Unveiling the Role of SlRNC1 in Chloroplast Development and Global Gene Regulation in Tomato Plants
Abstract
1. Introduction
2. Results
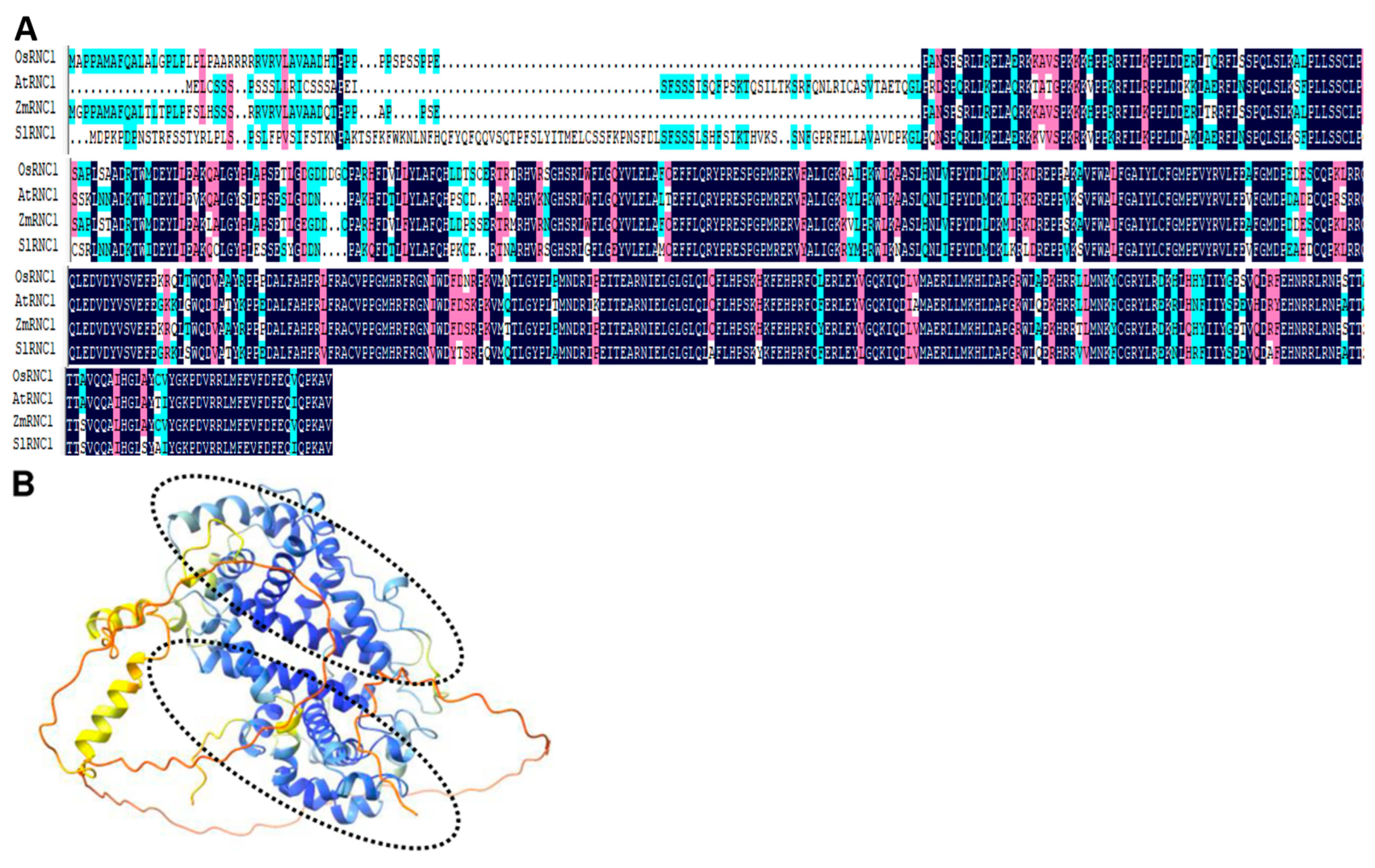
2.1. Identification of RNC1 in Tomato and Sequence Alignment RNC1 Proteins across Four Plants
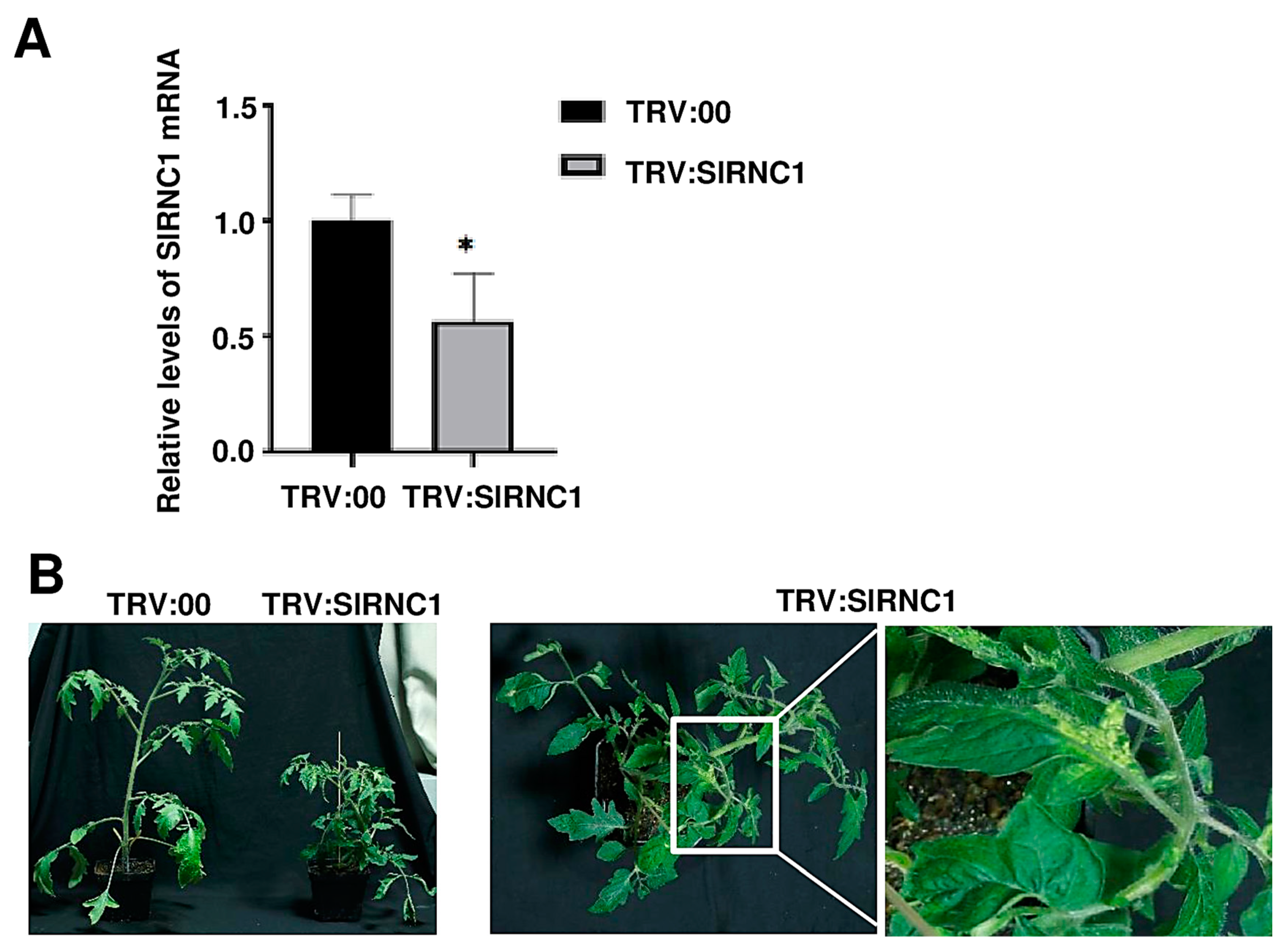
2.2. VIGS-Induced SlRNC1 Silencing Results in Dwarf Tomato Plants with Yellowish Leaves
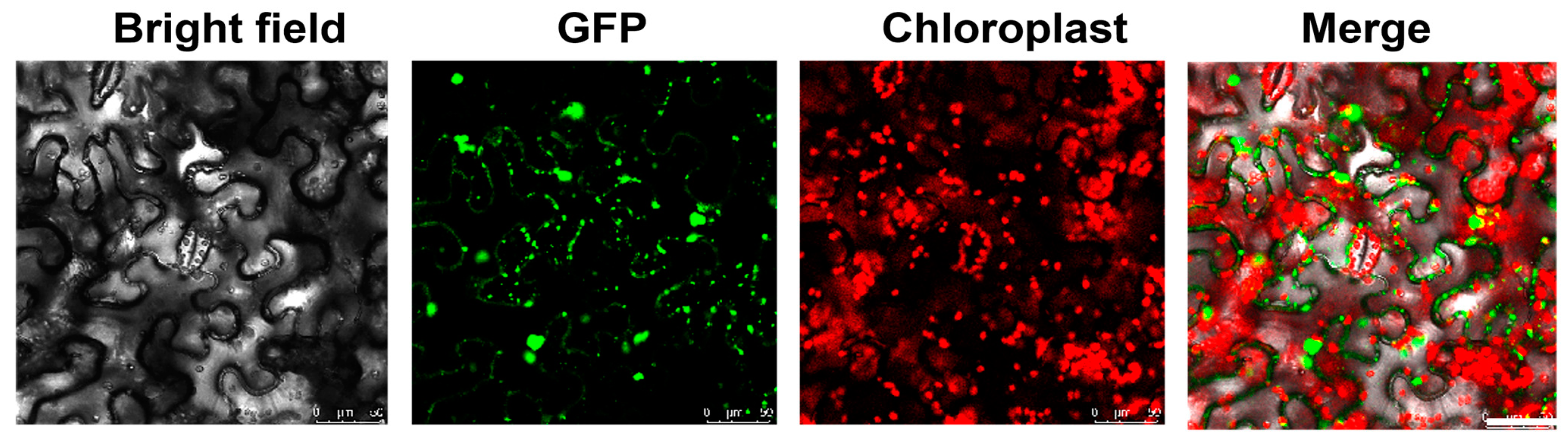
2.3. Subcellular Localization of SlRNC1
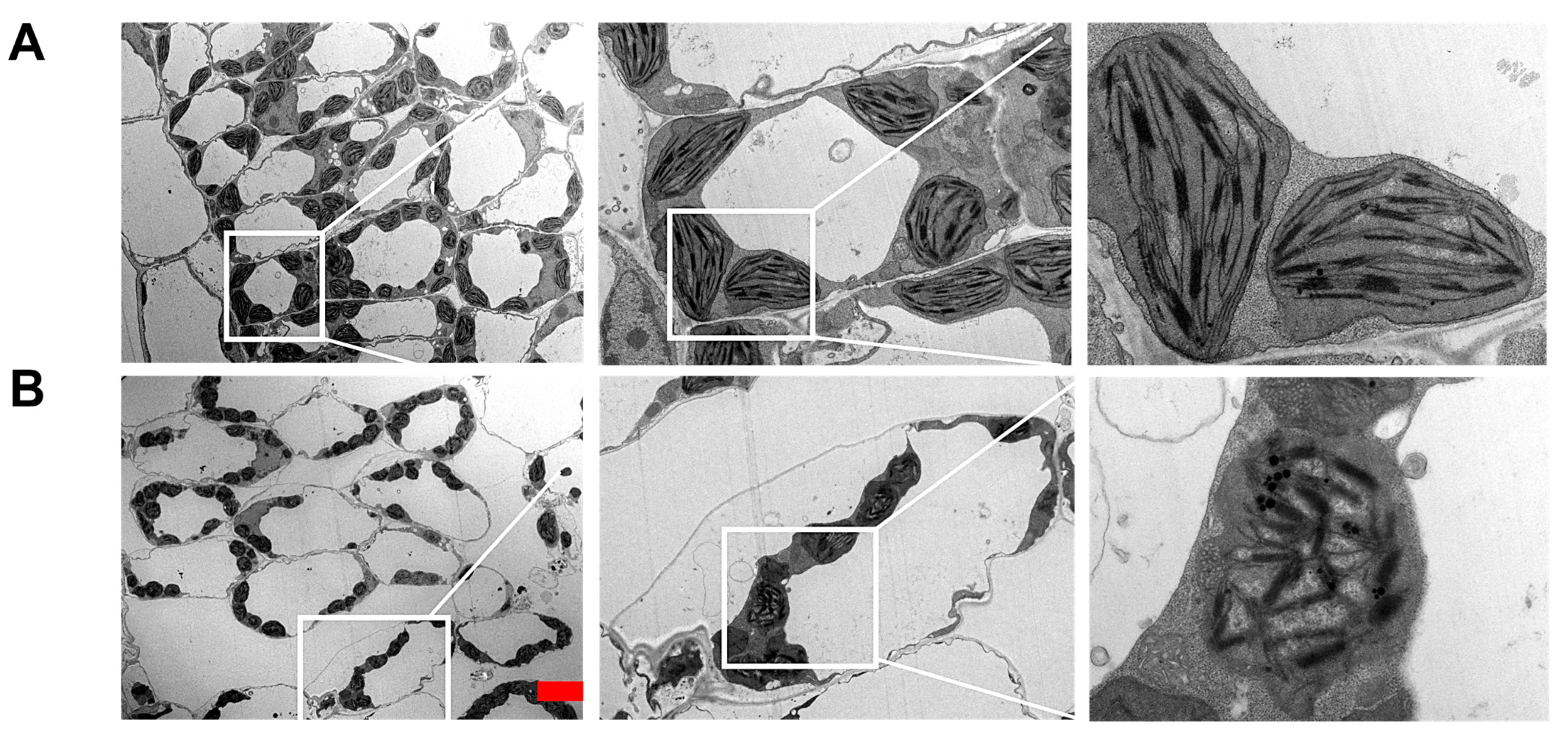
2.4. Silencing of SlRNC1 Caused Ultrastructural Changes in Chloroplasts
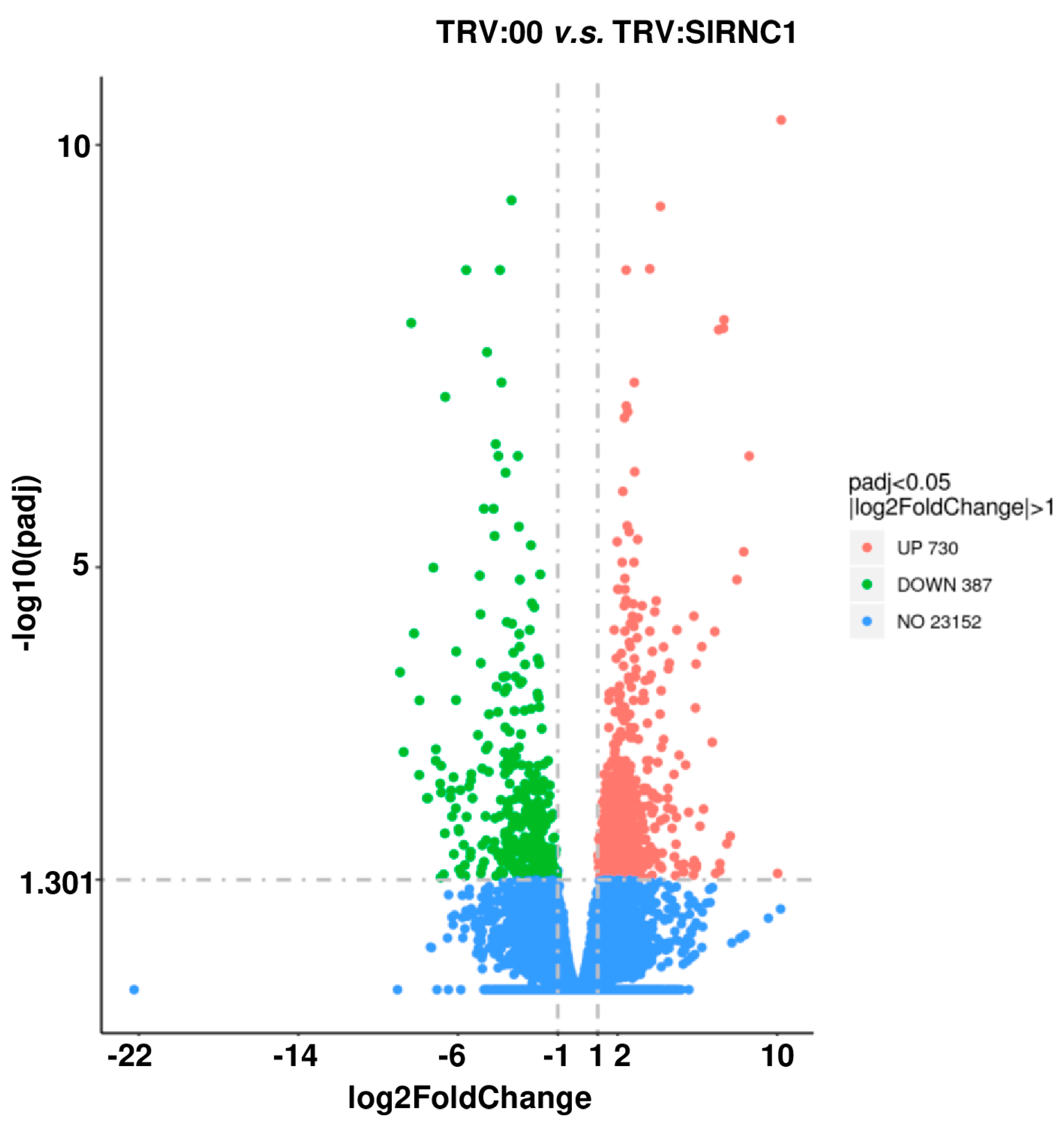
2.5. Analysis of Differentially Expressed Genes Caused by SlRNC1 Silencing
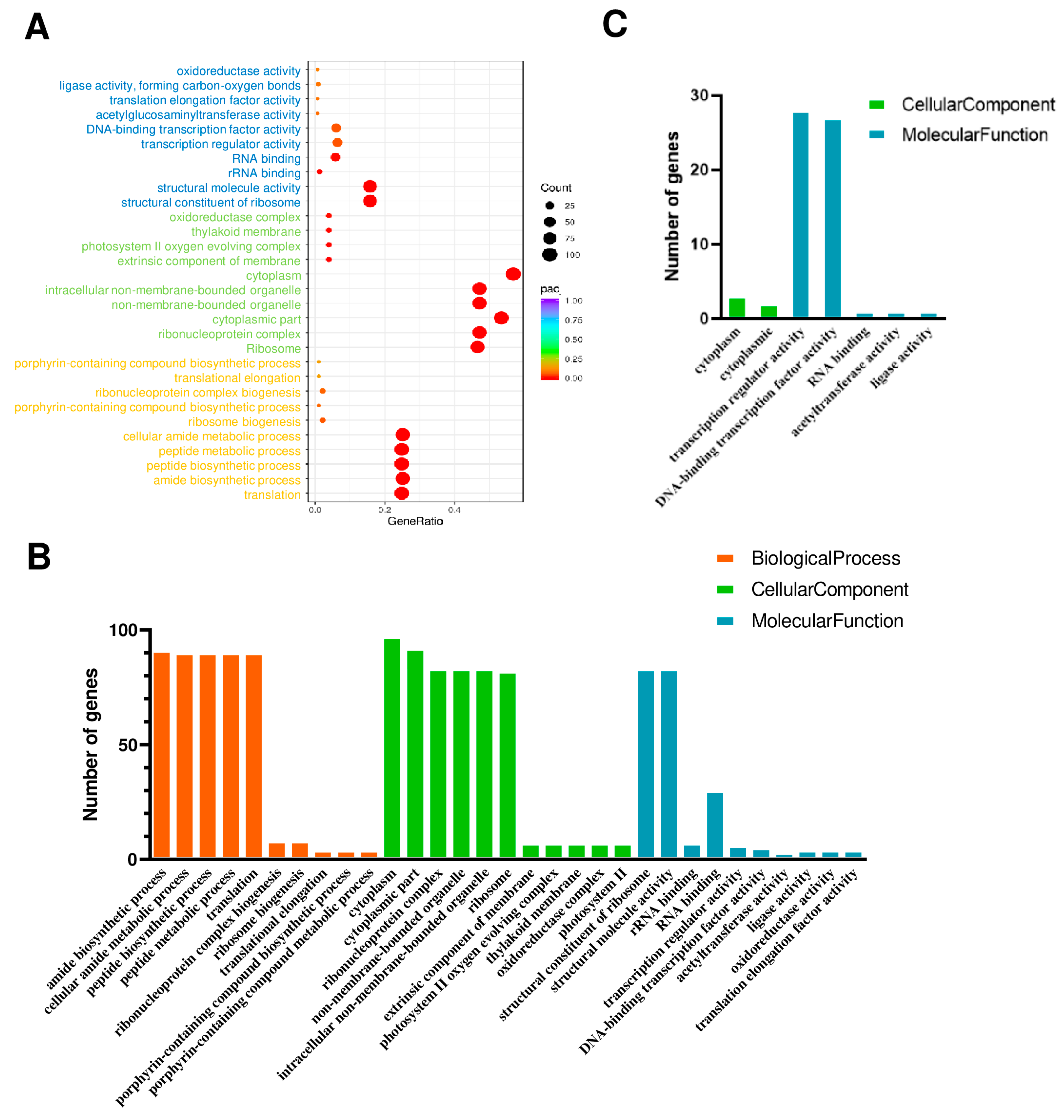
2.6. Gene Ontology (GO) and KEGG Analysis of DEGs
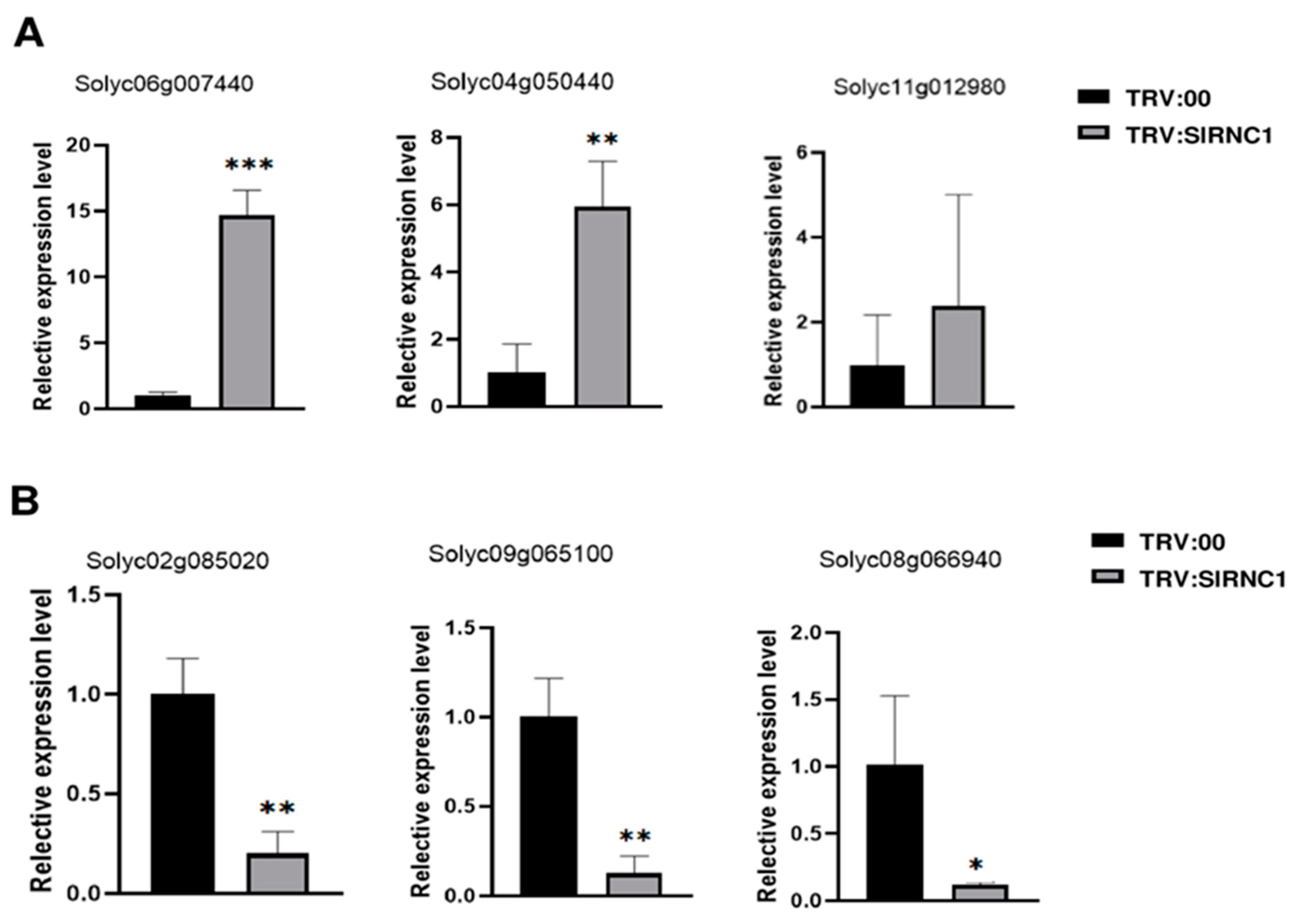
2.7. qRT-PCR Confirmation of DEGs
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Agrobacterium Transformation-Mediated Transient Gene Expression
4.3. RNA Extraction, RT-PCR, and RT-qPCR
4.4. Plasmid Construction and Inoculation
4.5. SlRNC1 Gene Identification, Sequence Alignment, and Protein Structure Modeling
4.6. Subcellular Localization Analysis and TEM Analysis
4.7. RNA-Seq Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bowles, A.M.; Williamson, C.J.; Williams, T.A.; Lenton, T.M.; Donoghue, P.C. The origin and early evolution of plants. Trends Plant Sci. 2023, 28, 312–329. [Google Scholar] [CrossRef]
- Ádám, A.L.; Nagy, Z.Á.; Kátay, G.; Mergenthaler, E.; Viczián, O. Signals of systemic immunity in plants: Progress and open questions. Int. J. Mol. Sci. 2018, 19, 1146. [Google Scholar] [CrossRef] [PubMed]
- Azim, M.F.; Burch-Smith, T.M. Organelles-nucleus-plasmodesmata signaling (ONPS): An update on its roles in plant physiology, metabolism and stress responses. Curr. Opin. Plant Biol. 2020, 58, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Mamaeva, A.; Taliansky, M.; Filippova, A.; Love, A.J.; Golub, N.; Fesenko, I. The role of chloroplast protein remodeling in stress responses and shaping of the plant peptidome. New Phytol. 2020, 227, 1326–1334. [Google Scholar] [CrossRef] [PubMed]
- Cackett, L.; Luginbuehl, L.H.; Schreier, T.B.; Lopez-Juez, E.; Hibberd, J.M. Chloroplast development in green plant tissues: The interplay between light, hormone, and transcriptional regulation. New Phytol. 2022, 233, 2000–2016. [Google Scholar] [CrossRef] [PubMed]
- Kachroo, P.; Burch-Smith, T.M.; Grant, M. An emerging role for chloroplasts in disease and defense. Annu. Rev. Phytopathol. 2021, 59, 423–445. [Google Scholar] [CrossRef] [PubMed]
- Medina-Puche, L.; Tan, H.; Dogra, V.; Wu, M.; Rosas-Diaz, T.; Wang, L.; Ding, X.; Zhang, D.; Fu, X.; Kim, C.; et al. A defense pathway linking plasma membrane and chloroplasts and co-opted by pathogens. Cell 2020, 182, 1109–1124.e25. [Google Scholar] [CrossRef] [PubMed]
- Han, K.; Jia, Z.; Zhang, Y.; Zhou, H.; Bu, S.; Chen, J.; Yan, D.; Qi, R.; Yan, F.; Wu, J. Chloroplast clustering around the nucleus induced by OMP24 overexpression unexpectedly promoted PSTVd infection in Nicotiana benthamiana. Mol. Plant Pathol. 2023, 24, 1552–1559. [Google Scholar] [PubMed]
- Han, K.; Zheng, H.; Yan, D.; Zhou, H.; Jia, Z.; Zhai, Y.; Wu, J.; Lu, Y.; Wu, G.; Rao, S.; et al. Pepper mild mottle virus coat protein interacts with pepper chloroplast outer envelope membrane protein OMP24 to inhibit antiviral immunity in plants. Hortic. Res. 2023, 10, uhad046. [Google Scholar] [CrossRef]
- de Longevialle, A.F.; Small, I.D.; Lurin, C. Nuclearly encoded splicing factors implicated in RNA splicing in higher plant organelles. Mol. Plant 2010, 3, 691–705. [Google Scholar] [CrossRef]
- Bonen, L.; Vogel, J. The ins and outs of group II introns. Trends Genet. 2001, 17, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Jan, M.; Liu, Z.; Rochaix, J.-D.; Sun, X. Retrograde and anterograde signaling in the crosstalk between chloroplast and nucleus. Front. Plant Sci. 2022, 13, 980237. [Google Scholar] [CrossRef] [PubMed]
- Mullineaux, P.M.; Exposito-Rodriguez, M.; Laissue, P.P.; Smirnoff, N.; Park, E. Spatial chloroplast-to-nucleus signalling involving plastid–nuclear complexes and stromules. Philos. Trans. R. Soc. B 2020, 375, 20190405. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, Y.; Zhu, X.; Ren, Y.; Dong, H.; Duan, E.; Teng, X.; Zhao, H.; Chen, R.; Chen, X.; et al. Tetrapyrrole biosynthesis pathway regulates plastid-to-nucleus signaling by controlling plastid gene expression in plants. Plant Commun. 2023, 4, 100411. [Google Scholar] [CrossRef] [PubMed]
- Small, I.; Melonek, J.; Bohne, A.-V.; Nickelsen, J.; Schmitz-Linneweber, C. Plant organellar RNA maturation. Plant Cell 2023, 35, 1727–1751. [Google Scholar] [CrossRef] [PubMed]
- Dobrogojski, J.; Adamiec, M.; Luciński, R. The chloroplast genome: A review. Acta Physiol. Plant. 2020, 42, 98. [Google Scholar] [CrossRef]
- Watkins, K.P.; Kroeger, T.S.; Cooke, A.M.; Williams-Carrier, R.E.; Friso, G.; Belcher, S.E.; Van Wijk, K.J.; Barkan, A. A ribonuclease III domain protein functions in group II intron splicing in maize chloroplasts. Plant Cell 2007, 19, 2606–2623. [Google Scholar] [CrossRef] [PubMed]
- Kroeger, T.S.; Watkins, K.P.; Friso, G.; van Wijk, K.J.; Barkan, A. A plant-specific RNA-binding domain revealed through analysis of chloroplast group II intron splicing. Proc. Natl. Acad. Sci. USA 2009, 106, 4537–4542. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Li, J.; Jia, J.; Yue, L.; Dong, X. Comprehensive analysis of the pre-ribosomal RNA maturation pathway in a methanoarchaeon exposes the conserved circularization and linearization mode in archaea. RNA Biol. 2020, 17, 1427–1441. [Google Scholar] [CrossRef]
- Gawroński, P.; Burdiak, P.; Scharff, L.B.; Mielecki, J.; Górecka, M.; Zaborowska, M.; Leister, D.; Waszczak, C.; Karpiński, S. CIA2 and CIA2-LIKE are required for optimal photosynthesis and stress responses in Arabidopsis thaliana. Plant J. 2021, 10, 619–638. [Google Scholar] [CrossRef]
- Kleine, T.; Leister, D. Retrograde signaling: Organelles go networking. Biochim. Biophys. Acta 2016, 185, 1313–1325. [Google Scholar] [CrossRef] [PubMed]
- Fernández, A.P.; Strand, A. Retrograde signaling and plant stress: Plastid signals initiate cellular stress responses. Curr. Opin. Plant Biol. 2008, 1, 509–513. [Google Scholar] [CrossRef]
- Satoh, R.; Hara, N.; Kawasaki, A.; Takasaki, T.; Sugiura, R. Distinct modes of stress granule assembly mediated by the KH-type RNA-binding protein Rnc1. Genes Cells 2018, 23, 778–785. [Google Scholar] [CrossRef]
- Lin, D.; Kong, R.; Chen, L.; Wang, Y.; Wu, L.; Xu, J.; Piao, Z.; Lee, G.; Dong, Y. Chloroplast development at low temperature requires the pseudouridine synthase gene TCD3 in rice. Sci. Rep. 2020, 10, 8518. [Google Scholar] [CrossRef]
- Griffin, J.H.C.; Prado, K.; Sutton, P.; Toledo-Ortiz, G. Coordinating light responses between the nucleus and the chloroplast, a role for plant cryptochromes and phytochromes. Physiol. Plant. 2020, 169, 515–528. [Google Scholar] [CrossRef]
- Zhou, J.; Liu, H.; Lin, Y.; Zhao, J. Membrane occupation and recognition nexus (MORN) motif controls protein localization and function. FEBS Lett. 2022, 596, 1839–1850. [Google Scholar] [CrossRef]
- Bhattacharya, O.; Ortiz, I.; Hendricks, N.; Walling, L.L. The tomato chloroplast stromal proteome compendium elucidated by leveraging a plastid protein-localization prediction Atlas. Front. Plant Sci. 2023, 14, 1020275. [Google Scholar] [CrossRef]
- Schmid, L.M.; Manavski, N.; Chi, W.; Meurer, J. Chloroplast Ribosome Biogenesis Factors. Plant Cell Physiol. 2024, 65, 516–536. [Google Scholar] [CrossRef]
- Chen Y, Zhou QX, Tao ZX; et al. Huan Jing Ke Xue. 2024, 45, 3446–3458.
- Wang, X.; Liu, Y.; Ouyang, L.; Yao, R.; Yu, T.; Yan, L.; Chen, Y.; Huai, D.; Zhou, X.; Wang, Z.; et al. Full-length transcriptome sequencing provides insights into alternative splicing under cold stress in peanut. Front. Plant Sci. 2024, 15, 1362277. [Google Scholar] [CrossRef] [PubMed]
- Chantarachot, T.; Bailey-Serres, J. Polysomes, stress granules, and processing bodies: A dynamic triumvirate controlling cytoplasmic mRNA fate and function. Plant Physiol. 2018, 176, 254–269. [Google Scholar] [CrossRef] [PubMed]
- Ivanov, P.; Kedersha, N.; Anderson, P. Stress granules and processing bodies in translational control. Cold Spring Harb. Perspect. Biol. 2019, 11, a032813. [Google Scholar] [CrossRef] [PubMed]
- Solis-Miranda, J.; Rubio-Ramos, R.; Gonzalez-Rodriguez, S.; Gutierrez-Beltran, E. Isolation and Visualization of Plant Stress Granule-Associated Components via On-Beads Digestion and Co-localization Analysis. Methods Mol. Biol. 2022, 83, 57–66. [Google Scholar]
- He, S.L.; Li, B.; Zahurancik, W.J.; Arthur, H.C.; Sidharthan, V.; Gopalan, V.; Wang, L.; Jang, J.C. Overexpression of stress granule protein TZF1 enhances salt stress tolerance by targeting ACA11 mRNA for degradation in Arabidopsis. Front Plant Sci. 2024, 15, 1375478. [Google Scholar] [CrossRef] [PubMed]
- Satoh, R.; Tanaka, A.; Kita, A.; Morita, T.; Matsumura, Y.; Umeda, N.; Takada, M.; Hayashi, S.; Tani, T.; Shinmyozu, K.; et al. Role of the RNA-binding protein Nrd1 in stress granule formation and its implication in the stress response in fission yeast. PLoS ONE. 2012, 7, e29683. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bisaro, D.M. Cell-cell communication and initial population composition shape the structure of potato spindle tuber viroid quasispecies. Plant Cell 2024, 36, 1036–1055. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bisaro, D.M. Biased Pol II fidelity contributes to conservation of functional domains in the Potato spindle tuber viroid genome. PLoS Pathog. 2020, 16, e1009144. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Bisaro, D.M. Tobacco mosaic virus movement protein complements a Potato spindle tuber viroid RNA mutant impaired for mesophyll entry but not mutants unable to enter the phloem. PLoS Pathog. 2022, 18, e1011062. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Leontis, N.B.; Zirbel, C.L.; Bisaro, D.M.; Ding, B. A three-dimensional RNA motif mediates directional trafficking of Potato spindle tuber viroid from epidermal to palisade mesophyll cells in Nicotiana benthamiana. PLoS Pathog. 2019, 15, e1008147. [Google Scholar] [CrossRef]
- Wu, J.; Zhang, Y.; Nie, Y.; Yan, F.; Zirbel, C.L.; Bisaro, D.M. RNA three-dimensional structure drives the sequence organization of potato spindle tuber viroid quasispecies. PLoS Pathog. 2024, 20, e1012142. [Google Scholar] [CrossRef]
- Wu, J.; Zhou, C.; Li, J.; Li, C.; Tao, X.; Leontis, N.B.; Zirbel, C.L.; Bisaro, D.M.; Ding, B. Functional analysis reveals G/U pairs critical for replication and trafficking of an infectious non-coding viroid RNA. Nucleic Acids Res. 2020, 48, 3134–3155. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Zhang, Z.; Wan, Q.; Zhou, H.; Jiao, M.; Zheng, H.; Lu, Y.; Rao, S.; Wu, G.; Chen, J. Selection and validation of reference genes for RT-qPCR analysis of gene expression in Nicotiana benthamiana upon single infections by 11 positive-sense single-stranded RNA viruses from Four Genera. Plants 2023, 12, 857. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Pozo, N.; Menda, N.; Edwards, J.D.; Saha, S.; Tecle, I.Y.; Strickler, S.R.; Bombarely, A.; Fisher-York, T.; Pujar, A.; Foerster, H.; et al. The Sol Genomics Network (SGN)—From genotype to phenotype to breeding. Nucleic Acids Res. 2015, 43, D1036–D1041. [Google Scholar] [CrossRef] [PubMed]
- Amari, K.; Lerich, A.; Schmitt-Keichinger, C.; Dolja, V.V.; Ritzenthaler, C. Tubule-guided cell-to-cell movement of a plant virus requires class XI myosin motors. PLoS Pathog. 2011, 7, e1002327. [Google Scholar] [CrossRef]
- Wang, D.; Chen, M.; Peng, J.; Zheng, H.; Lu, Y.; Wu, G.; Wu, J.; Li, J.; Chen, J.; Yan, F.; et al. Transcriptome Analysis of Tomato Leaves Reveals Candidate Genes Responsive to Tomato Brown Rugose Fruit Virus Infection. Int. J. Mol. Sci. 2024, 25, 4012. [Google Scholar] [CrossRef]

| Samples | Libraries | Raw_Reads | Raw_Bases | Clean_Reads | Clean_Bases | Error_Rate | Q20 | Q30 | GC_pct |
|---|---|---|---|---|---|---|---|---|---|
| TRV:SlRNC1-1 | SRAS240002246-1a | 47923554 | 7.19G | 46986592 | 7.05G | 0.01 | 98.80 | 96.58 | 43.48 |
| TRV:SlRNC1-2 | SRAS240002247-1a | 50772296 | 7.62G | 49650698 | 7.45G | 0.01 | 98.24 | 95.70 | 41.04 |
| TRV:SlRNC1-3 | SRAS240002248-1a | 46462328 | 6.97G | 43268944 | 6.49G | 0.01 | 98.83 | 96.70 | 43.41 |
| TRV:00-1 | SRAS240002249-1a | 44465486 | 6.67G | 41516244 | 6.23G | 0.01 | 98.67 | 96.26 | 42.58 |
| TRV:00-2 | SRAS240002250-1a | 44624792 | 6.69G | 44557430 | 6.68G | 0.01 | 98.53 | 95.86 | 43.27 |
| TRV:00-3 | SRAS240002251-1a | 45080082 | 6.76G | 43849266 | 6.58G | 0.01 | 98.80 | 96.62 | 42.41 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nie, Y.; Zhang, Y.; Wang, L.; Wu, J. Unveiling the Role of SlRNC1 in Chloroplast Development and Global Gene Regulation in Tomato Plants. Int. J. Mol. Sci. 2024, 25, 6898. https://doi.org/10.3390/ijms25136898
Nie Y, Zhang Y, Wang L, Wu J. Unveiling the Role of SlRNC1 in Chloroplast Development and Global Gene Regulation in Tomato Plants. International Journal of Molecular Sciences. 2024; 25(13):6898. https://doi.org/10.3390/ijms25136898
Chicago/Turabian StyleNie, Yuxin, Yuhong Zhang, Luyou Wang, and Jian Wu. 2024. "Unveiling the Role of SlRNC1 in Chloroplast Development and Global Gene Regulation in Tomato Plants" International Journal of Molecular Sciences 25, no. 13: 6898. https://doi.org/10.3390/ijms25136898
APA StyleNie, Y., Zhang, Y., Wang, L., & Wu, J. (2024). Unveiling the Role of SlRNC1 in Chloroplast Development and Global Gene Regulation in Tomato Plants. International Journal of Molecular Sciences, 25(13), 6898. https://doi.org/10.3390/ijms25136898




