Anticoagulant Oligonucleotide–Peptide Conjugates: Identification of Thrombin Aptamer Conjugates with Improved Characteristics
Abstract
1. Introduction
2. Results
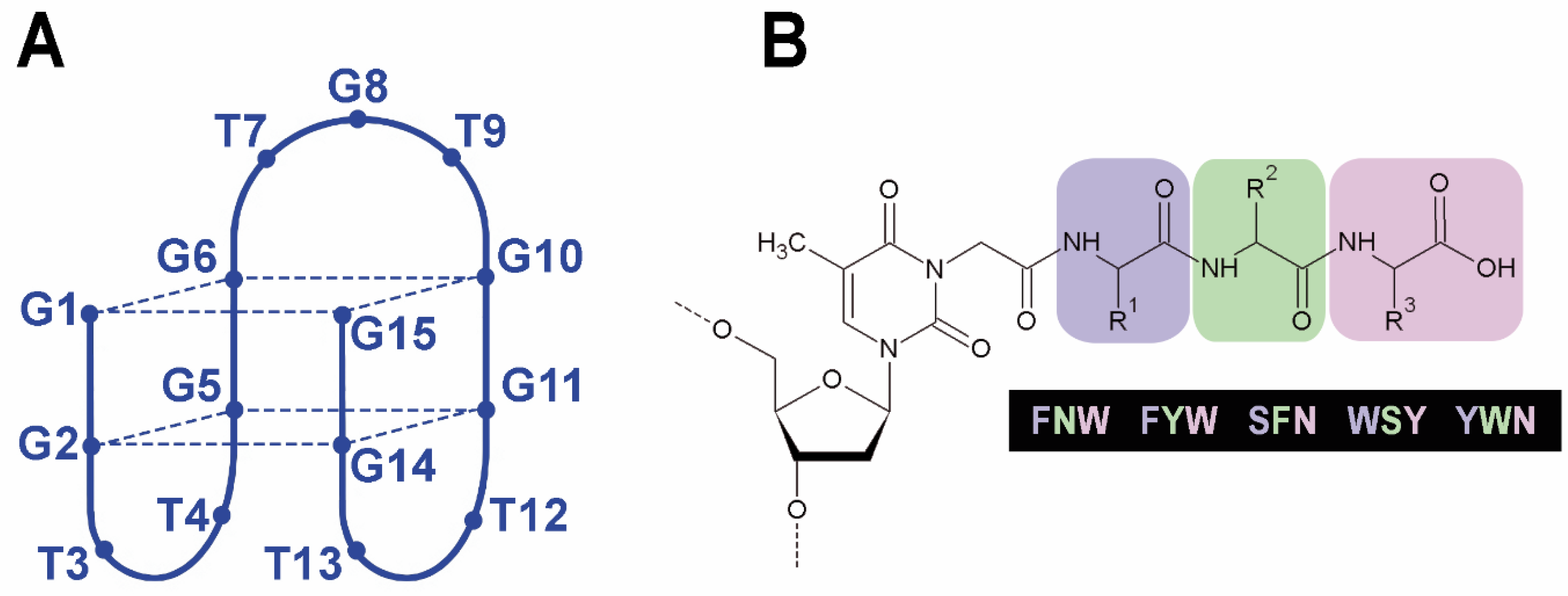
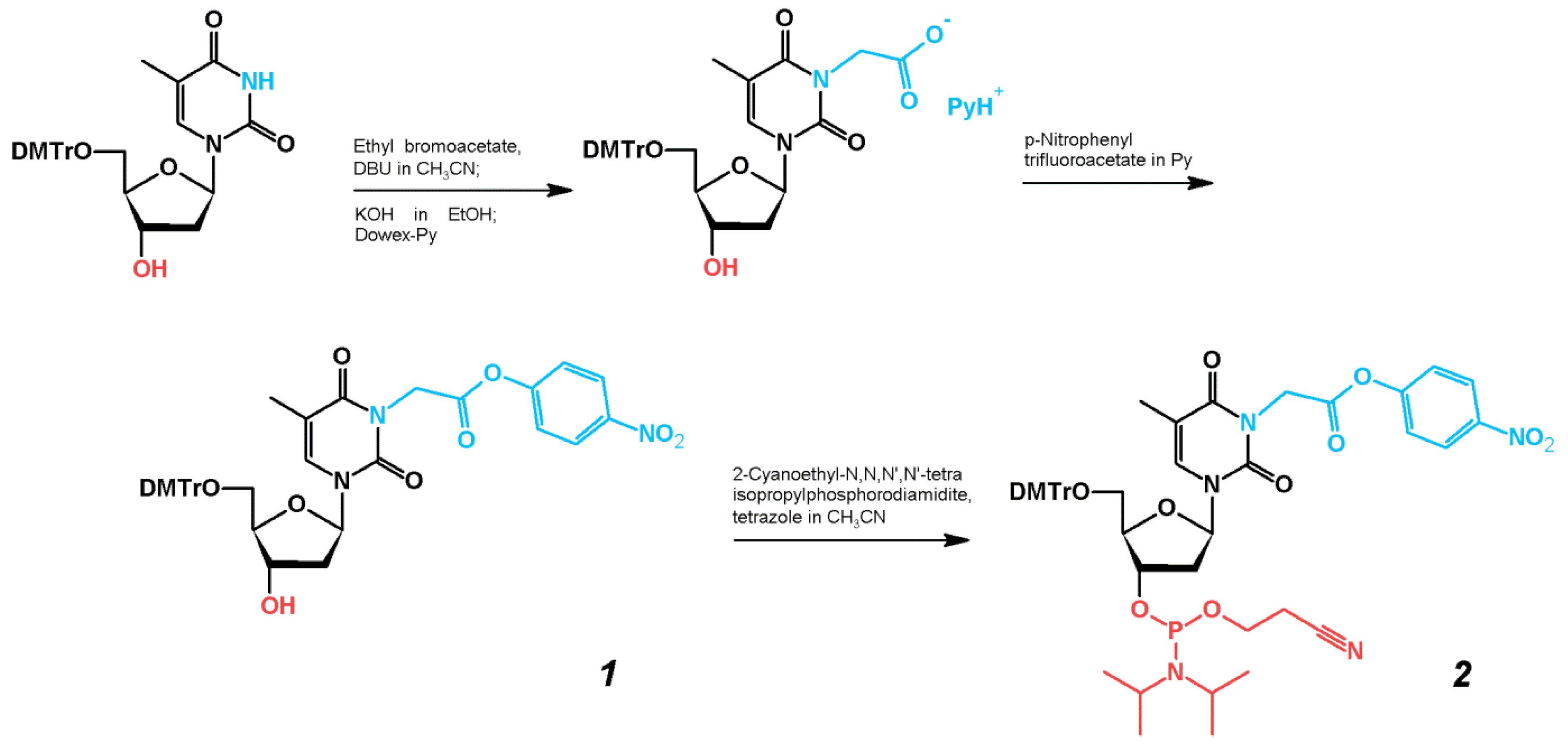
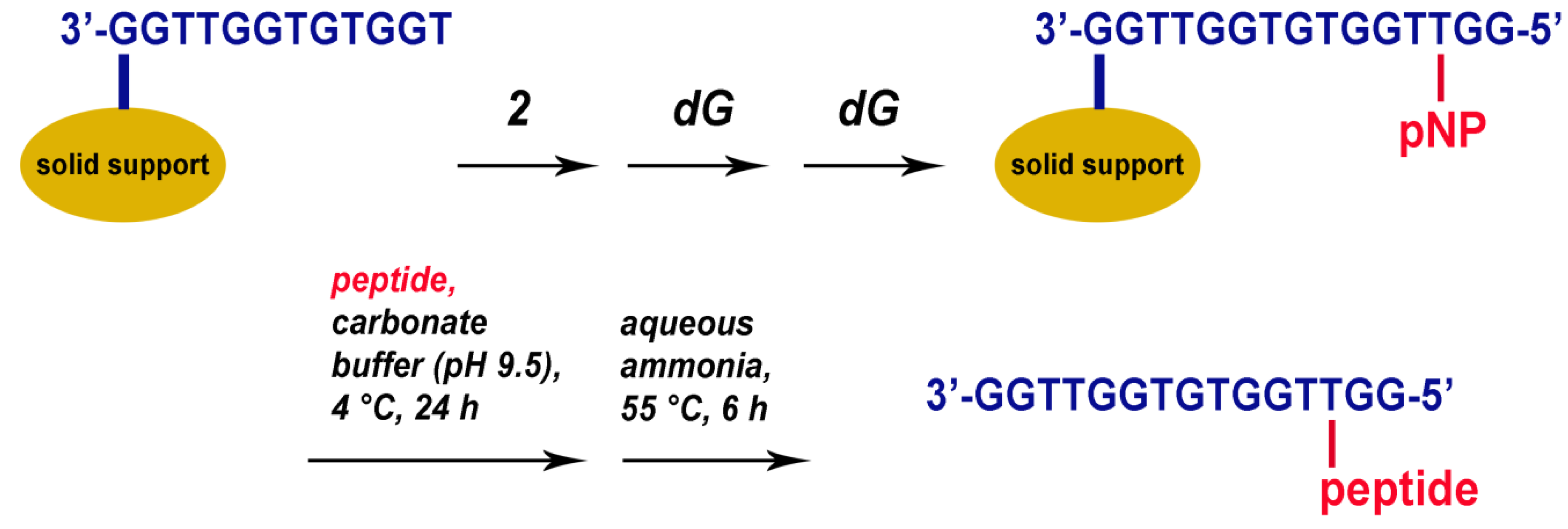
2.1. Synthesis of TBA–Peptide Conjugates
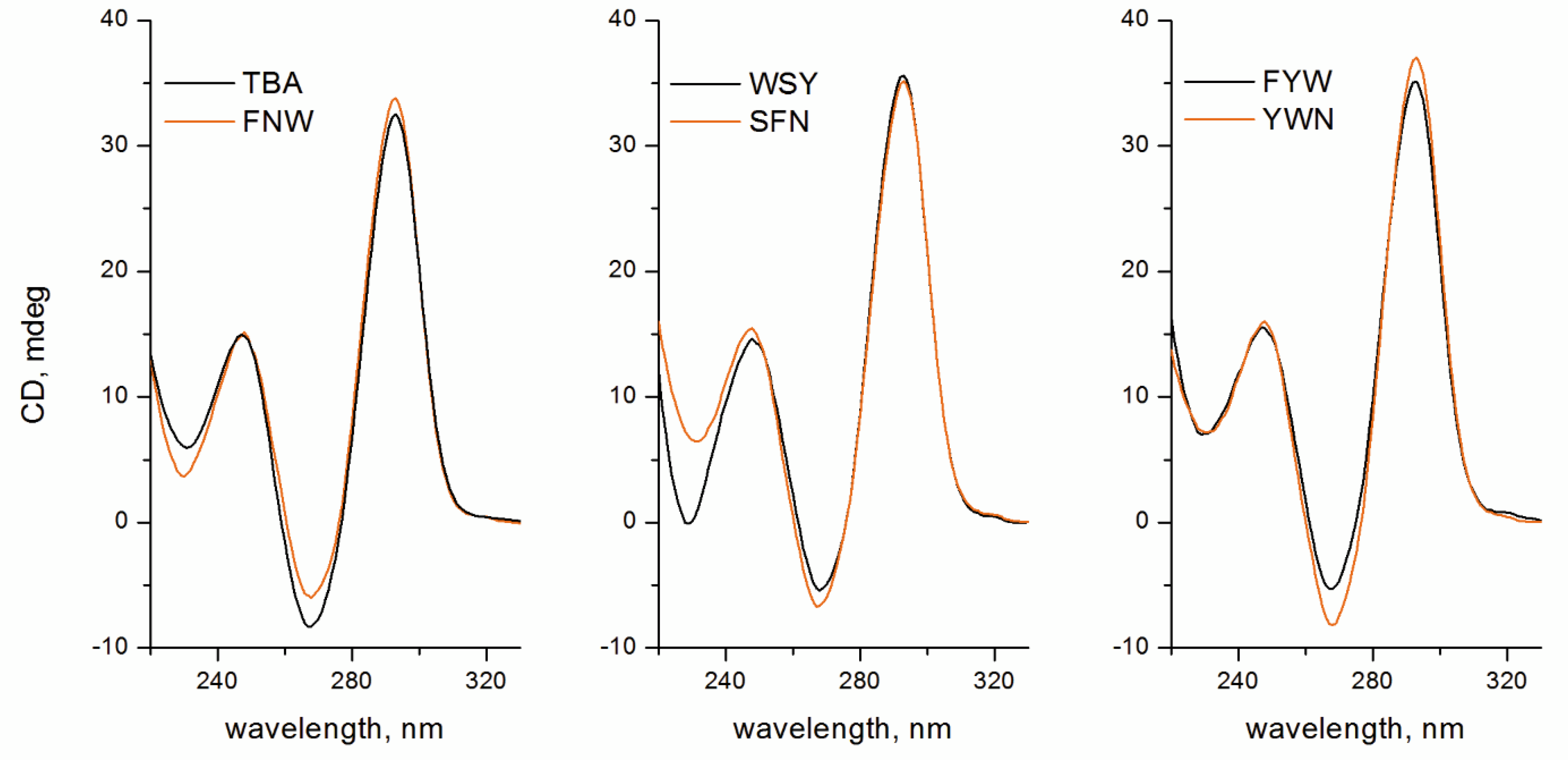
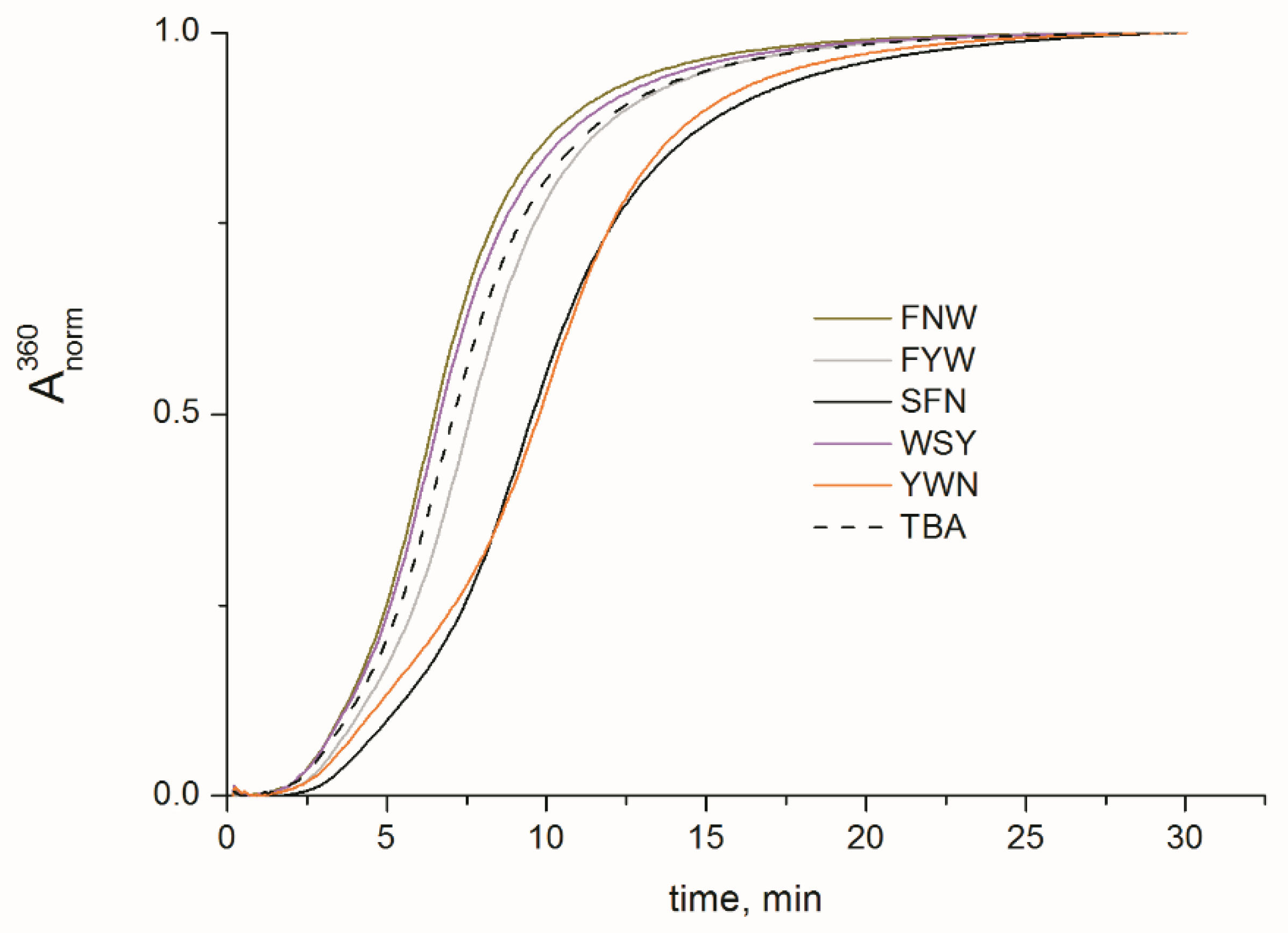
2.2. Folding, Stability, and Anticoagulant Activity of TBA–Peptide Conjugates
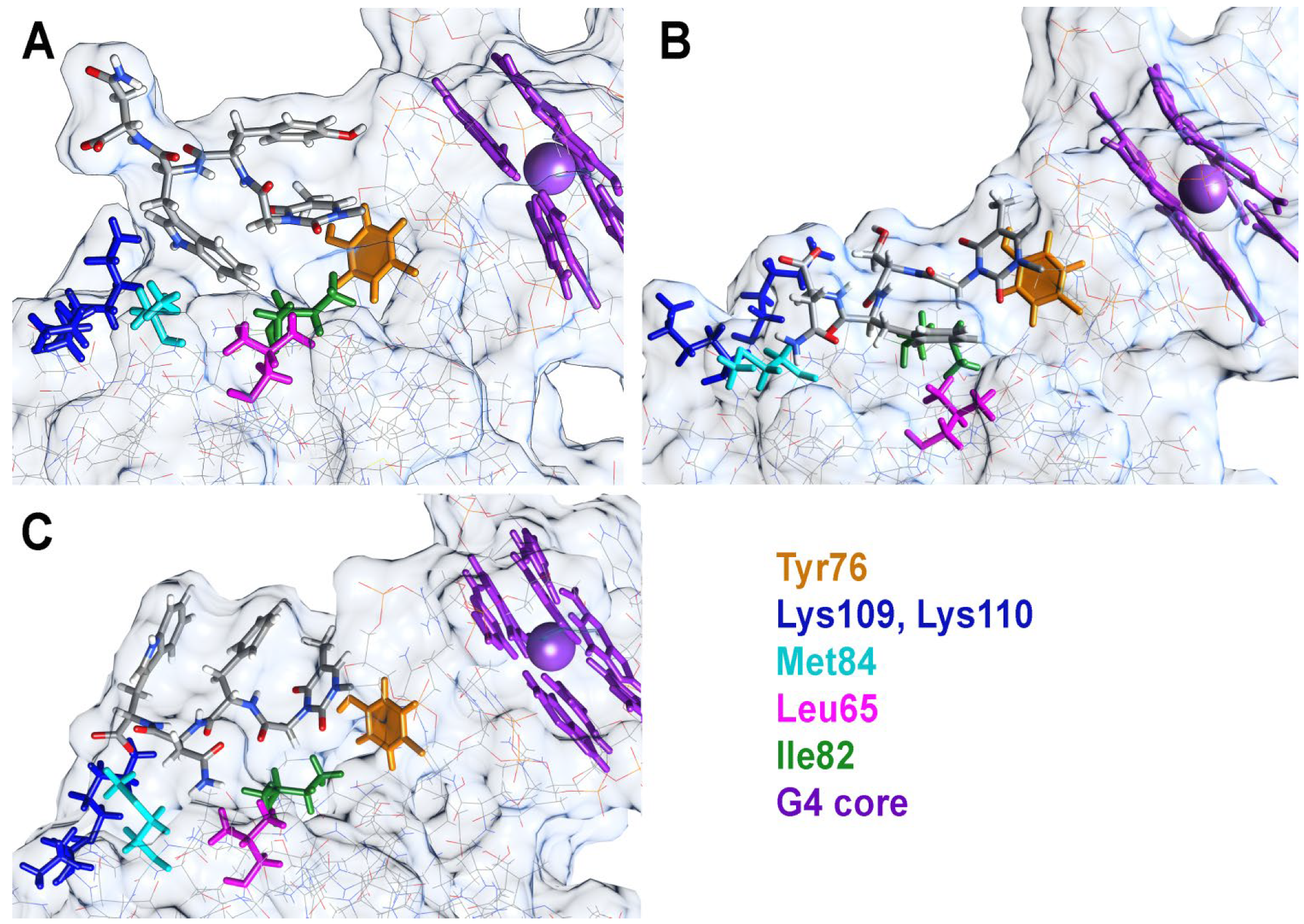
2.3. Molecular Dynamics Studies of TBA OPC Complexes with Thrombin
3. Materials and Methods
3.1. Materials and Instrumentation
3.2. Syntheses of Compounds 1 and 2
3.3. Automated Oligonucleotide Synthesis and Conjugation
3.4. Ultraviolet Thermal Denaturation
3.5. Circular Dichroism Spectroscopy
3.6. Fibrinogen Clotting in the Presence of Aptamers
3.7. Molecular Dynamics Simulations of TBA–OPC Complexes with Thrombin
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hoyer, J.; Neundorf, I. Peptide vectors for the nonviral delivery of nucleic acids. Acc. Chem. Res. 2012, 45, 1048–1056. [Google Scholar] [CrossRef]
- Taskova, M.; Mantsiou, A.; Astakhova, K. Synthetic Nucleic Acid Analogues in Gene Therapy: An Update for Peptide-Oligonucleotide Conjugates. ChemBioChem 2017, 18, 1671–1682. [Google Scholar] [CrossRef] [PubMed]
- Gait, M.J.; Arzumanov, A.A.; McClorey, G.; Godfrey, C.; Betts, C.; Hammond, S.; Wood, M. Cell-Penetrating Peptide Conjugates of Steric Blocking Oligonucleotides as Therapeutics for Neuromuscular Diseases from a Historical Perspective to Current Prospects of Treatment. Nucleic Acid Ther. 2019, 29, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Klabenkova, K.; Fokina, A.; Stetsenko, D. Chemistry of Peptide-Oligonucleotide Conjugates: A Review. Molecules 2021, 26, 5420. [Google Scholar] [CrossRef] [PubMed]
- Hammond, S.M.; Aartsma-Rus, A.; Alves, S.; Borgos, S.E.; Buijsen, R.A.M.; Collin, R.W.J.; Covello, G.; Denti, M.A.; Desviat, L.R.; Echevarría, L.; et al. Delivery of oligonucleotide-based therapeutics: Challenges and opportunities. EMBO Mol. Med. 2021, 13, e13243. [Google Scholar] [CrossRef]
- Smirnov, I.; Kolganova, N.; Troisi, R.; Sica, F.; Timofeev, E. Expanding the recognition interface of the thrombin-binding aptamer HD1 through modification of residues T3 and T12. Mol. Ther. Nucleic Acids 2021, 23, 863–871. [Google Scholar] [CrossRef] [PubMed]
- Kysilka, J.; Vondrášek, J. Towards a better understanding of the specificity of protein-protein interaction. J. Mol. Recognit. 2012, 25, 604–615. [Google Scholar] [CrossRef] [PubMed]
- Lafont, V.; Schaefer, M.; Stote, R.H.; Altschuh, D.; Dejaegere, A. Protein-protein recognition and interaction hot spots in an antigen-antibody complex: Free energy decomposition identifies “efficient amino acids”. Proteins 2007, 67, 418–434. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Nussinov, R. Trp/Met/Phe hot spots in protein-protein interactions: Potential targets in drug design. Curr. Top. Med. Chem. 2007, 7, 999–1005. [Google Scholar] [CrossRef] [PubMed]
- Ofran, Y.; Rost, B. Protein-protein interaction hotspots carved into sequences. PLoS Comput. Biol. 2007, 3, e119. [Google Scholar] [CrossRef]
- Chen, C.T.; Peng, H.P.; Jian, J.W.; Tsai, K.C.; Chang, J.Y.; Yang, E.W.; Chen, J.B.; Ho, S.Y.; Hsu, W.L.; Yang, A.S. Protein-protein interaction site predictions with three-dimensional probability distributions of interacting atoms on protein surfaces. PLoS ONE 2012, 7, e37706. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Smith, J.K.; Jiang, S.; Pfaendtner, J. Redefining the Protein-Protein Interface: Coarse Graining and Combinatorics for an Improved Understanding of Amino Acid Contributions to the Protein-Protein Binding Affinity. Langmuir 2017, 33, 11511–11517. [Google Scholar] [CrossRef]
- Gallet, X.; Charloteaux, B.; Thomas, A.; Brasseur, R. A fast method to predict protein interaction sites from sequences. J. Mol. Biol. 2000, 302, 917–926. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Martinez, D.; Platero, H.; Lane, T.; Werner-Washburne, M. Exploiting amino acid composition for predicting protein-protein interactions. PLoS ONE 2009, 4, e7813. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Duan, Q.P.; Ma, L.; Zhao, D.X. Chemical strategies for the synthesis of peptide-oligonucleotide conjugates. Bioconjug. Chem. 2010, 21, 187–202. [Google Scholar] [CrossRef] [PubMed]
- Dastpeyman, M.; Karas, J.A.; Amin, A.; Turner, B.J.; Shabanpoor, F. Modular Synthesis of Trifunctional Peptide-oligonucleotide Conjugates via Native Chemical Ligation. Front. Chem. 2021, 9, 627329. [Google Scholar] [CrossRef] [PubMed]
- Smirnov, I.P.; Kolganova, N.A.; Vasiliskov, V.A.; Chudinov, A.V.; Timofeev, E.N. Mass-spectrometry analysis of modifications at DNA termini induced by DNA polymerases. Sci. Rep. 2017, 7, 6674. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sakakibara, S.; Inukai, N. The trifluoroacetate method of peptide synthesis. I. The synthesis and use of trifluoroacetate reagents. Bull. Chem. Soc. Jpn. 1965, 38, 1979–1984. [Google Scholar] [CrossRef] [PubMed]
- Zeskind, B.J.; Jordan, C.D.; Timp, W.; Trapani, L.; Waller, G.; Horodincu, V.; Ehrlich, D.J.; Matsudaira, P. Nucleic acid and protein mass mapping by live-cell deep-ultraviolet microscopy. Nat. Methods. 2007, 4, 567–569. [Google Scholar] [CrossRef] [PubMed]
- Abagyan, R.; Totrov, M.; Kuznetsov, D. ICM, a new method for protein modeling and design: Applications to docking and structure prediction from the distorted native conformation. J. Comput. Chem. 1994, 15, 488–506. [Google Scholar] [CrossRef]
- Arnautova, Y.A.; Jagielska, A.; Scheraga, H.A. A new force field (ECEPP-05) for peptides, proteins, and organic molecules. J. Phys. Chem. B 2006, 110, 5025–5044. [Google Scholar] [CrossRef] [PubMed]
- Case, D.A.; Belfon, K.; Ben-Shalom, I.Y.; Brozell, S.R.; Cerutti, D.S.; Cheatham, T.E., III; Cruzeiro, V.W.D.; Darden, T.A.; Duke, R.E.; Giambasu, G.; et al. AMBER 2020; University of California: San Francisco, CA, USA, 2020. [Google Scholar]
- Izadi, S.; Onufriev, A.V. Accuracy limit of rigid 3-point water models. J. Chem. Phys. 2016, 145, 074501. [Google Scholar] [CrossRef] [PubMed]
- Zgarbová, M.; Luque, F.J.; Sponer, J.; Cheatham, T.E., III; Otyepka, M.; Jurečka, P. Toward Improved Description of DNA Backbone: Revisiting Epsilon and Zeta Torsion Force Field Parameters. J. Chem. Theory Comput. 2013, 9, 2339–2354. [Google Scholar] [CrossRef] [PubMed]
- Zgarbová, M.; Sponer, J.; Otyepka, M.; Cheatham, T.E., III; Galindo-Murillo, R.; Jurečka, P. Refinement of the Sugar-Phosphate Backbone Torsion Beta for AMBER Force Fields Improves the Description of Z- and B-DNA. J. Chem. Theory Comput. 2015, 11, 5723–5736. [Google Scholar] [CrossRef] [PubMed]
- Maier, J.A.; Martinez, C.; Kasavajhala, K.; Wickstrom, L.; Hauser, K.E.; Simmerling, C. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 2015, 11, 3696–3713. [Google Scholar] [CrossRef] [PubMed]
| TBA OPC | Mcalc/Mobsvd | Y a, % | Tm, °C b | t1/2, min c |
|---|---|---|---|---|
| Unmodified TBA | 4726.1/4721.9 | n/a | 52.5 | 7.1 |
| TBA–SFN | 5132.8/5138.5 | 47 | 54.2 | 9.6 |
| TBA–FYW | 5280.7/5284.0 | 66 | 57.9 | 7.6 |
| TBA–FNW | 5231.6/5241.6 | 63 | 57.5 | 6.5 |
| TBA–YWN | 5247.6/5250.1 | 72 | 56.8 | 9.8 |
| TBA–WSY | 5220.6/5222.8 | 70 | 54.9 | 6.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tsvetkov, V.B.; Varizhuk, I.V.; Kurochkin, N.N.; Surzhikov, S.A.; Smirnov, I.P.; Stomakhin, A.A.; Kolganova, N.A.; Timofeev, E.N. Anticoagulant Oligonucleotide–Peptide Conjugates: Identification of Thrombin Aptamer Conjugates with Improved Characteristics. Int. J. Mol. Sci. 2022, 23, 3820. https://doi.org/10.3390/ijms23073820
Tsvetkov VB, Varizhuk IV, Kurochkin NN, Surzhikov SA, Smirnov IP, Stomakhin AA, Kolganova NA, Timofeev EN. Anticoagulant Oligonucleotide–Peptide Conjugates: Identification of Thrombin Aptamer Conjugates with Improved Characteristics. International Journal of Molecular Sciences. 2022; 23(7):3820. https://doi.org/10.3390/ijms23073820
Chicago/Turabian StyleTsvetkov, Vladimir B., Irina V. Varizhuk, Nikolay N. Kurochkin, Sergei A. Surzhikov, Igor P. Smirnov, Andrey A. Stomakhin, Natalia A. Kolganova, and Edward N. Timofeev. 2022. "Anticoagulant Oligonucleotide–Peptide Conjugates: Identification of Thrombin Aptamer Conjugates with Improved Characteristics" International Journal of Molecular Sciences 23, no. 7: 3820. https://doi.org/10.3390/ijms23073820
APA StyleTsvetkov, V. B., Varizhuk, I. V., Kurochkin, N. N., Surzhikov, S. A., Smirnov, I. P., Stomakhin, A. A., Kolganova, N. A., & Timofeev, E. N. (2022). Anticoagulant Oligonucleotide–Peptide Conjugates: Identification of Thrombin Aptamer Conjugates with Improved Characteristics. International Journal of Molecular Sciences, 23(7), 3820. https://doi.org/10.3390/ijms23073820







