving shrimp were randomly sampled on the day of termination (day 14). The hepatopancreas of each shrimp was collected aseptically; next, 30 mg of the hepatopancreas tissue was used for DNA extraction using the DNeasy
® Blood & Tissue Kit. Using the extracted DNA, quantitative PCR was performed to quantify the AHPND toxin gene
pirA in the hepatopancreas in the groups using the method described by Han et al. [
37].
2.4. Genome Sequencing and Phylogenetic Analysis of the Selected Bacillus Strains
The genomes of two selected
Bacillus strains (B1 and B3) were sequenced using a hybrid approach on a PacBio RS II system (Pacific Biosciences Inc., Menlo Park, CA, USA) by constructing a 20 kb SMRTbellTM template library and on the HiSeq X-10 platform (Illumina Inc., San Diego, CA, USA) by preparing a DNA library using the TruSeq Nano DNA Library Prep Kit (Illumina). Genome assembly of the filtered PacBio reads was performed using the HGAP (v3.0) pipeline, the 150-bp Illumina paired-end reads were mapped using BWA-MEM (v0.7.15), and errors were corrected using Pilon (v1.21) using the default parameters. Annotation was performed using the NCBI Prokaryotic Genome Annotation Pipeline (
http://www.ncbi.nlm.nih.gov/books/NBK174280/ (accessed on August 2022) [
38]. The regions and clusters of secondary metabolites present in the genomes of both strains were predicted using antibiotics & Secondary Metabolite Analysis Shell (anti-SMASH) v.6.1.1 [
39]. The phylogenetic trees of the two
Bacillus strains based on the 16S ribosomal RNA (rRNA) genes and whole-genome sequences were constructed using selected 20-type species of the genus
Bacillus. First, the 16S rRNA sequences of the two isolates were aligned with 20 representative species of the genus
Bacillus using Clustal X (ver. 2.0) [
40] and BioEdit (ver. 7.0) [
41], and the maximum-likelihood phylogenetic tree based on the concatenated sequences was generated using MEGA X [
42] with 1000 bootstrap replicates. Second, the whole genome-based phylogenetic tree was generated using the Type (Strain) Genome Server and inferred with FastME 2.1.6.1 [
43] from Genome BLAST Distance phylogeny approach (GBDP) distances calculated using genome sequences. The branch lengths were scaled in terms of GBDP distance formula D5, and the numbers above the branches were GBDP pseudo-bootstrap support values >60% from 100 replications. The regions and clusters of secondary metabolites present in the genomes of both the B1 and B3 strains and
B. velezensis CR-502
T (=NRRL B-41580
T) were predicted using antibiotics & Secondary Metabolite Analysis Shell (anti-SMASH) v.6.1.1 [
42] and compared.
2.5. Accession Numbers of Nucleotide Sequences and Strain Deposition
The 16S rRNA sequences of Bacillus B1 and B3 were deposited in GenBank under the accession numbers OP364972 and OP364977, respectively. The complete genome sequences of B1 and B3 were deposited in GenBank under the accession numbers CP100040 and CP100041, respectively.
2.6. Statistical Analysis
Survival data in the challenge test were analyzed via one-way analysis of variance (ANOVA) using SPSS version 24.0 (SPSS Inc., Chicago, IL, USA). The mean differences were compared using Duncan’s multiple range test when a significant difference was identified using ANOVA. For the comparison of means, the significance level was set at p ˂ 0.05. Data are presented as the mean ± SD, and the percentage data were arcsine-transformed before the comparisons.
4. Discussion
In this study, we evaluated the antimicrobial activity of five
Bacillus isolates against 12 shrimp
Vibrio strains (10 AHPND
Vibrio strains [9
V. parahaemolyticus and 1
V. campbellii] and 2 non-AHPND
Vibrio strains [1
V. parahaemolyticus and 1
V. harveyi]).
Bacillus spp. are usually isolated from soil, fermented soybean paste (cheonggukjang), plants, and pond water, and are incubated at 30–37 °C [
45,
46,
47]. The
Bacillus strains described in this study were isolated from seawater and were found to grow well at 28–37 °C. Additionally, all
Bacillus strains exhibited growth in both TSA and TSA+ (supplemented with 2% NaCl), indicating that these strains could be applied to water with wide ranges of salinity.
In the dot-spot test, B1, B3, B5, B7, and B8 exerted inhibitory effects on at least one of tested Vibrio strain. In addition, these strains showed inhibitory effects against isolates from both South Korea and several other countries (Mexico, Vietnam, Latin America, Thailand, and the USA). This indicates that the Bacillus strains used in this study can be used globally in various shrimp-farming countries to control AHPND. Management of AHPND, a disease which results in extensive mortality in shrimp, could increase shrimp production and decrease economic losses in shrimp farming.
In the challenge test, the B1 treatment group (100%) exhibited a significantly higher survival rate than the non-
Bacillus treatment group (64.3%) at 60 h. In a previous study by Han et al. [
48],
VcAHPND was highly pathogenic to shrimp, similar to
VpAHPND, and the accumulative mortality in shrimp was as high as 100% within 2 days of
VcAHPND laboratory infection. In this study, two
Bacillus strains (B1 and B3) displayed prominent antimicrobial effects within 2–3 days (48–60 h) of
VcAHPND infection compared with the findings in the positive control group (
VcAHPND immersion without B1 and B3 treatment); thus, both strains are expected to emerge as alternatives to antibiotics for controlling
VcAHPND. Moreover, among the live shrimp collected on the day of experiment termination, the
Ct value was higher in samples of the
Bacillus-treated groups than in the positive control group. Therefore, these results suggested that the two
Bacillus strains identified in this study exhibited antimicrobial activity against pathogenic
VcAHPND. Additionally, the histopathology of the hepatopancreas was examined after exposure to
Bacillus spp. for 14 days in our preliminary study. The structure of the hepatopancreas was found to be similar between the
Bacillus treatment groups and the control group (not exposed to
VcAHPND and
Bacillus), indicating that
Bacillus strains are harmless to shrimp.
The two strains (B1 and B3), which showed antimicrobial activity using the dot-spot test (in vitro) and challenge test, were finally classified as
B. velezensis based on their whole genome-based phylogeny. Several studies have examined the probiotic effects of
B. velezensis in various organisms. For example, Chauyod et al. [
49] demonstrated that
B. velezensis significantly inhibited the growth of
Vibrio spp. isolated from shrimp, including
VpAHPND, using the disk diffusion test. Li et al. [
50] reported that the expression of immune-related genes such as IL-8 and IgM was upregulated in the hybrid grouper fed a feed supplemented with
B. velezensis (1 × 10
7 CFU g
−1) compared with the findings in the control group, and the former also exhibited increased resistance to
V. harveyi. Other studies described the antibacterial activity of
B. velezensis against
V. parahaemolyticus isolated from shrimp [
51] and
V. anguillarum isolated from seabass [
52]. Although the predicted secondary metabolites derived from the B1 and B3
Bacillus strains were relatively similar to those previously reported from related
Bacillus species [
39], plipastatin and surfactin were only found in the two isolates, and they were not detected during our in silico analysis of the type strain of
B. velezensis. These results suggest that the newly isolated B1 and B3 strains will have additional advantageous characteristics in terms of their potential use in the aquaculture industry. Till date, most previously reported secondary metabolites produced by
Bacillus spp. were known to have surfactant and antibiotic activity [
53]. In particular, the potential presence of surfactin, which was previously reportedly associated with antibacterial activity against multidrug-resistant
Vibrio spp. [
54] in the two
Bacillus strains might explain their antimicrobial activity against pathogenic
VcAHPND in this study; however, further studies are warranted regarding the predicted presence of surfactin in the isolates because of its relatively low similarity with previously reported compounds. Moreover, the potential presence of iturin and fengycin, which have been associated with the antifungal activity of some
Bacillus strains [
55], may contribute to the potential usability of the
Bacillus strains identified in this study.