Abstract
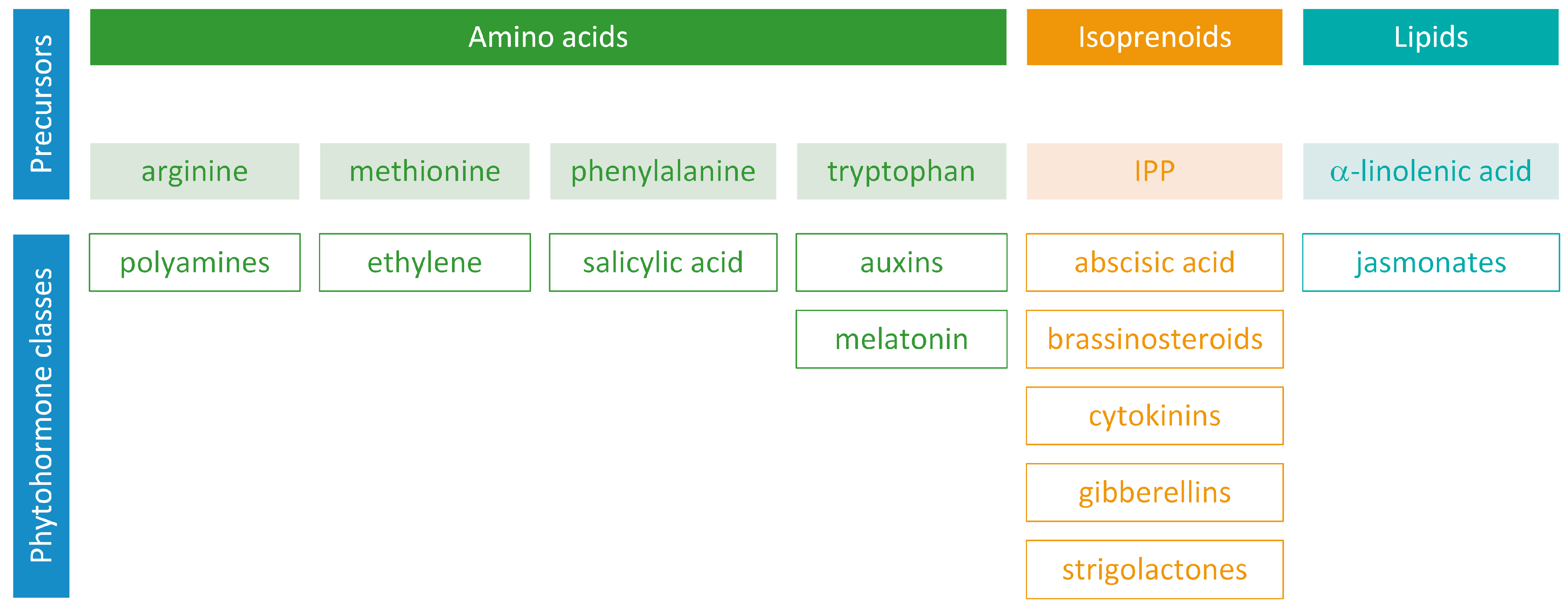
Phytohormones exhibit a wide range of chemical structures, though they primarily originate from three key metabolic precursors: amino acids, isoprenoids, and lipids. Specific amino acids, such as tryptophan, methionine, phenylalanine, and arginine, contribute to the production of various phytohormones, including auxins, melatonin, ethylene, salicylic acid, and polyamines. Isoprenoids are the foundation of five phytohormone categories: cytokinins, brassinosteroids, gibberellins, abscisic acid, and strigolactones. Furthermore, lipids, i.e., α-linolenic acid, function as a precursor for jasmonic acid. The biosynthesis routes of these different plant hormones are intricately complex. Understanding of these processes can greatly enhance our knowledge of how these hormones regulate plant growth, development, and physiology. This review focuses on detailing the biosynthetic pathways of phytohormones.
1. Introduction
Phytohormones, which are also known as plant hormones, are small, naturally occurring organic compounds that significantly influence the growth, development, defense, productivity, and physiological mechanisms of plants. They also orchestrate various cellular activities within the plant. Even at minimal concentrations, they are operative in plant cells, tissues, and organs. They are found in all vascular plants and a substantial number of non-vascular species (Table 1). From the initial discovery of auxin to the most recent unearthing of strigolactones (SLs), 12 groups of phytohormones—auxins, cytokinins (CKs), gibberellins (GAs), abscisic acid (ABA), ethylene, brassinosteroids (BRs), salicylic acid (SA), jasmonates, polyamines (PAs), melatonin, SLs, and peptide hormones—have been identified in numerous plant species. The various chemical structures of phytohormones are pivotal for their diverse biological functions and biosynthesis (Figure 1):
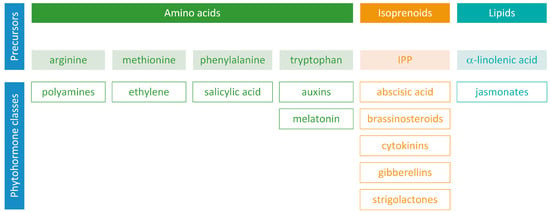
Figure 1.
Precursors of phytohormone biosynthesis.
- auxins and melatonin are indole derivatives;
- ABA is a sesquiterpene;
- ethylene is the simplest alkene;
- CKs are adenine analogues;
- GAs are tetracyclic diterpenoid acids;
- BRs are polyhydroxysteroids;
- jasmonates are derived from fatty acids;
- PAs are aliphatic nitrogenous bases;
- SA is a phenolic organic acid;
- SLs are terpenoid lactones [1,2].
In addition, peptide hormones regulate many developmental and defense processes, such as meristem maintenance, xylem and phloem differentiation, stomata patterning, pollination, embryo and endosperm development, cell division, nodulation, and systematic response [3]. They are divided into secreted and non-secreted types. Secreted peptide hormones are further divided into post-translationally modified peptides and cysteine-rich peptides. Peptide hormones are synthesized as larger precursor molecules, which are then cleaved to produce active peptides, e.g., CLAVATA3 (CLV3)/Embryo Surrounding Region-Related (CLE), Phytosulfokine (PSK), Plant Peptide-Containing Sulfated Tyrosine (PSY) peptides belong to post-translationally modified peptides; Rapid Alkalinization Factor (RALF) peptides belong to cysteine-rich peptides; Plant Elicitor Peptides (PEP)—to non-secreted peptides. Peptide hormones are also generated by plant pathogens, symbionts, and microbes that interact with plants. They are crucial in establishing a molecular interface that allows them to co-exist with the host plant. These organisms produce effectors that mimic peptide phytohormones and other effectors of pathogens and symbionts. Plant receptors recognize these effectors, which primarily regulate growth rather than defense responses. However, the origin of non-plant peptide phytohormones is still a subject of controversy [3,4,5,6]. Although peptide hormones are considered a legitimate group of plant hormones, their biosynthesis is not covered by the scope of this article because their biosynthesis shares all of the typical characteristics of peptide biosynthesis.

Table 1.
The localization of hormones in plants [7,8].
Table 1.
The localization of hormones in plants [7,8].
| Phytohormone Class | Occurrence and Site of Biosynthesis |
|---|---|
| Abscisic acid | Roots, mature leaves, particularly in response to water stress, seeds |
| Auxins | Leaf primordia, young leaves, developing seeds |
| Brassinosteroids | Leaves, shoots, roots, fruits, seeds, pollen |
| Cytokinins | Root tips, developing seeds, leaves, stem, flowers, siliques, fruits, shoot meristem |
| Ethylene | Tissues undergoing ripening (fruits), roots, shoots, particularly in response to stress |
| Gibberellins | Young tissues of the shoot, developing seeds |
| Jasmonates | Leaves, roots |
| Melatonin | Leaves, stems, roots, fruits, seeds |
| Polyamines | Most tissues, particularly in response to stress, in tissues undergoing senescence or ripening |
| Salicylic acid | Leaves, particularly in response to pathogenic attack |
| Strigolactones | Roots, shoots |
As such, plants host diverse phytohormone pathways. Significant advancements have been made in the study of phytohormone biology and synthesis over the past decade. An assortment of new tools and methods has been developed, resulting in the discovery of phytohormone substrates, intermediates, and final products. The biosynthetic pathways of plant hormones have been elucidated, and extensive research has been carried out into the genes found in the plant genome, encoding the enzymes that catalyze the various stages of phytohormone synthesis [1,2].
2. Polyamines
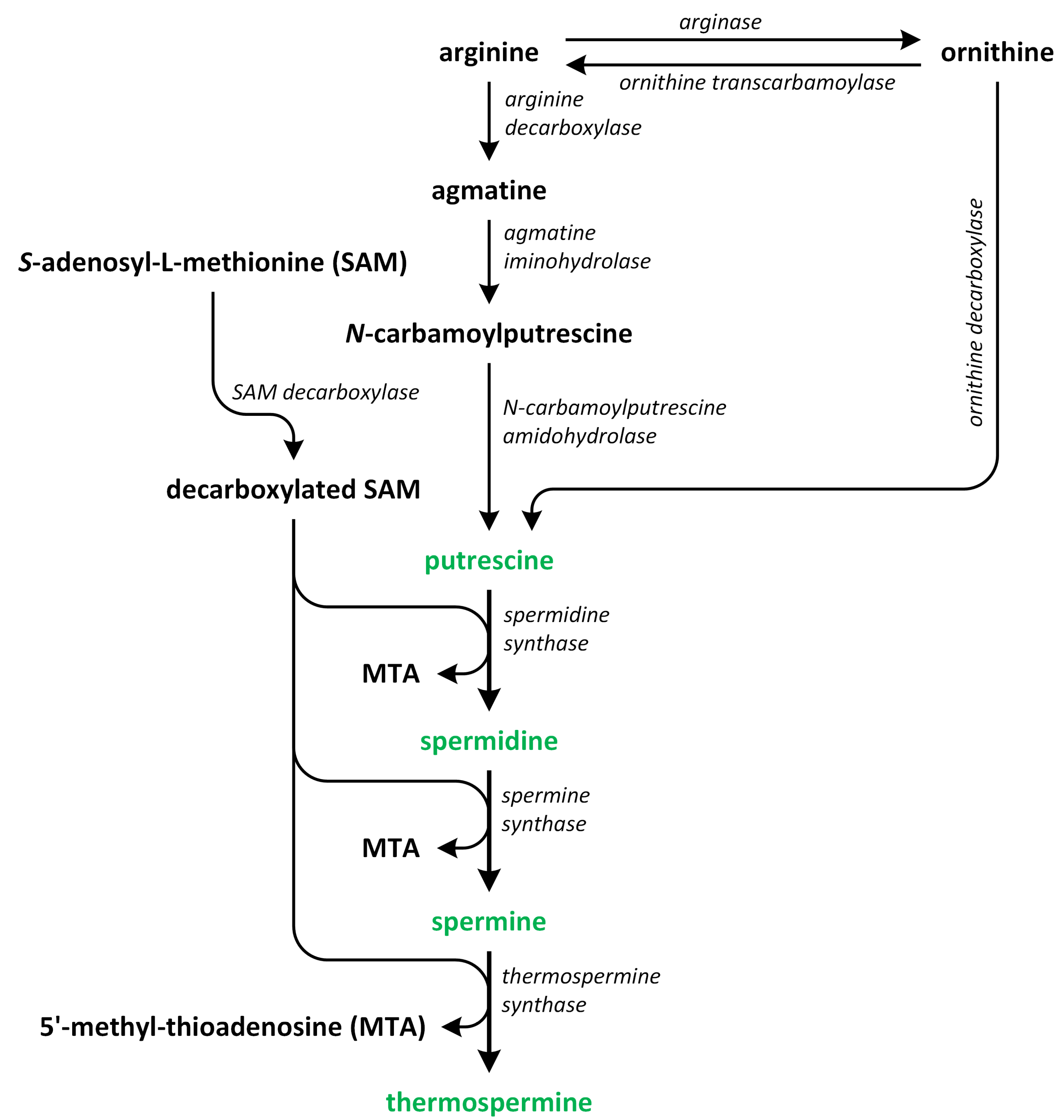
Polyamines (PAs) play a crucial role in plant growth, metabolism, and development. Structurally, these compounds are organic polycationic alkylamines that contain two or more amino groups. PAs such as putrescine (Put), spermidine (Spd), and spermine (Spm) are the most well-studied and recognized PAs found in plants. These three PAs regulate various physiological activities, such as photosynthesis, flower generation, embryogenesis, and organogenesis. They are also accountable for maintaining the stability of nucleic acids, various protein molecules, and the membrane structure. Additionally, they play a substantial role in enhancing the tolerance of many plants to the presence of a variety of abiotic and biotic stress factors, thereby impacting crop yield. Putrescine is a key compound involved in the biosynthesis of PAs. It serves as a common intermediate in the creation of Spd, Spm, and thermospermine. The biosynthesis pathways of Put have been identified in many plants (Figure 2) [9,10,11,12].
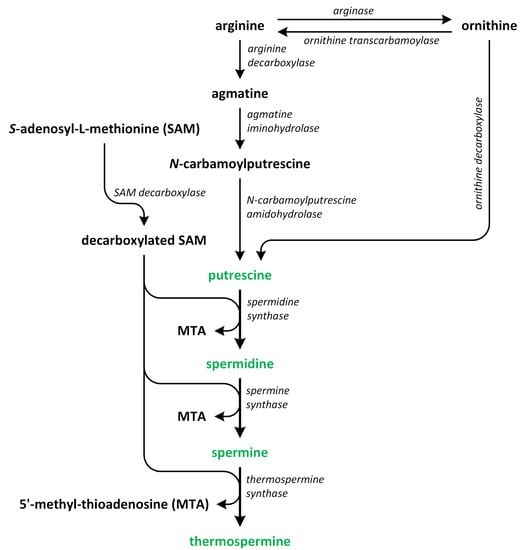
Figure 2.
The biosynthetic pathways of polyamines (phytohormones are green).
In the first pathway, arginine (Arg) is decarboxylated through the action of enzyme argi-nine decarboxylase (ADC). This enzyme acts on the carbon at position 8. In this reaction, agmatine (Agm) and CO2 (the ADC pathway) are produced. Subsequently, Agm is used to synthesize N-carbamoyl putrescine (NCPA) and ammonia (NH3), which occurs following the removal of nitrogen. In the succeeding reaction, NCPA amidohydrolase hydrolyzes NCPA, removing its carbamoyl group to produce NH3, CO2, and the first PA—Put. This process is the most prevalent pathway for Put synthesis in plants [13,14].
In the second pathway of Put biosynthesis, ornithine (Orn) is formed from Arg in the reaction catalyzed by arginase [15]. Next, the enzyme ornithine decarboxylase (ODC) removes the carboxyl group bonded to C-1 of Orn to form products such as Put and CO2 [13]. On the other hand, the ODC gene that encodes this enzyme has not been identified in Arabidopsis thaliana and other plants from the Brassicaceae family. Therefore, there is a high possibility that the ornithine pathway, which is known also as ODC pathway, is not important for normal plant growth and development in relation to the ADC pathway [16,17]. However, other authors speculate that the ODC pathway is mainly involved in regulation of plant growth, development, differentiation of organs, and reproduction, while the ADC pathway is activated in plants that grow under unfavorable environmental conditions. These two pathways are commonly found in plants [10,16,17,18,19,20].
The third pathway is the least known route of Put formation in plants. Other PAs, such as Spd and Spm, are biosynthesized from Put. Decarboxylated S-adenosyl-L-methionine (dcSAM) is a source of aminopropyl residues, which are the indispensable intermediates involved in the polyamine biosynthetic pathway [10,21]. For dcSAM production, S-adenosyl-L-methionine (SAM) is decarboxylated through the activity of the enzyme S-adenosyl-L-methionine decarboxylases (SAMDC) [22]. Aminopropyl groups are added to the Put molecule in a reaction catalyzed by spermidine synthase (SPDS), and Spd is produced. In A. thaliana, two genes, i.e., SPDS1 (At1g23820) and SPDS2 (At1g70310) that encode SPDS, as well as four SAMDC1-4 genes, i.e., At3g02470, At3g25570, At5g15959, and At5g18930, that encode SAMDC, which are essential for plant embryogenesis, have been identified [23]. The last step of the biosynthesis of PAs, i.e., the transfer of aminopropyl groups to propylamine acceptor Spd, leads to the formation of Spm. In this reaction, this Spm is formed via the action of spermine synthase (SPMS) [18,24,25,26]. The biosynthesis of PAs in plants is closely connected to the biosynthetic pathways of other plant hormones. For example, SAM, which is present in the PA biosynthetic pathway, plays also a key role as a precursor involved in ethylene biosynthesis, and past studies showed that the production of PAs can compete with ethylene [27,28].
3. Ethylene
The volatile phytohormone ethylene plays a key role in controlling various developmental and physiological activities, including seed dormancy and germination, growth of vegetation, flowering, maturation of climacteric fruit, and aging. Moreover, ethylene is seen as a critical component that protects plants from both biological and non-biological stressors. Being a gaseous plant hormone, ethylene has the ability to easily spread from its creation sites without needing any biochemical alterations or metabolic processes, allowing it to be sensed promptly [29].
Ethylene production is intricately controlled by internal cues throughout its development. Additionally, ethylene production can be swayed by environmental triggers, including biological triggers, like an onslaught of pathogens, and non-biological triggers, like injury, low oxygen levels, exposure to ozone, cold temperatures, or freezing conditions [30].
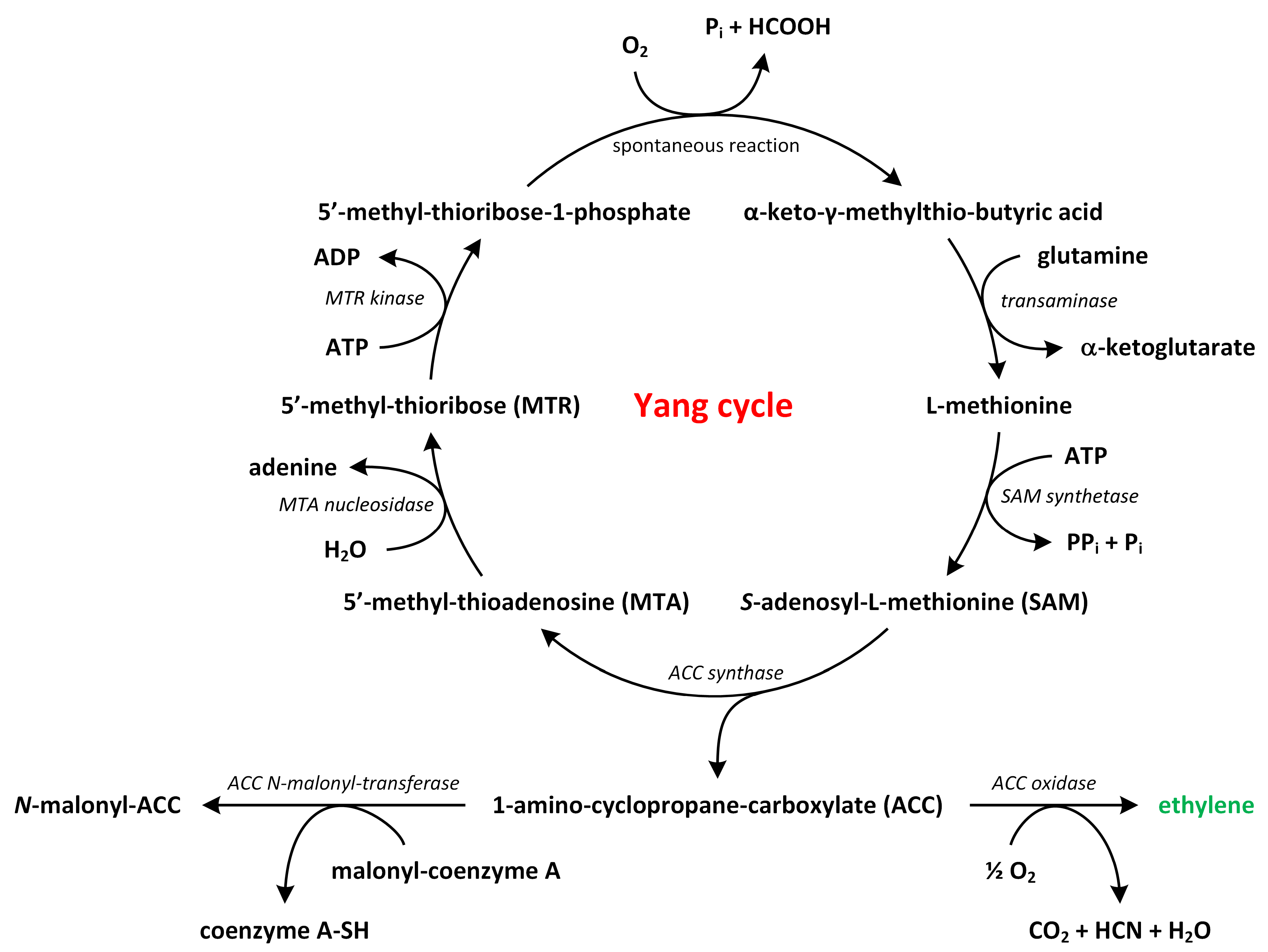
The biosynthesis of ethylene is a relatively straightforward process that involves two key enzymatic reactions (Figure 3).
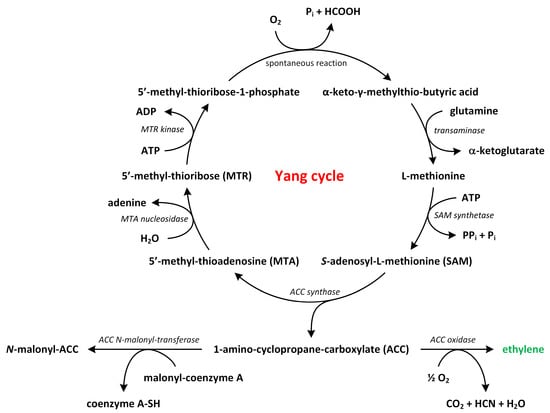
Figure 3.
The biosynthetic pathways of ethylene (phytohormone is green).
Methionine, which is the precursor of ethylene, is converted to S-adenosylmethionine (SAM) by the enzyme S-adenosylmethionine synthetase. Around 80% of cellular methionine is directed towards SAM synthesis, which is involved in various pathways, including polyamines (PAs) and ethylene biosynthesis [31,32,33]. SAM also participates in methylation reactions that modify lipids, proteins, and nucleic acids [30]. Next, SAM is transformed into 1-aminocyclopropane-1-carboxylic acid (ACC) and 5′-methyl-thioadenosine (MTA) by the enzyme ACC synthase (ACS) [34,35,36]. This conversion represents the rate-limiting step in the ethylene biosynthetic pathway, as the expression of ACS genes is tightly regulated by environmental factors. ACS is present at low levels under normal conditions, though its activity increases in response to biotic or abiotic stress, suggesting tight control over ethylene production [37,38].
In the second step, ACC is further converted into ethylene and other by-products, such as CO2 and cyanide, by the enzyme ACC oxidase (ACO) (Figure 3). In A. thaliana, ACO proteins are encoded by five genes (ACO1–5) and belong to a superfamily of oxygenases/oxidases. The toxic cyanide by-product is rapidly detoxified through its conversion into β-cyanoalanine by β-cyanoalanine synthase [39]. Alternatively, ACC can be conjugated to malonyl-ACC, and this process regulates the availability of ACC for conversion to ethylene. By controlling the ratio of malonyl-ACC to ACC, plants can modulate ethylene biosynthesis [29,40,41].
4. Salicylic Acid
Salicylic acid (SA) participates in plant growth and development. In heat-generating plants, it initiates the process of heat production by triggering an alternative respiration pathway, leading to the release of foul-smelling compounds that lure pollinators. In terms of plant defense, SA acts as a messenger, managing the expression of plant pathogenesis-related genes and fostering disease resistance. Additionally, it helps to orchestrate plant reactions to diverse types of non-biological stress factors, like extreme temperatures, saline conditions, and oxidative stress. SA also has a role in controlling a range of other activities in plants, including photosynthesis, respiration, growth of vegetation, seed germination, flowering, and aging [42,43,44].
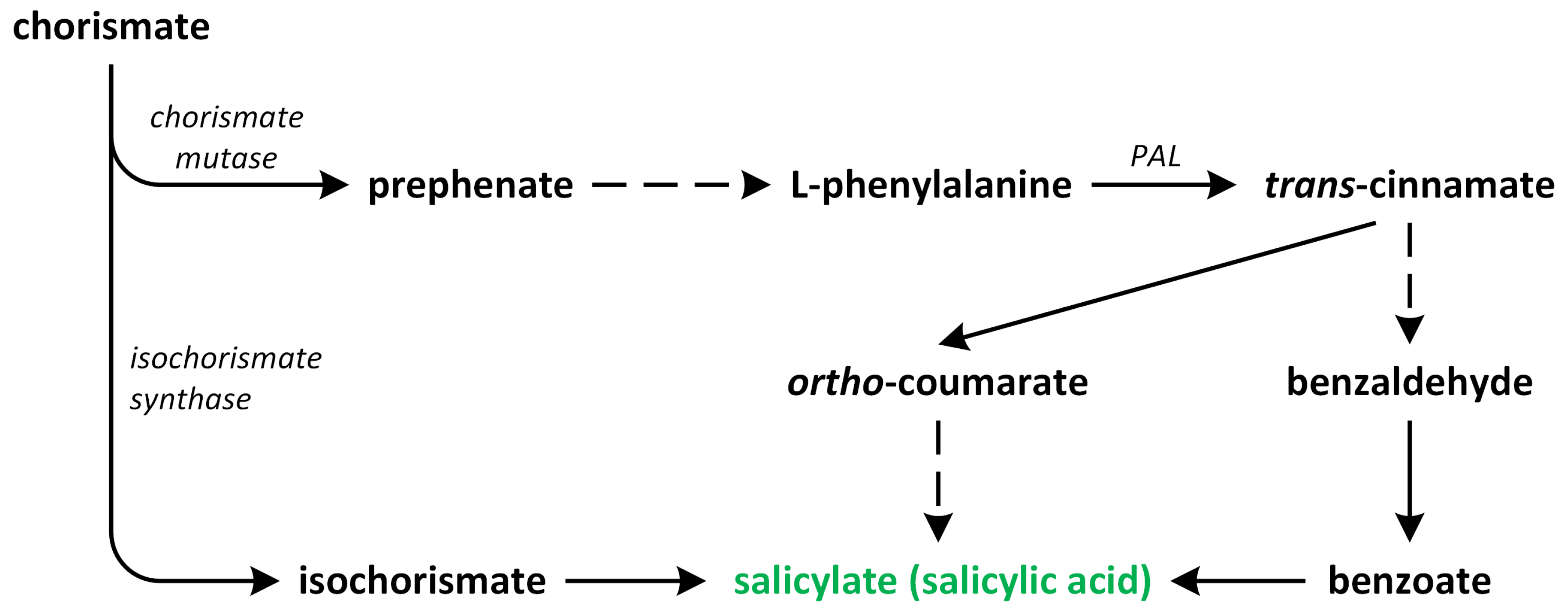
In plants, SA occurs via two pathways: (i) the isochorismate synthase (ICS) (major fraction) and (ii) the phenylalanine ammonia-lyase (PAL) pathways (Figure 4). Both biosynthetic pathways start in chorismate plastids and vary between plant species [42]. Although plants employ both pathways simultaneously, the ICS pathway is the primary contributor, accounting for approximately 95% of SA synthesis. Chorismate is a primary metabolic precursor of both pathways [45]. This compound is the end product of the shikimate pathway, which is initiated by erythrose-4-phosphate and phosphoenolpyruvate [46,47].
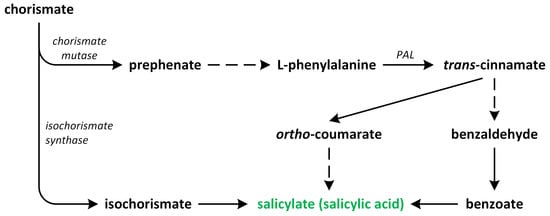
Figure 4.
The biosynthetic pathways of salicylic acid (phytohormone is green).
The ICS pathway was first discovered in bacteria. Later, its presence was confirmed in vascular plants [48,49]. In the ICS pathway, the first identified step is the conversion of chorismate into its isomer, which is known as isochorismate, via the reaction catalyzed by ICS or its homologous. These enzymes are common to both bacteria and plants. A. thaliana and soybean (Glycine max) possess two genes that encode ICS, while rice (Oryza sativa) has only one gene that encodes this enzyme. Although the activities of ICS enzymes may vary between plant species, their primary structures are almost conserved [47]. Isochorismate is, indeed, produced in plastids and subsequently transported to the cytosol. The transportation of isochorismate from plastids to the cytosol is facilitated by the Enhanced Disease Susceptibility 5 (EDS5) protein, which functions as a MATE transporter [48]. Once it is present in the cytosol, isochorismate undergoes conjugation with L-glutamate and is converted into isochorismate-9-glutamate (ICS-Glu) through the action of a cytosolic amidotransferase. In the last step, the spontaneous decomposition of ICS-Glu leads to accumulation of the following products: SA and 2-hydroxy-acryloyl-N-glutamate. The activity of the cytosolic amidotransferase, which converts isochorismate into isochorismate-9-glutamate (ICS-Glu), is, indeed, inhibited by salicylic acid (SA), which acts as a negative feedback regulation mechanism. This feedback inhibition by SA helps to regulate the biosynthesis of SA and control its levels within the plant [42]. In A. thaliana, the ICS pathway plays a significant role in the accumulation of SA in response to pathogen infection. The ICS pathway is known to be a major contributor to SA biosynthesis, particularly during pathogen-induced defense responses. Upon pathogen recognition, the activation of defense signaling pathways leads to an increase in ICS enzyme activity, resulting in the production of isochorismate and its subsequent conversion into SA. This pathogen-induced SA accumulation plays a crucial role in triggering and amplifying plant defense responses against pathogens [47].
The second route of SA biosynthesis, which is known as the PAL pathway, is responsible for the synthesis of only a small percentage of SA in plants such as A. thaliana and Nicotiana benthamiana, which were infected by pathogens or treated with UV during the reviewed studies [50]. This pathway occurs entirely within the cytosol. However, the PAL pathway has been known for much longer in relation to its role in the ICS pathway [48,51]. PAL is an upstream enzyme that leads to the production and accumulation of many other potentially defense-related compounds that are synthesized during the phenylpropanoid pathway [52]. The conversion of phenylalanine into trans-cinnamic acid is catalyzed by PAL, which has been identified in numerous plant species. The activity of this enzyme is tightly regulated by various biotic and abiotic stress factors [53]. Depending on the order of the hydroxylation of the aromatic ring in trans-cinnamate, SA can also be synthesized from o-coumaric acid or benzoic acid through chain-shortening reactions. The specific pathway that leads to SA formation is determined based on whether hydroxylation occurs before or after the chain-shortening reactions [50]. In certain plants, such as sunflower (Helianthus annuus), potato (Solanum tuberosum), and pea (Pisum sativum), SA is derived from benzoic acid. This result is achieved via the chain-shortening of cinnamate through a β-oxidation process, leading to the synthesis of benzoic acid, which is subsequently converted into SA [54]. On the other hand, Primula acaulis and Gaultheria procumbens can synthesize SA from o-coumarate [55].
5. Auxins
Auxins play a crucial role in the regulation of various aspects of plant growth and development. They are involved in controlling processes such as cell division, elongation, differentiation, tropisms (response to external stimuli), flowering, apical dominance (suppression of lateral bud growth), lateral root formation, senescence (aging), abscission (shedding of plant parts), and responses to environmental stresses [56]. The most well-studied auxin in plants is indole-3-acetic acid (IAA). The process of producing auxins in plants is highly intricate, and understanding this process can significantly enhance our comprehension of auxins’ biological role and contribution to the regulation of plant growth and physiology [57]. Although different plant species employ distinct strategies to optimize their metabolic pathways, it seems that there could be shared mechanisms for auxin biosynthesis, given that IAA is an phytohormone essential to plant life. Auxins are primarily generated in the plant’s apical meristems, young leaves, and flower buds, and they are then quickly transported throughout the plant via the phloem. Nonetheless, some research suggests that auxins may also be locally synthesized within the roots [58,59,60].
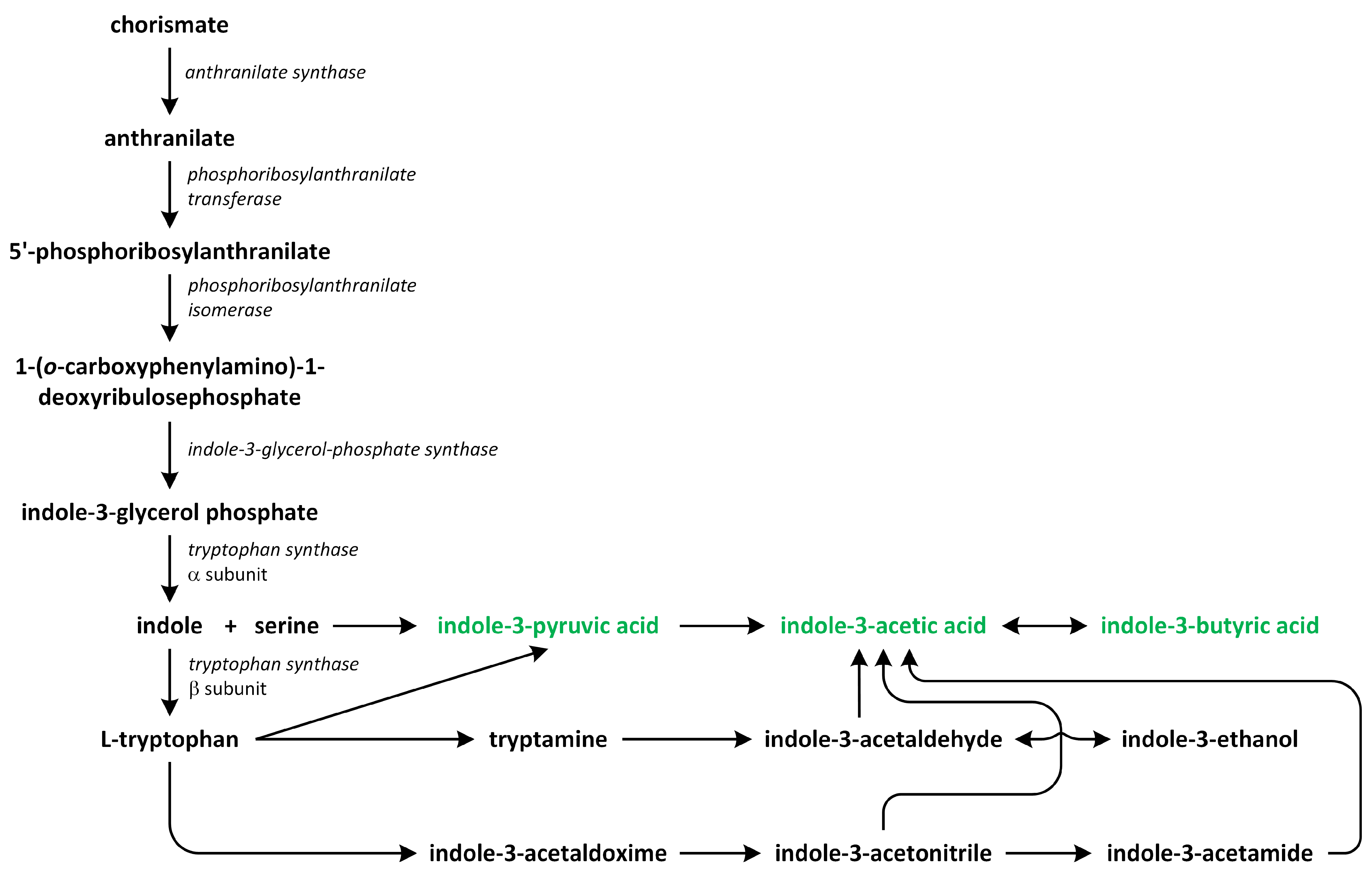
Two major routes have been assumed to contribute to de novo IAA biosynthesis in plants: the tryptophan (Trp)-dependent and Trp-independent pathways (Figure 5). Several pathways for Trp-dependent IAA biosynthesis have been proposed, including:
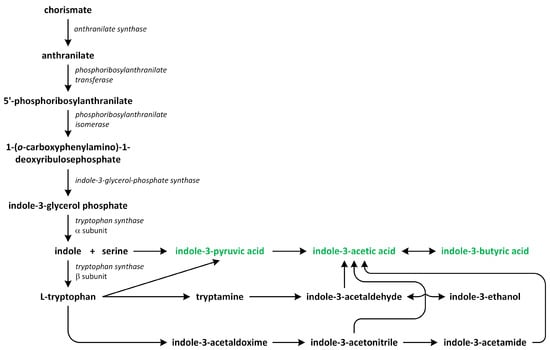
Figure 5.
The biosynthetic pathways of auxins (phytohormones are green).
- the indole-3-acetamide (IAM) pathway;
- the indole-3-pyruvic acid (IPA) pathway;
- the tryptamine (TAM) pathway;
- the indole-3-acetaldoxime (IAOX) pathway [61,62,63,64,65].
Even though IAA was the first natural auxin to be identified, our understanding of the genes that encode all of the enzymes involved in auxin biosynthesis remains limited. It is also uncertain whether all of these pathways exist in all plant species [66].
Tryptophan, which is a precursor of the production of various indole-containing compounds in plants, including IAA, indole glucosinolates, phytoalexins, and tryptamine (TAM) derivatives, is synthesized from chorismate through the action of indole-3-glycerol phosphate synthase in the chloroplast. The A. thaliana genes Anthranilate synthase1 (AtASA1) and AtASA2 encode the α-subunit of the enzyme that is responsible for catalyzing the initial reaction through the Trp synthesis pathway. Indole-3-glycerol phosphate synthase, on the other hand, facilitates the conversion of 1-(O-carboxyphenylamino)-1-deoxyribulose-5-phosphate into indole-3-glycerol phosphate [67,68].
In hairy plant roots, IAA is synthesized from Trp via a reaction that consists of two steps. Tryptophan is first converted into IAM by the enzyme Trp-2-mono-oxygenase, and IAM is then transformed to IAA by indole-3-acetamide hydrolase [66]. This IAA biosynthetic pathway via IAM is a bacteria-specific synthesis because plant pathogens, such as Pseudomonas and Agrobacterium, have a large root-inducing plasmid and are able to induce hairy-root disease, which is characterized by a high root proliferation ratio from the place of infection, which occurs as a result of a high rate of IAA biosynthesis and accumulation [69]. Pseudomonas and Agrobacterium use Trp-2-mono-oxygeanse to convert Trp into IAM, which is then hydrolyzed into IAA. The Trp-dependent IAA biosynthesis pathway is the best-known and studied pathway used in this process [65]. It was reported that plants can also synthesize IAA via the IAM pathway. Therefore, enzymes amidases that are able to hydrolyze IAM into IAA have been identified in A. thaliana. This observation suggests that IAM may be a key substrate for IAA biosynthesis in plants [70]. In addition, overexpression of the IAM gene that encodes the enzyme tryptophan-2-mono-oxygeanse, which is involved in the conversion of Trp into IAM, increases the content of IAA and results in high-auxin phenotypes being present in plants, leading to increased hypocotyl elongation under light, severe rosette leaf epinasty, and apical dominance. Thus, plants are able to transform IAM into IAA, as has been shown in in vivo studies performed using A. thaliana [71]. IAM, which is an intermediate of auxin biosynthesis, can also be detected in many other plant species, regardless of whether they are monocots or dicots [66].
The indole-3-pyruvic acid (IPA) pathway occurs using intermediate IPA, which is crucial for auxin biosynthesis and plant development [60]. Tryptophan Aminotransferase of Arabidopsis 1 (TAA1) is a member of aminotransferases that can convert tryptophan into IPA. These enzymes have been identified in many plants. This observation indicates that this pathway is highly conserved. Mutations in the TAA1 gene that encodes this protein lead to a high reduction in IAA levels. Thus, IPA-dependent IAA biosynthesis is an important route for the biosynthesis of free IAA. The TAA1 protein belongs to the superfamily of enzymes that is dependent on the α class of PLP and exhibits Trp aminotransferase activity, which uses L-Trp, but not D-Trp, as a substrate. This enzyme can also react with other amino acids, such as L-phenylalanine, tyrosine, leucine, alanine, methionine, and glutamine [72].
Tryptophan can also be converted into TAM through the action of cytosolic Trp decarboxylase (TDC). TAM is a protoalkaloid that is involved in an early step of the biosynthetic pathway that influence terpenoid indole alkaloids [66]. TDC has been functionally and structurally characterized. It participates in synthesis of indole alkaloids and serotonin in many plants. However, TDC members probably do not fully contribute to IAA biosynthesis. As observations show, overexpression of the TDC gene in Catharanthus roseus has been indicated to generate increased levels of TAM, but not IAA, accumulation in tobacco (Nicotiana tabacum) [73]. Moreover, genetically modified rice plants with overexpression of the TDC gene displayed a typical phenotype and contained serotonin levels that were 25 times higher in leaves and 11 times higher in seeds than those found in their wild-type equivalents [74]. In rice, TDC can be involved in the production of metabolites that are derivatives of the TAM-inducing disease known as Sekiguchi lesions or the accumulation of serotonin, which is a well-known neurotransmitter in mammals and a signalling molecule that is present in many plants [75].
Another possible pathway involved in the synthesis of IAA from Trp consists of the conversion of indole-3-acetaldoxime into indole-3-acetonitrile (IAN). In this reaction, amino nitrogen is removed from tryptophan via hydrolysis of the nitrile to form acid [76]. Indole-3-acetaldoxime (IAOX) is a precursor of the biosynthesis pathway of IAA and its conjugates, and it has been proposed as the branch point between the biosynthesis of IAA and indole-3-methyl glucosinolate [77].
Studies of A. thaliana mutant lines show that IAOX is a key intermediate involved in IAA biosynthesis. This compound can be converted into IAN by eliminating one molecule of water. IAN is subsequently transformed into IAA by a family of nitrilases. IAOX can be hydrolyzed to form indole-3-acetaldehyde, and the aldehyde can then be converted into IAA via an action of two enzymes: aldehyde oxidase (AO1) or aldehyde dehydrogenase. AO1 is important enzyme for auxin biosynthesis [78]. The enzyme AO1 shows substrate specificity to indole-3-acetaldehyde, and the regulation of the activity of this enzyme is unaffected by the high levels of IAA. However, plant mutants that are unable to perform molybdopterin biosynthesis, which is an essential cofactor of all aldehyde oxidases, did not show phenotypes that were characteristic of auxin deficiency. This result suggests that aldehyde oxidases are probably not essential for auxin biosynthesis, as was previously believed [79].
6. Melatonin
Melatonin, which is formally known as N-acetyl-5-methoxytriptamine, is an indolamine plant hormone that plays a key role in controlling growth and various physiological responses. Affected functions include root structure and development, flowering, leaf aging, fruit maturation, and determining the levels of chlorophyll, proline, and carbohydrates. It also functions as a signaling molecule during non-biological stress situations, like cold, drought, heavy metal exposure, UV radiation, and salinity, and it helps to direct plant defense responses against pathogen invasions [80,81]. The highest levels of melatonin in plants are found in the mitochondria, endoplasmic reticulum, and chloroplasts, suggesting that these organelles are the primary sites involved in its biosynthesis. The varied locations of melatonin within the plant could potentially influence its mode of action in different ways [82].
The biosynthesis of melatonin begins with an essential aromatic amino acid tryptophan, which is synthesized de novo via the shikimate pathway (Figure 6).
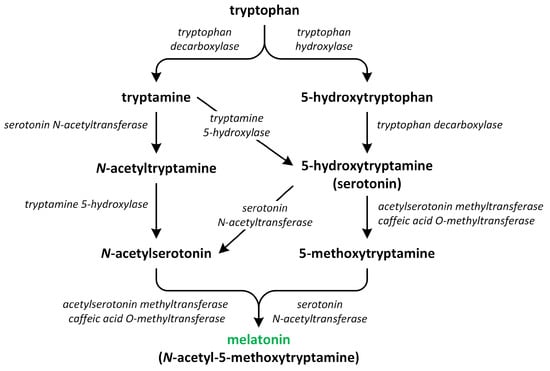
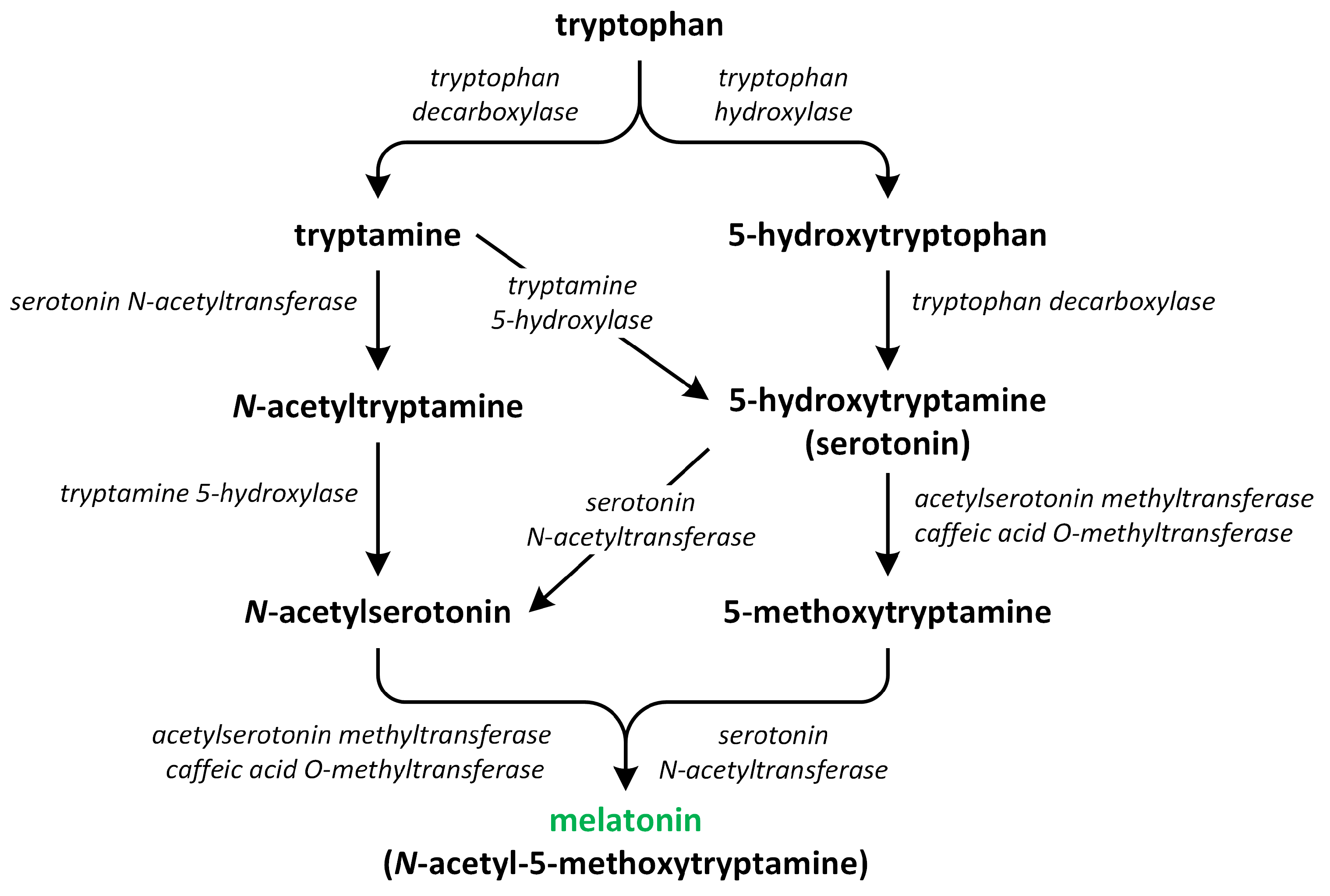
Figure 6.
The biosynthetic pathways of melatonin (phytohormone is green).
The pathway involved in the synthesis of this phytohormone occurs through various four-step reactions catalyzed by six enzymes: tryptophan decarboxylase (TDC), tryptophan hydroxylase (TPH), tryptamine 5-hydroxylase (T5H), serotonin N-acetyltransferase (SNAT), N-acetylserotonin O-methyltransferase (ASMT), and caffeic acid O-methyltransferase (COMT) [83]. These enzymes have different cellular localization. Chloroplasts and mitochondria contain SNAT. TPH, ASMT/COMT, and TDC are present in the cytoplasm. T5H can be only found in the endoplasmic reticulum [84]. Recent research suggests that the synthesis of melatonin in plants primarily takes place in chloroplasts and mitochondria. These organelles have inherited the ability to produce melatonin from their bacterial forebears. Under normal circumstances, melatonin production predominantly occurs in the chloroplasts. However, if the pathway in the chloroplasts is obstructed, the production of melatonin shifts to the mitochondria. Therefore, under stress conditions, melatonin is chiefly synthesized in the mitochondria [85,86].
The first step of melatonin biosynthesis is the formation of serotonin from Trp. Two different pathways involved in Trp and serotonin synthesis were identified in vascular plants. The first pathway involves the decarboxylation of tryptophan into tryptamine via the reaction catalyzed by the enzyme TDC. Next, tryptamine is hydroxylated into serotonin via the action of an enzyme called T5H. Another possibility is that the hydroxylation of Trp into 5-hydroxytryptophan via the action of enzyme TPH occurs. Next, the decarboxylation of 5-hydrotryptophan into serotonin by TDC takes place. The enzyme TDC is localized in the cytoplasm and characterized by a good affinity for two substrates: tryptophan and 5-hydroxytriptophan; thus, both pathways can be present in plants at levels of equal importance. The activity of TDC is considered to be the rate-limiting step in melatonin biosynthesis [80,87,88,89].
Thereafter, serotonin is transformed into melatonin via a two-step reaction. The roles of three different enzymes—SNAT, ASMT, and COMT—in this reaction were confirmed. These enzymes may possess various biochemical isoforms. SNAT is responsible for N-acetylation of serotonin, which is further methylated by the methyltransferases ASMT or/and COMT, leading to production of melatonin [90]. In addition, SNAT can convert serotonin directly into N-acetylserotonin through alternative minor pathways. The enzymes ASMT and COMT catalyze the conversion of serotonin into 5-methoxytryptamine. This compound is then changed into melatonin by the enzyme SNAT. These three enzymes are characterized by an affinity for substrates such as serotonin, N-acetylserotonin, and 5-methoxytryptamine; therefore, the order in which the different enzymes act differs [81]. SNAT is localized in the chloroplasts, whereas ASMT and COMT are present in the cytoplasm [91]. The biosynthetic ratio connected to the transformation of tryptophan into serotonin is considerably higher in relation to the conversion of serotonin into melatonin, leading to a low level of this phytohormone being present in plants [83].
When plants grow under normal environmental conditions and/or serotonin is pre-sent in low levels in cells, two following pathways exist in plants: (i) tryptophan → trypta-mine → serotonin → N-acetylserotonin → melatonin; and (ii) tryptophan → 5-hydroxytryptophan → serotonin → N-acetylserotonin → melatonin. However, the following reactions, which lead to the production of melatonin from tryptophan, were detected in plants that produce large amounts of serotonin during the senescence process. Furthermore, experiments performed on rice seedlings showed that exogenously applied N-acetylserotonin can be converted into serotonin by the enzyme N-acetylserotonin deacetylase, resulting in a reduction in the content of melatonin in plants. These different pathways and sites of melatonin synthesis suggest that the production of melatonin is highly diverse in plants [80,85,86,88].
7. Abscisic Acid
Abscisic acid (ABA) also plays an important role in many cellular processes, including seed development, dormancy, germination, vegetative growth, and responses to environmental stresses [92]. ABA is ubiquitous in lower and higher plants. It was reported that young tissues contain high levels of ABA [93]. The biosynthesis of ABA predominantly takes place in vascular tissues, and it is subsequently transported into target tissues. This transportation process occurs through both the xylem and phloem, enabling bidirectional transport between the roots and shoots. The naturally occurring form of ABA is S-(+)-ABA, which has a side chain defined as 2-cis, 4-trans. The trans-ABA is biologically inactive, whereas R-(−)-ABA, which can potentially result from racemization through the catabolite ABA-trans-diol, exhibits biological activity [94,95].
The initial stages of ABA biosynthesis occur within plastids. In plants, two distinct ABA biosynthesis pathways have been identified. The first pathway is a direct route, while the second pathway is indirect and involves the conversion of a C15 compound (i.e., farnesyl pyrophosphate) and a C40 carotenoid [96]. Through the characterization of ABA-deficient mutants and the isolation of relevant mutated genes, it has been determined that the indirect pathway is the primary route involved in ABA biosynthesis in plants [96,97]. Investigations into the biosynthesis of carotenoids, as well as their pathways and mechanisms, have indicated that mevalonate (MVA) plays a minor role as a precursor of carotenoid biosynthesis in chloroplasts [98,99].
In plants, there are two distinct pathways involved in isoprenoid production (Figure 7). The first pathway, which is known as the MVA pathway, is shared by animals, fungi, and a few bacteria. It operates in both the cytosol and mitochondria, and it is responsible for generating precursor molecules used in the synthesis of sterols, certain sesquiterpenes, and the side chain of ubiquinone. On the other hand, the second pathway, which is called the methylerythritol phosphate (MEP) pathway, is localized within plastids [100].
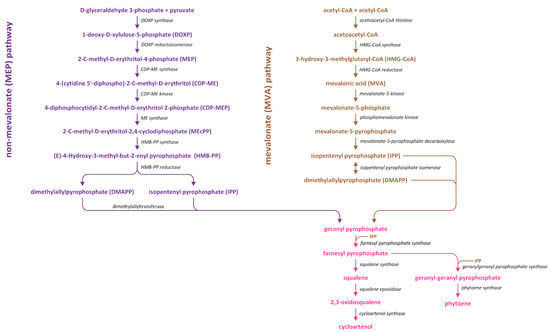
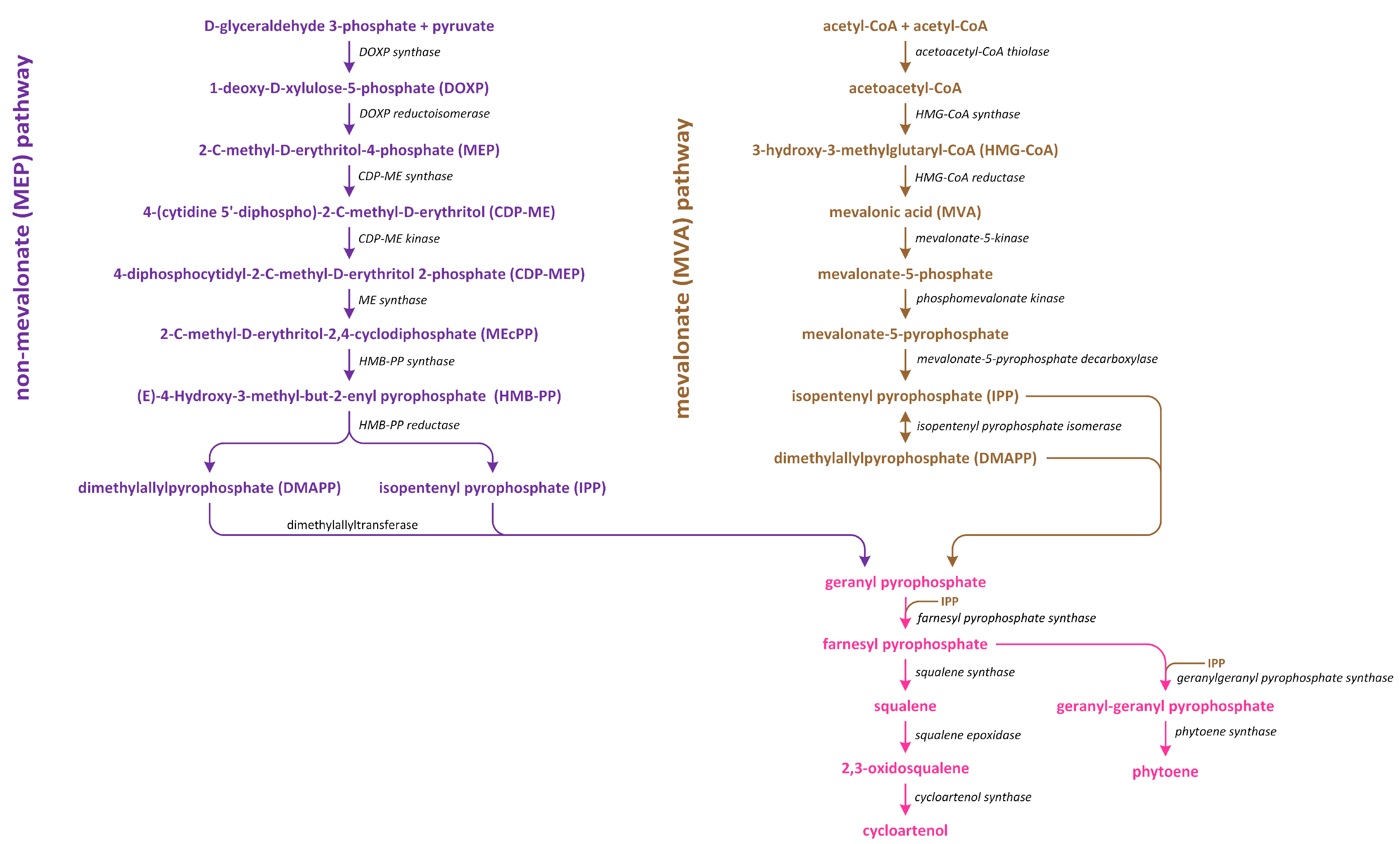
Figure 7.
The methylerythritol phosphate (MEP) and mevalonate (MVA) pathways lead to the conversion of isopentenyl diphosphate to cycloartenol and phytoene.
The MVA pathway initiates in tandem with the condensation of three acetyl~CoA molecules (Figure 7). Subsequently, a reduction process leads to the formation of MVA, which undergoes two phosphorylation reactions to produce mevalonate-5-diphosphate. Further phosphorylation, which is followed by decarboxylation-driven dephosphorylation, yields isopentenyl pyrophosphate (IPP). This compound can be isomerized to create dimethylallyl pyrophosphate (DMAPP), which serves as a precursor of the biosynthesis of various isoprenoid-derived phytohormones [101]. On the other hand, the MEP pathway initiates in tandem with the condensation of pyruvate and glyceraldehyde 3-phosphate (Figure 7). This condensation reaction gives rise to 1-deoxy-D-xylulose-5-phosphate (DOXP). Subsequently, DOXP is converted into methylerythritol phosphate. Methylerythritol phosphate then undergoes conjugation with the cytidyl phosphate moiety, followed by another phosphorylation step, i.e., the release of CMP, and cyclization. The resulting product is subjected to a reduction process, leading to the formation of (E)-4-hydroxy-3-methyl-but-2-enyl diphosphate (HMBDP). HMBDP can be further reduced to produce IPP and, to a lesser extent, DMAPP. Both HMBDP and DMAPP can serve as precursors of cytokinin synthesis, though it is worth noting that the level of HMBDP in plastids is approximately five times higher than that of DMAPP [102]. MVA can be converted into an intermediate IPP (in cytosol), which is an intermediate in both carotenoid and steroid pathways. Furthermore, in plastids in which carotenoid synthesis occurs, IPP is generated through the MEP pathway using pyruvate and glyceraldehyde-3-phosphate. The enzyme responsible for catalyzing the initial step in the non-MEP biosynthetic pathway is called DOXP synthase (DXS). After IPP is formed, it is converted into a C20 compound known as geranylgeranyl pyrophosphate (GGPP) [103].
The conversion of geranylgeranyl pyrophosphate (GGPP) into phytoene, which is a C40 carotenoid, is the initial and rate-limiting step involved in carotenoid synthesis. This reaction is catalyzed by the enzyme phytoene synthase (Figure 8).
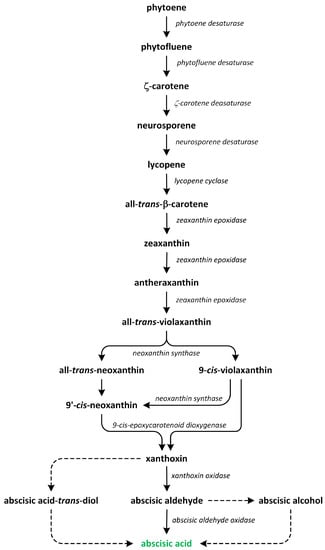
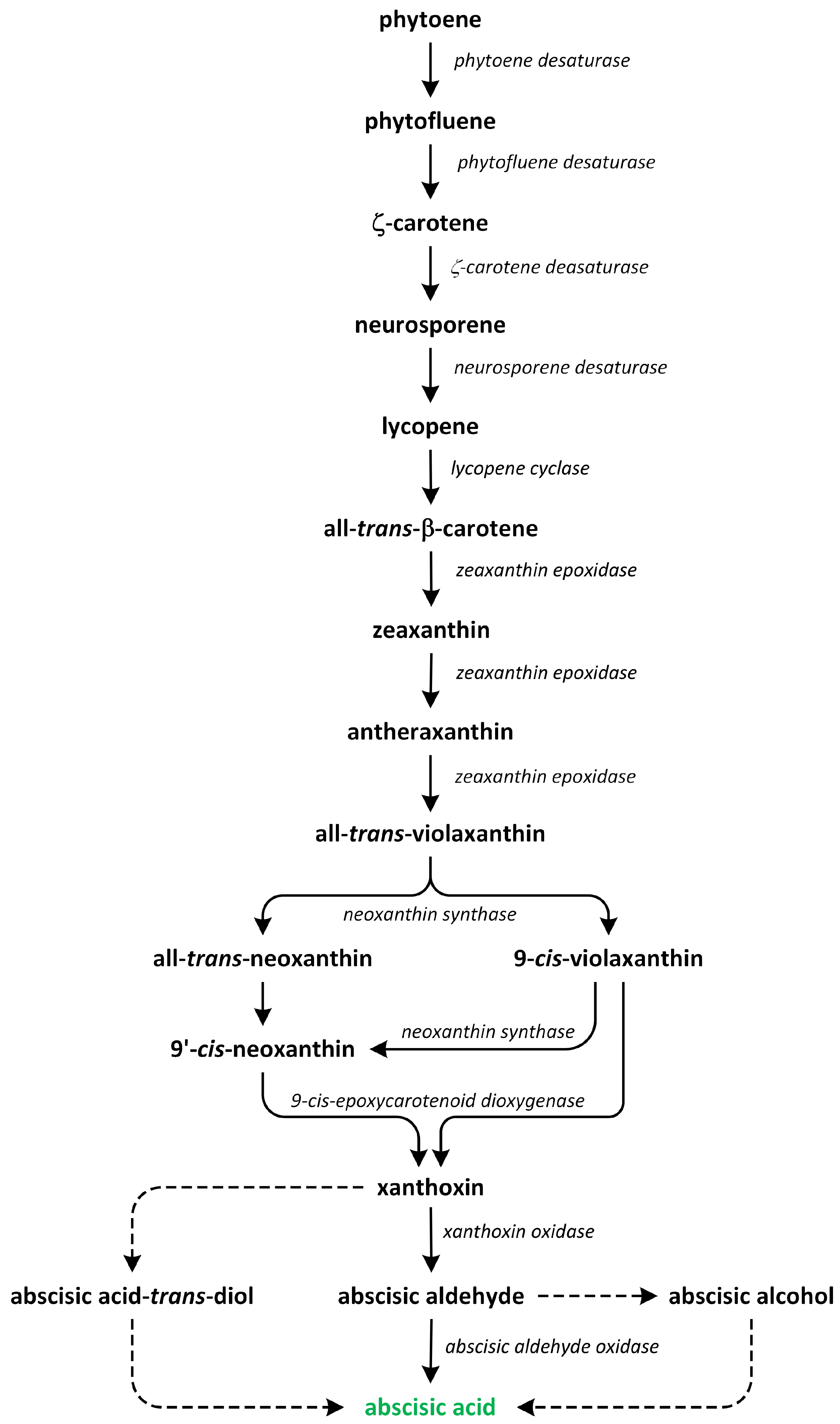
Figure 8.
The biosynthetic pathways of abscisic acid (phytohormone is green).
Subsequently, phytoene undergoes a series of enzymatic conversions in a sequential order, resulting in the formation of ζ-carotene, lycopene, β-carotene, and zeaxanthin [104]. The subsequent step, which is specific to the ABA biosynthesis pathway and occurs within plastids, involves the epoxidation of zeaxanthin and antheraxanthin to produce violaxanthin. This enzymatic conversion is mediated by zeaxanthin epoxidase. Following this step, the oxidative cleavage of xanthophylls, such as 9-cis-violaxanthin and/or 9’-cis-neoxanthin, occurs, leading to the formation of xanthoxin. This step is catalyzed by 9-cis-epoxycarotenoid dioxygenase and subsequently exported to the cytosol. Once 9-cis-epoxycarotenoids are cleaved, xanthoxin is further converted into ABA [96].
Three possible pathways have been proposed for the final stages of ABA biosynthesis that involves abscisic aldehyde, xanthoxic acid, or abscisic alcohol acting as intermediates. In the first proposed pathway, the enzyme encoded by the AtABA2 gene, which is a short-chain alcohol dehydrogenase/reductase, catalyzes the initial step and produces ABA aldehyde. The final step in this biosynthetic pathway is catalyzed by ABA aldehyde oxidase. The activities of aldehyde oxidase and xanthine dehydrogenase require molybdenum cofactor sulfurase, which is responsible for the sulfurylation of a dioxo form into a sulfurylated mono-oxo form, with L-cysteine used as a sulfur donor [94,105].
The second pathway of ABA biosynthesis that involves xanthoxic acid was discovered in ripening avocado (Persea americana) fruits. In these experiments, inhibition of aldehyde oxidase activity by tungstate, which is a potent inhibitor of molybdoenzymes in plants, led to the accumulation of xanthoxin. This result indicates that xanthoxin is a substrate for aldehyde oxidase. It is possible to speculate that the two ABA biosynthetic pathways may operate in an organ- and/or developmental stage-dependent manner, suggesting a potential regulatory mechanism [96,98,106].
The last ABA biosynthetic pathway, which involves abscisic alcohol, can be activated in certain mutants. It was shown that in the flacca and sitiens mutants of tomato (Solanum lycopersicum), when abscisic aldehyde was supplied externally, it was converted into abscisic alcohol. Subsequently, abscisic alcohol was oxidized to ABA (Figure 8) [107]. This reaction is a non-specific source of ABA in wild-type plants, though it may play a more significant role in mutants that have impaired capacity to directly convert abscisic aldehyde into ABA. This finding suggests that the shunt pathway serves as an alternative route for ABA production in tomato mutants [96].
8. Brassinosteroids
Brassinosteroids (BRs) are steroid phytohormones that elicit a wide spectrum of morphological and physiological responses, as well as a tolerance of abiotic and biotic stress [108,109,110,111,112,113,114]. BRs are categorized into three primary groups based on the number of carbon atoms present in each steroid molecule, and BRs are divided into three main groups on the basis of each steroid molecule, i.e., C27, C28, and C29 [115,116]. The basic structure of C27-BRs is the 5α-cholestane skeleton; 5α-ergostane serves as the foundational structure of C28-BRs, while 5α-stigmastane forms the basis of C29-BRs. The structures of these hormones vary due to the type and orientation of oxygenated functions in the A- and B-rings, as well as the number and position of functional groups in the side chain. These variations arise from oxidation and reduction reactions during their synthesis in plants [117,118,119].
Three pathways of BR biosynthesis that lead to the production of BRs of type C27, C28, or C29 were identified in plants. Early steps involved in their synthesis, which are common for all BRs, occur via the MVA or non-MVA (MEP) pathways (Figure 7). Later steps differentiate between the biosynthesis pathways (cycloartenol- and cycloartanol-dependent) of this group of plant hormones (Figure 9) [116,120,121].
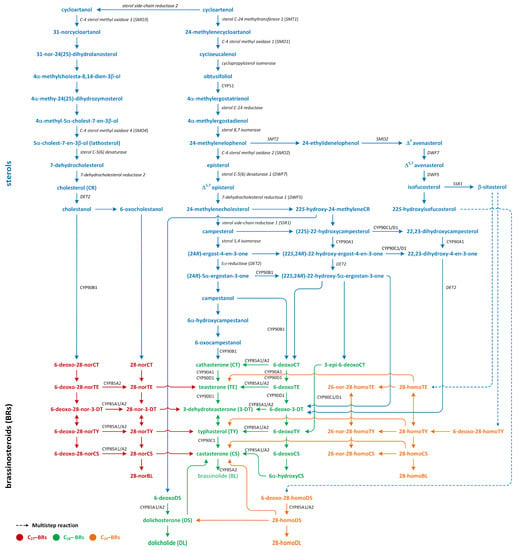
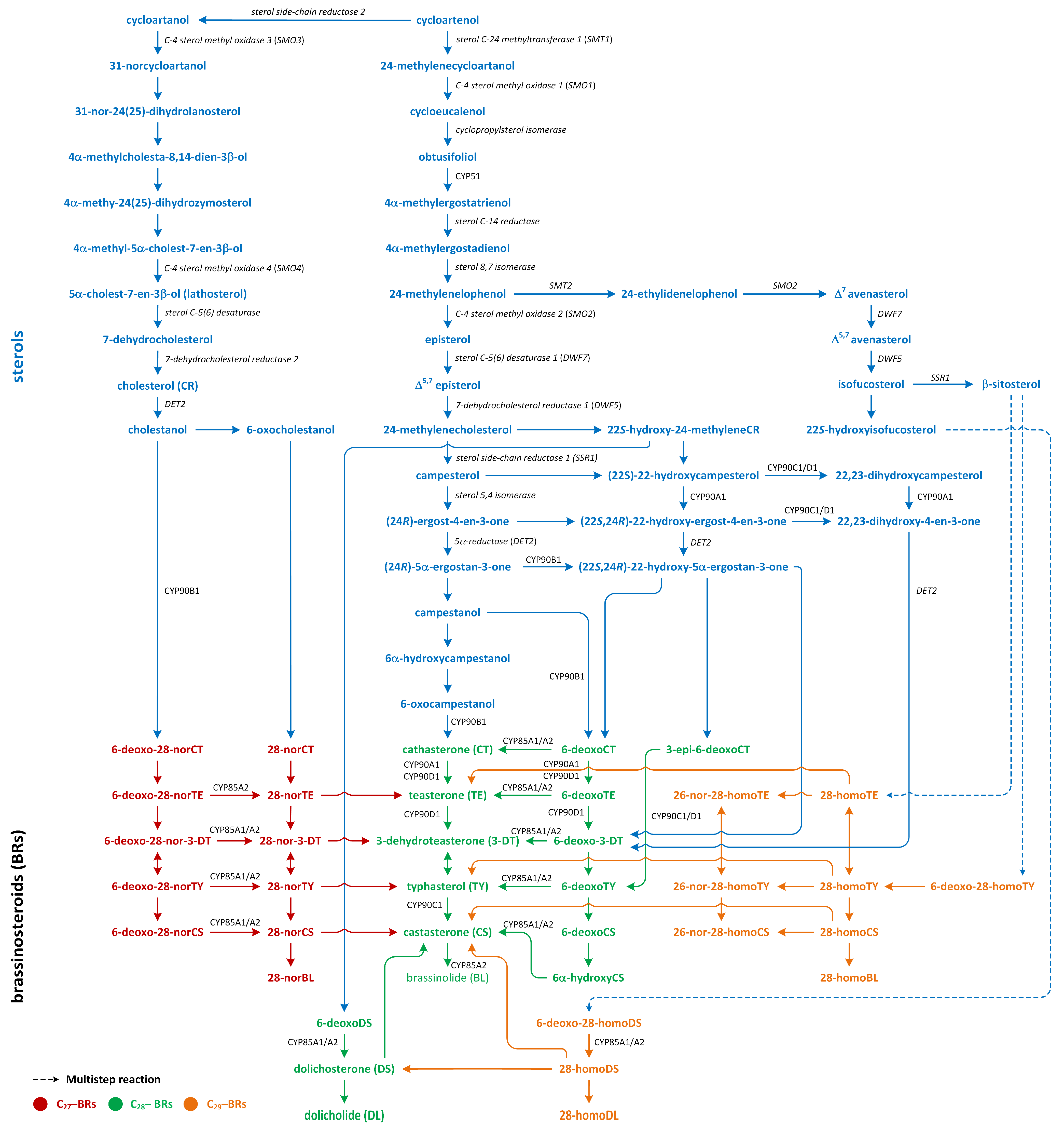
Figure 9.
The biosynthetic pathways of different types of brassinosteroids (phytohormones are red, green, and orange).
Isopentenyl pyrophosphate (IPP), which is an intermediary compound involved in the steroid synthesis pathway (for example, cholesterol), can be created via two routes: the MEP pathway (in lower plants) and the MVA pathway (in most higher plants) (Figure 7) [122]. The conversion of IPP into geranyl pyrophosphate and, subsequently, farnesyl pyrophosphate leads to the creation of squalene (Figure 7). The squalene is then oxidized to create squalene-2,3-oxide under the influence of the enzyme squalene epoxidase. Following this step, cycloartenol synthase transforms squalene-2,3-oxide into cycloartenol, which is a crucial substrate used in the synthesis of C27, C28, and C29-BRs. Cycloartenol can then be transformed into cycloartanol and, through a series of reactions, into cholesterol/cholestanol and/or 6-oxocholestanol, leading to the creation of C27-BR (Figure 9). In another process, cycloartenol may serve as a substrate for the C-24 methylation reaction, which is catalyzed by sterol C-24 methyltransferase (SMT1), resulting in the production of 24-methylenecycloartanol. Subsequent reactions, which are driven by enzymes such as C-4 sterol methyl oxidase (SMO1), cyclopropylsterol isomerase, obtusifoliol 14α-demethylase (CYP51), and sterol C-14 reductase, lead to the formation of 4α-methylergostatrienol. The intermediates of these reactions include cycloeucalenol and obtusifoliol. Next, the enzyme sterol C-14 reductase catalyzes the reduction in 4α-methylergostatrienol to create 4α-methylergostadienol, which is subsequently transformed into 24-methylenelophenol by the enzyme sterol Δ7-isomerase. 24-methylenelophenol can serve as a substrate involved in two distinct sterol biosynthesis pathways. The first pathway leads to the synthesis of isofucosterol/β-sitosterol, which are precursors of C29-BRs. 24-methylenelophenol is converted by sterol methyltransferase 2 (SMT2) into 24-ethylidenelophenol. This compound is then converted into avenasterol by sterol methyl oxidase 2. Next, avenasterol is transformed into isofucosterol and β-sitosterol, which is hydroxylated into 6-deoxo-28-homoTY and then oxygenated into 28-homoTY by CYP724B2 and CYP90B3 C-22 hydroxylase, respectively [123]. 28-homoTY can also be generated from 28-homoTE, though the intermediates involved in this reaction have not yet been identified [116]. 28-homoTY is transformed into 28-homoCS and 28-homoBL via the actions of enzymes CYP85A1/A2 oxidases. Furthermore, 28-homoDL and CS can be produced from isofucosterol using a series of intermediates: 22-hydroxyisofucosterol, 6-deoxo-28-homoDS, and 28-homoDS. Castasterone can also be synthesized from β-sitosterol using intermediary compounds such as 22-homositosterol, 6-deoxohomositosterol, and 28-homoCS. It has been discovered that 28-homoTE, 28-homoTY, and 28-homoCS can be converted into 26-nor-28-homoTE, 26-nor-28-homoTY, and 26-nor-28-homoCS, respectively [116,124,125,126,127,128].
In the second pathway, 24-methylenelophenol is converted into campesterol, which acts as a precursor for C28-BRs (Figure 9) [122,129,130]. In this C28-BRs biosynthesis, two parallel pathways, i.e., the late and early C-22 oxidation pathways, occur. Campesterol is then converted into (24R)-ergostan-4-en-3β-one and campestanol (CN). In the parallel early C-22 oxidation pathway, C-22α hydroxylation of 24-methyleneCR by the enzyme C-22α hydroxylase leads to the biosynthesis of 22-hydroxy-24-methyleneCR [131]. Subsequent processes resemble the late C-22 oxidation pathway, resulting in the generation of 22-hydroxy variants of the related substances. The distinguishing aspect between the C-22 oxidation sub-pathways is the formation of 6-deoxocathasterone (6-deoxoCT) from (22S,24R)-22-hydroxy-5α-ergostan-3-one, circumventing the synthesis of CN. This process is referred to as the CN-independent route of BR biosynthesis. Nonetheless, during each phase of the late C-22 oxidation pathway, the compound involved could undergo hydroxylation via C-22α hydroxylase, leading to the formation of hydroxylated versions of compounds that are part of the early C-22 pathway [132]. Transformation in 22-hydroxymethyleneCR results in the creation of 6-deoxodolichosterone, which is converted into dolichosterone (DS), dolicholide (DL), castasterone (CS), and brassinolide (BL) [120,128].
The biosynthesis of C27-BR involves the late C6 oxidation pathway. In these reactions, cholesterol is converted into cholestanol by the 5α-reductase enzyme DET2. Consecutive reactions then produce a sequence of compounds: 6-deoxo-28-norcathasterone, 6-deoxo-28-norteasterone, 6-deoxo-28-nor-3-dehydroteasterone, 6-deoxo-28-nortyphasterol, and 6-deoxo-28-norcastasterone. In addition, the early C6 oxidation pathway starts with the oxidation of cholestanol to create 6-oxocholestanol. This process leads to the synthesis of a series of compounds, including 28-norcathasterone, 28-norteasterone, 28-nor-3-dehydroteasterone, 28-nortyphasterol, 28-norcastasterone, and, eventually, 28-norbrassinolide [124,133,134].
The late C6 pathway begins with the hydroxylation of CN to form 6-deoxocathasterone (6-deoxoCT), which is a process catalyzed by the enzyme 22α-hydroxylase. Interestingly, 6-deoxoCT can also be directly synthesized from 22-hydroxy-5α-ergostan-3-one. Following this step, 6-deoxoCT can be hydroxylated by C-23 hydroxylase to form 6-deoxoteasterone. This compound then undergoes C-3 oxidation to form 3-dehydro-6-deoxoteasterone (6-deoxo-3-DT) under the action of CYP90D C3-oxidase. Next, 6-deoxo-3-DT is converted into 6-deoxotyphasterol (6-deoxoTY) by the D11 CYP724B1 enzyme. In the subsequent step, 6-deoxoTY is hydroxylated to form 6-deoxocastasterone (6-deoxoCS) by the enzyme 2α-hydroxylase. 6-deoxoCS is then converted into castasterone (CS) by the enzymes BR-6-oxidase1 and BR-6-oxidase2. Afterward, CS is transformed into brassinolide (BL) via a process called Baeyer–Villiger oxidation, which is facilitated by the enzyme BR-6-oxidase2 (CYP85A2). At the start of the early oxidation pathway, CN is hydrated to form 6α-hydroxyCN. This compound is then oxidized to create 6-oxo-CN. The enzyme 22α-hydroxylase is responsible for the conversion of 6-oxo-CN into cathasterone (CT). CT then undergoes a series of transformations to form teasterone (TE), 3-dehydroteasterone, typhasterol (TY), CS, and, finally, BL [112,116,120,124,125,132,135].
9. Cytokinins
Cytokinins (CKs) regulate plant development, nutrient uptake, cell division, cell differentiation, chlorophyll senescence, apical dominance, embryonic development, and general aspects of plant growth [136]. Cytokinins are N6-substituted adenine derivative compounds that can be divided into two groups: (i) isoprenoid and (ii) aromatic CKs. Isoprenoid CKs more abundant than aromatic compounds [137,138]. Isoprenoid CKs include compounds such as N6-(∆2-isopentenyl adenine (iP), dihydrozeatin, and trans- and cis-zeatin (tZ and cZ, respectively), while N6-benzyladenine and its hydroxyl derivatives, such as ortho- and meta-topolin, are representatives of aromatic CKs [139].
Cytokinin levels are spatially and temporally regulated. Cytokinins are abundant in the apical meristems, root tips, and immature seeds. It is generally assumed that the root tip is the main site of the biosynthesis of CKs. On the other hand, literature data indicate that the shoot apex, cambium, and immature seeds are also capable of synthesizing CKs [140]. Changes in CK levels are related to the phase of cell cycle, consequences of influence of environmental factors, presence or lack of mineral nutrients, and effect of abiotic or biotic stress factors. The levels of active CKs in plants are highly regulated by the rates of production, interconversion, transport, and degradation [141].
Cytokinins can be synthesized de novo as nucleotide mono-, di-, or tri-phosphates, which are characterized by low biological activity. Moreover, tRNA is a source of CKs. The release of cytokinins from this nucleic acid leads to the creation of nucleotide monophosphates. The conversions of nucleotides, nucleosides, and free bases are catalyzed by enzymes involved in the metabolism of adenine [142]. In the next step, an isoprenoid moiety is added to the adenine present in the ATP and/or ADP molecule [143]. An alternative pathway, in which a hydroxylated side chain is added to the adenine moiety, has also been proposed [144].
DMAPP and HMBDP are the common isoprenoid side chain donors in biosynthetic pathway of CKs [102,145]. In the case of formation of isopentenyladenine-type CKs via the precipitation of DMAPP, the side chain is hydroxylated by cytochrome P450 mono-oxygenase (Figure 7) [146]. The CK nucleotides that include nucleotides released from tRNA are then hydrolyzed to free bases [102].
Cytokinin biosynthesis is catalyzed by the enzyme isopentenyl transferase (IPT) (Figure 10). There are two types of the adenylate IPT, which are responsible for attachment of an isopentenyl group to the N6 atom present in AMP, ADP, or ATP and tRNA. IPT does not add isopentenyl group to adenosine or adenine. IPT acts in the same way on adenine present in the structure of tRNA [143]. Adenylate IPT can synthesize iP and tZ nucleotides [145,147].
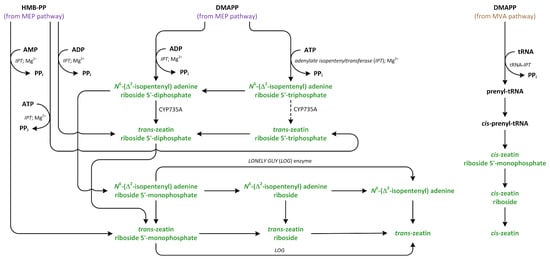
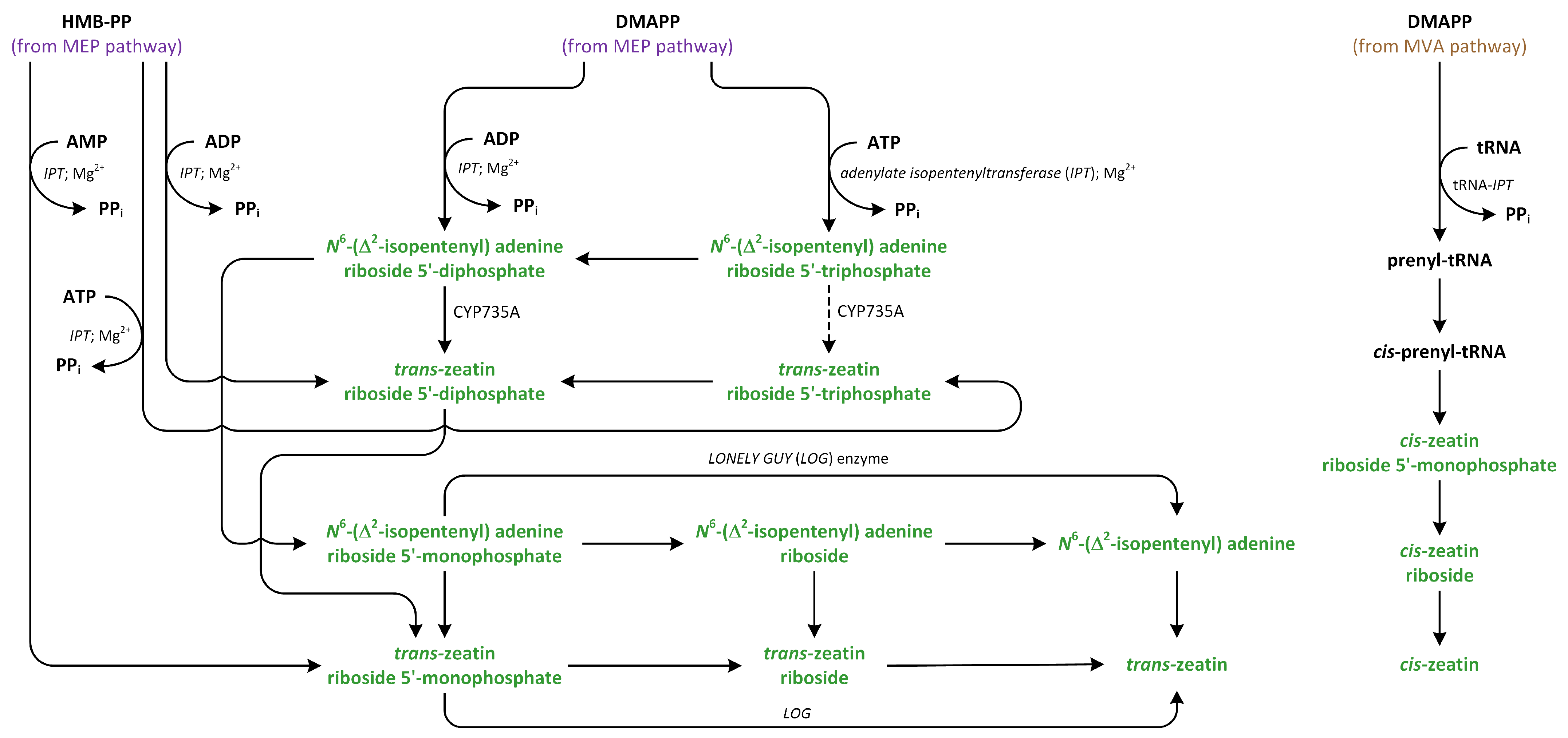
Figure 10.
The biosynthetic pathways of cytokinins (phytohormones are green).
Isopentenyl transferase isolated from vascular plants preferentially acts on ADP or ATP, rather than AMP, as prenyl acceptors and almost exclusively uses DMAPP as a prenyl donor, which, in turn, can synthesize iP riboside 5′-diphosphate or iP riboside 5′-triphosphate. Past kinetic studies of the activity of IPT in A. thaliana showed that the Km values for AMP are much higher than those of ADP and ATP, which were used as substrates. Moreover, the cellular level of AMP is low, indicating that AMP is not an essential substrate involved in IPT activity [143]. Interestingly, IPT isolated from mulberry (Morus alba) can also recognize and react with dATP, dADP, CDP, and GDP as substrates. In the case of GDP, the isopentenyl moiety is added to N2 group (the exocyclic nitrogen), rather than N1 group (the endocyclic nitrogen), which is closer to C6 [148,149]. However, IPT that is isolated from Agrobacterium cells, which is called Tmr, uses AMP as an acceptor and HMBDP or DMAPP as donors in vitro, while HMBDP is predominantly utilized in vivo, leading to the production of tZ, which is a highly active CK, in plastids of host plants [150]. IPTs are transported into different organelles, such as mitochondria, plastids, and the cytosol. Therefore, isoforms of IPT, such as AtIPT1, AtIPT3, AtIPT5, and AtIPT8, are present in the plastids and involved in the production of CKs from DMAPP that originate from the MEP pathway [151]. On the other hand, AtIPT4 is localized in cytosol, whereas AtIPT7 can be found in mitochondria. These observations suggest that DMAPP involved in CK biosynthesis is also derived via the MVA pathway [147].
Studies suggest that tRNA isopentenyltransferase (tRNA IPT) is responsible for the transportation of DMAPP via MVA pathway to adenine in tRNA, which results in the creation of cZ [143]. Studies of the synthesis of the isopentenyl side chain of the CK present in the tRNA molecule in vitro demonstrated that the isopentenyl group was derived from CKs MVA, and the turnover of CK-containing tRNA may serve as a minor source of free cZ in plant cells. It has been proposed that the target adenine is released from the tRNA structure in a similar way to that modified by other nucleic acid-editing enzymes. This type of IPT has been identified in almost all living organisms, including bacteria, yeast, animals, and plants. However, tRNA IPT was not found in Archaea. The isopentenylation of adenine present in tRNA has an effect on translational efficiency and precision. It improves proofreading during the translation process by decreasing misreading at the first position of the codon [146,152]. The different origins of large amounts of DMAPP utilized in biosynthesis of CKs, such as tZ and cZ, suggest that plants can precisely regulate the homeostasis of these CKs [147].
10. Gibberellins
Gibberellins (GAs) are tetracyclic and diterpenoid plant hormones that form a large group of carboxylic acids [153]. GAs are present in all vascular plants and lower plants, such as lycophytes and ferns. GAs are involved in the promotion of organ growth, enhance cell elongation and/or division, and activate most developmental processes (i.e., seed germination, induction of flowering, and maturation) in many plant species [154,155]. GAs were first isolated from the fungus Gibberella fujikuroi (reclassified as Fusarium fujikuroi), which stimulated growth in infected vascular plants. Next, their presence in plants as phytohormones was confirmed in the late 1950s. The system that numbers GAs, starting with GA1, assigns GAs in order of their discovery and structural characterization [156]. The most well-known and biologically active GAs in plants are GA1, GA3, GA4, and GA7. There are three structural properties that alter these GAs: the presence of hydroxyl group in C-3β, carboxyl group in C-6, and lactone between C-4 and C-10. The 3β-hydroxyl group can be replaced by other functional groups at positions C-2 and/or C-3. However, GA5 and GA6 are representatives of bioactive GAs that lack a hydroxyl group at C-3β [153].
The knowledge of GA biosynthesis has developed rapidly in recent years. Quantita-tive analysis shows that biosynthesis of GAs occurs in actively growing tissues, i.e., young leaves, shoot apices, and flowers. In contrast, there are some reports that indicate that xylem and phloem exudates contain Gas, suggesting the presence of long-distance transportation of these phytohormones in plants [154,157,158].
In plants, GAs are formed from GGPP through IPP, which represents the C-5 building moiety involved in the synthesis of terpenoid (isoprenoid) compounds (Figure 7) [155,159]. In the green tissues of plants, IPP is produced via two pathways: (i) the MVA pathway in the cytoplasm and (ii) the methylerythritol phosphate (MEP) pathway in plastids in two-step reactions. Such a metabolism is ancient and leads to the synthesis of 12,000 natural diterpenoid products in plant tissues. Recent data suggest that this compound is cyclized to the tetracyclic hydrocarbon precursor ent-kaurene in plastids by ent-copalyl diphosphate, which predominantly occurs through the MEP pathway [157]. However, the MVA pathway may also be involved in the synthesis of isoprenoid intermediates of GGPP and their transport from the cytosol into the plastids [160]. The formation of ent-kaurene from GGPP takes place in the stroma of proplastids or in the developing chloroplasts, except those of mature chloroplasts [161]. Ten functional GGPP synthase (GGPPS) genes were discovered in A. thaliana. Among these genes, seven encode enzymes were localized in the plastid. However, GGPPS11 is expressed most strongly and constitutively in plants that produce most of the substrate utilized in the biosynthesis of various terpenoids in plastids [162]. In contrast to A. thaliana, rice is reported to contain one functional GGPPS that is present in the plastid, which is involved in the biosynthesis of all diterpenoid compounds, including ent-kaurene, which is a key intermediate involved in the formation of GAs [163].
Firstly, the synthesis of ent-kaurene from GGPP starts via cyclization, which initiated by the proton of the dicyclic ent-copalyl diphosphate (CPP), which is catalyzed by a type II diterpene cyclase: ent-copalyl diphosphate synthase (CPS) (Figure 11). CPS contains a conserved DXDD motif, in which aspartate gives a proton to initiate this reaction via cyclization. Additionally, a water molecule attached to amino acids (histidine and asparagine) plays role as the catalytic base that accepts a proton and terminates the reaction [164]. CPS activity is inhibited by high concentrations of both Mg2+ and GGPP, which are promoted by light, leading to reduced flux into the GA pathway during de-etiolation. This process is part of the mechanism used to decrease GA biosynthesis and its cellular concentration [165].
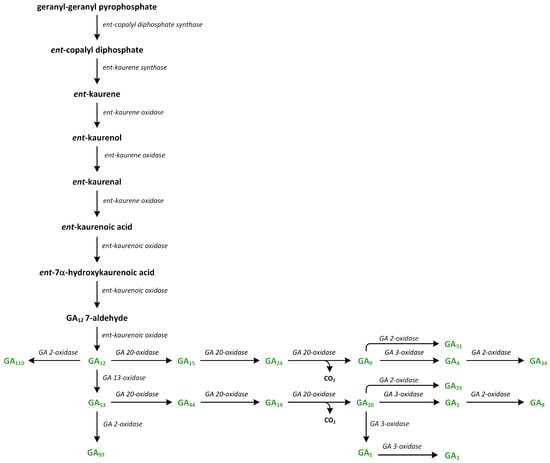
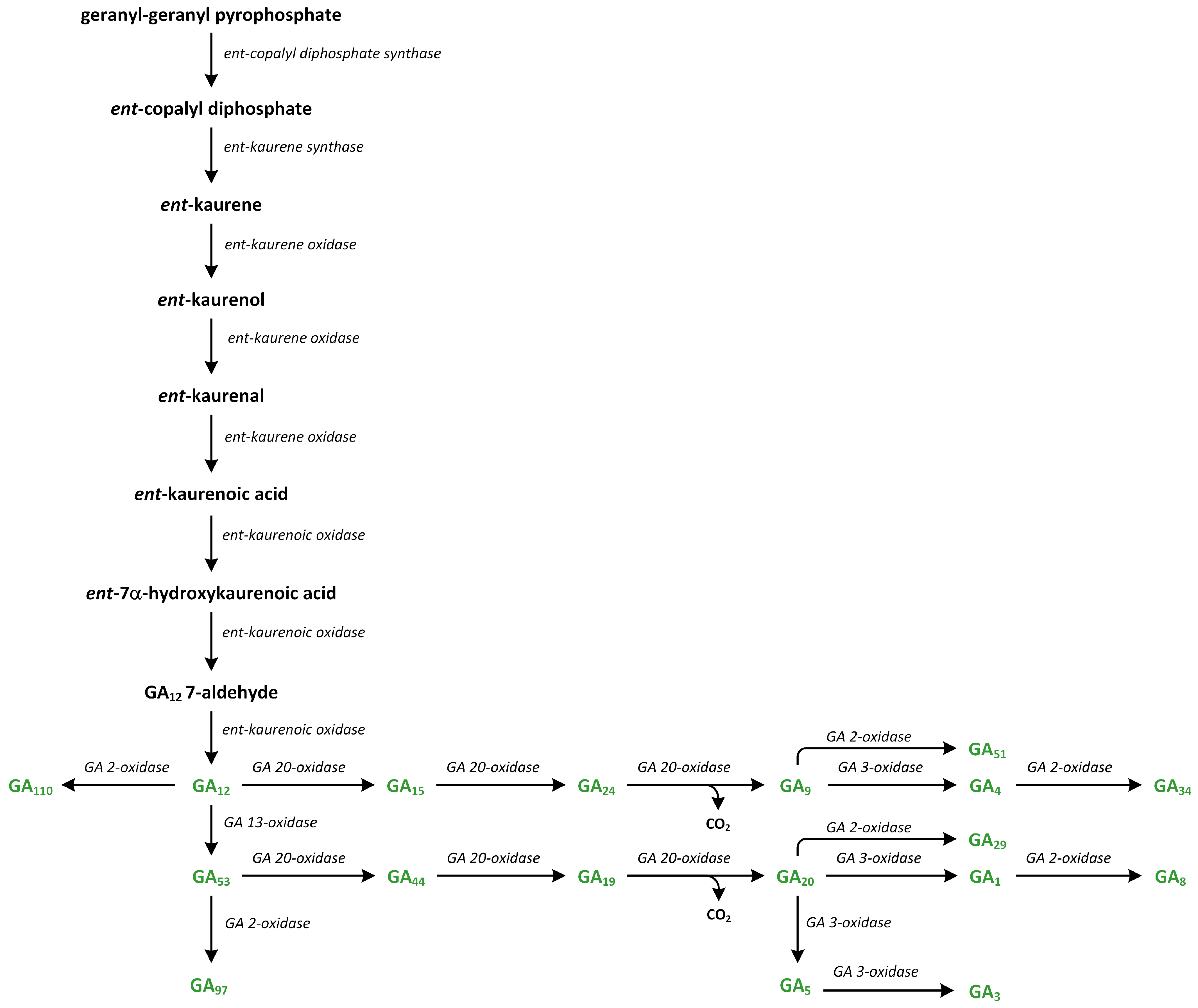
Figure 11.
The biosynthetic pathways of gibberellins (phytohormones are green).
The second step of the transformation of ent-copalyl diphosphate to ent-kaurene is catalyzed by a type I cyclase, which is known as ent-kaurene synthase (KS) (Figure 11). Cyclization reaction is initiated via metal-dependent heterolytic cleavage of the O-C bond. A pimeren-8-yl carbocation takes place, which then undergoes rearrangement. The loss of H+ is important step that leads to tetracyclic ent-kaurene [166]. Enzyme KS contains RLX(N,D)DXX(S,T,G)XXX(E,D) and DDXXD motifs, which can bind to ion Mg2+, which is associated with the diphosphate residue and participates in its ionization [167]. The lycophyte Selaginella moellendorfii, which is one of the first plants to have evolved the ability to synthesize GAs, possesses monofunctional CPS and KS enzymes, which are characteristic of vascular plants. In S. moellendorffii, KS is characterized by low substrate specificity, as it converts different stereoisomeric forms of CPP to various organic compounds, while KS enzymes in angiosperms are specific only for ent-CPP [168]. In angiosperms, CPS and KS possess the functional diversification required to produce the diterpenoids involved in plants’ defense responses [157,169].
GA12, which is the first C20-GA, is formed from ent-kaurene, and the reactions of these synthetic steps of GA productions are catalyzed by the following enzymes: ent-kaurene oxidase (KO), two cytochrome P450 mono-oxygenases, and ent-kaurenoic acid oxidase (KAO) (Figure 11) [170]. In A. thaliana, KO is associated with the outer chloroplast membrane and the endoplasmic reticulum. On the other hand, two KAOs are located in the endoplasmic reticulum. This observation suggests that ent-kaurene is oxidized and then transported from plastids to the endoplasmic reticulum [157]. KO catalyzes the oxidation of ent-kaurene into ent-kaurenoic acid via repeated hydroxylation of C-19. The intermediate aldehyde, i.e., ent-kaurenal, is present in this reaction [171]. The first reaction of hydroxylation to ent-kaurenol is the rate-limiting step in the GA biosynthesis pathway [172]. The reactions catalyzed by KAO have been studied in cells that were isolated from developing seeds, i.e., endosperm [158].
In plants, ent-kaurenoic acid is converted into GA12 in three steps that involve 7β-hydroxy-ent-kaurenoic acid and GA12-aldehyde. Enzyme KAO is involved in this conversion (Figure 11). The first step involves stereospecific hydroxylation of C-7β. In the second reaction (initiated via stereospecific loss of the 6β-H), ring B contracts between 5 and 6 carbon atoms via migration of the C-7–C-8 bond from C-7 to C-6. The extrusion of C-7 as the aldehyde is observed. In the third step, GA12-aldehyde is oxidized to GA12. Cucurbitaceae plants, such as Cucurbita maxima and Cucumis sativa, contain GA7 oxidases (GA7ox) that can convert GA12-aldehyde into GA12. These enzymes are present in the development of seeds and vegetative tissues. The pathway branches from GA12. The reaction of 13-hydroxylation to GA53 initiates the formation of 13-hydroxylated GAs, such as GA1. Additionally, a parallel non-13-hydroxylation route from GA12 results in the synthesis of GA4 [158].
In higher plants, GA12 is present at a branch point along the GA pathway. GA12-aldehyde can be converted into GA12 or transformed into GA53 via the hydroxylation at C-13 (13-hydroxylated GA) (Figure 11). GA12 and GA53 are precursors of the non-13-hydroxylation and 13-hydroxylation pathways, respectively. These GAs can be oxidized at C-20, producing GA9 and GA20, respectively [159].
In rice, two cytochrome P450s—CYP714B1 and CYP714B2—are responsible for 13-hydroxylation of GA12. Although vegetative rice tissues contain predominantly 13-hydroxylated GAs, the overexpression of the CYP714B gene, which resulted in an increase in the concentration of various 13-hydroxy types of GAs, including GA1, also caused semidwarfism [173]. However, A. thaliana has two members of the CYP714A subfamily that are highly expressed in developing seeds. CYP714A1 transforms GA12 into 16α-carboxy-17-norGA12, whereas CYP714A2 converts GA12 into 12α-hydroxyGA12 (GA111) and, at a lower level of efficiency, GA53 via the 13-hydroxylation process. This enzyme is also capable of hydrolyzing ent-kaurenoic acid in the C-13 position to steviol [174]. Another member of the CYP714 family that is present in rice—CYP714D1—catalyzes the reaction of the epoxidation of the 16,17-double bond of 13-deoxyGAs, including GA12. This reaction leads to synthesis of inactive compounds [157]. Thus, members of the CYP714 family are responsible for the inactivation of GAs and ent-kaurenoids via their oxidation at the C- and D-rings. A. thaliana contains eight tandem CYP72A genes. Among these genes, CYP72A9 was shown to 13-hydroxylate GA12, as well as GA9 and GA4. Therefore, CYP72A9 is the main source of the 13-hydroxy types of Gas [175].
The transformation of GA12 and GA53 into GA9 and GA20, respectively, are catalyzed in plants by a family of 2-oxoglutarate/Fe(II)-dependent dioxygenases, which are also known as GA 20-oxidases (GA20ox) (Figure 11). These enzymes are responsible for the cleavage of C-20 with 19,10-γ-lactone formation, which is a characteristic of C19-GA. Firstly, C-20 methyl is oxidized into alcohol form. Next, it is transformed into the aldehyde. During this oxidation process, C-20 is removed as CO2, and 20-oic acid is a minor by-product. However, this biochemical mechanism is not well understood [176]. Vascular plants contain a family of GA20ox genes, which have different developmental, environmental, and tissue expression patterns. In many plants, GA20ox proteins are involved in the regulation of the GA rate of the synthesis of GA, and expressions of GA20ox genes are tightly regulated by developmental and environmental signals to maintain GA homeostasis [177].
In the final step in the biosynthesis of bioactive GAs, the C19-GAs (i.e., GA9 and GA20) are 3β-hydroxylated to GA4 and GA1, respectively, by the enzyme GA 3-oxidase (GA3ox) (Figure 11) [178]. GA4 is the first biologically active GA. This GA is desaturated to GA7, which is then converted into GA3 via the late 13-hydroxylation reaction. Enzymes GA3ox are highly active in vegetative tissues and function only as 3β-hydroxylases. They are characterized by high regiospecificity in dicots. On the other hand, GA3ox in monocots are less specific to the substrates. Therefore, GA3 is synthesized from GA20 as a minor by-product via the GA1 production route. The oxidation at the 2β and 3β positions leads to formation of the 2,3-unsaturated intermediate GA5, which is then converted into GA3 via oxidation at C-1β. This reaction is catalyzed by the same enzyme [157]. Therefore, monocots contain low levels of GA3, which are usually less than 10% of the GA1 content, while they are generally undetectable in vegetative tissues of dicots. In plants, the last step in GA synthesis is the 3β-hydroxylation of GA9 and GA20 into GA4 and GA1, respectively. GA4 and GA1 are the main GAs. The GA3ox genes are present in small families. A. thaliana contains four members, while rice and barley (Hordeum vulgare) contain only two members. Two enzymes of GA3ox—AtGA3ox1 and AtGA3ox2—present in A. thaliana, as well as only one enzyme found in cereals, are involved in the development of vegetative organs [178].
11. Strigolactones
Strigolactones (SLs), which are carotenoid-derived terpenoids, were initially characterized as germination stimulants for root parasitic plant witchweed (Striga spp.). SLs are a class of carotenoid-derived terpenoids that are involved in diverse developmental processes, such as seed germination, shoot branching, leaf senescence, and root development, as well as responses to various environmental conditions (e.g., light stress and high-temperature stress) [179]. They are also known as rhizosphere chemical signals that can regulate interactions between arbuscular mycorrhizal fungi and root parasitic plants. These groups of phytohormones also affect photosynthesis, synthesis, and action of other phytohormones, as well as regulating the levels of different metabolic compounds [180,181].
According to their chemical structures, SLs can be classified into two groups: canon-ical and non-canonical SLs. Canonical SLs contain a tricyclic lactone structure composed of three rings (ABC-rings) connected to a butenolide group (D-rings) through an enol-ether bridge, which is critical for the performance of their biological activities in plants. In contrast, non-canonical SLs do not have typical ABC-rings, though they contain both an enol-ether bridge and D-ring moieties [182].
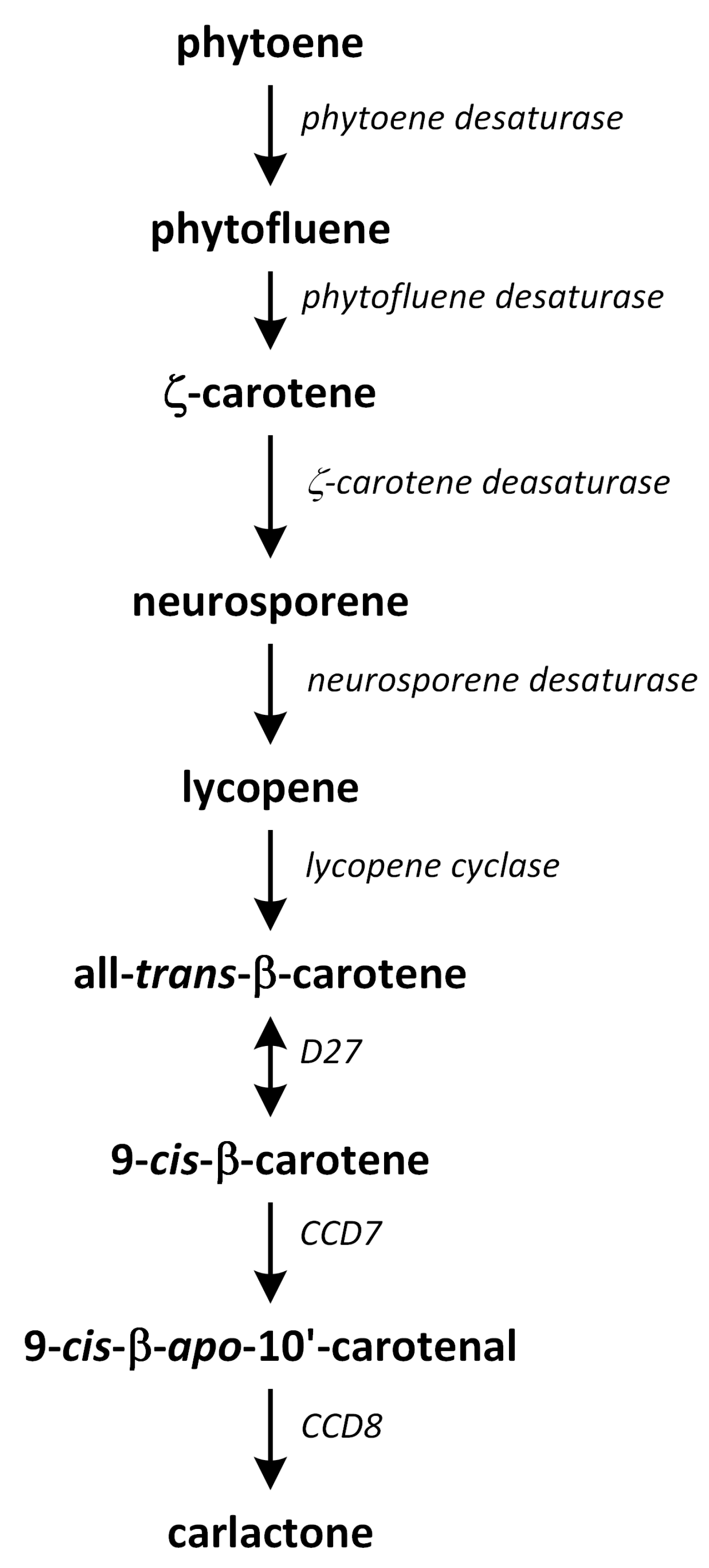
Strigolactone biosynthesis mainly occurs in roots. The intermediates in the SL synthetic pathway have been also found in the stem. The biosynthesis of this group of phytohormones is tightly regulated by environmental conditions, such as starvation of phosphate or drought stress [183]. The early steps of SL biosynthesis are common for each type and may occur via the MVA or MEP pathways that lead to IPP synthesis, which is sequentially condensed to create DMAPP, geranyl pyrophosphate (GPP), farnesyl pyrophosphate (FPP), and GGPP (Figure 7) [116]. Thus, the first part of the SL biosynthetic pathway starts in chloroplasts. The condensation of two GGPP molecules by phytoene synthase generates phytoene (Figure 12), which is the first uncolored carotenoid [184]. Next, phytoene is transformed via a sequence of reactions of desaturation and cis/trans-isomerization into all-trans-lycopene. Cyclization reactions include conversion of all-trans-lycopene into all-trans-β-carotene and all-trans-α-carotene [185,186]. Dwarf27 (D27) isomerizes all-trans-β-carotene to 9-cis-β-carotene, followed by cleavage induced by carotenoid removal of enzyme dioxygenase 7 (CCD7) from 9-cis-β-apo-10′-carotenal, which undergoes cleavage and rearrangement reactions to create carlactone (CL), which is the central intermediate in SL biosynthesis (Figure 12). This reaction is catalyzed by the CCD8 enzyme [187]. CCD7 is encoded by gene MAX3 and its orthologs genes: RMS5 and D17/HTD1. Gene MAX4 and its orthologs RMS1, D10, and Defective in Anther Dehiscence 1 (DAD1) encode CCD8. Both enzymes act in a progressive manner [188].
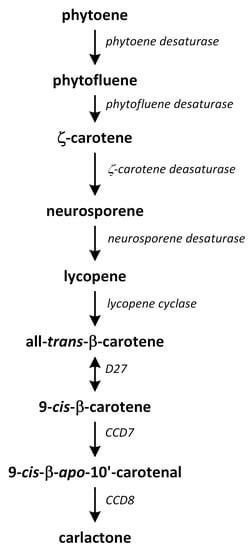
Figure 12.
The biosynthetic pathways of carlactone.
Carlactone (CL) contains only A- and D-rings that have an enol-ether bridge and exhibits SL-like properties. For example, CL inhibits shoot branching in SL biosynthetic mutants (i.e., rice, and A. thaliana) and promotes seed germination of Striga hermonthica. CL has been identified as an endogenous precursor in synthetic routes of both canonical and non-canonical SLs [186,189,190]. Furthermore, a new compound, which we identified as 3-hydroxyCL, is formed in vitro using 9-cis-3-hydroxy-β-apo-10′-carotenal in a reaction catalyzed by enzymes D27, CCD7, and CCD8. This compound was identified in plants such as rice, A. thaliana, and N. benthamiana [191].
The SL biosynthesis activates the biochemical conversions of CL through the participation of cytochrome P450 mono-oxygenases and other enzymes, which are responsible for the biological and structural diversity of these phytohormones. Subsequently, CL is transported from the chloroplast to the cytoplasm, where the reactions related to SL biosynthesis occur (Figure 13) [190].
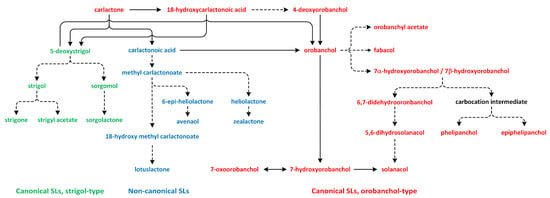
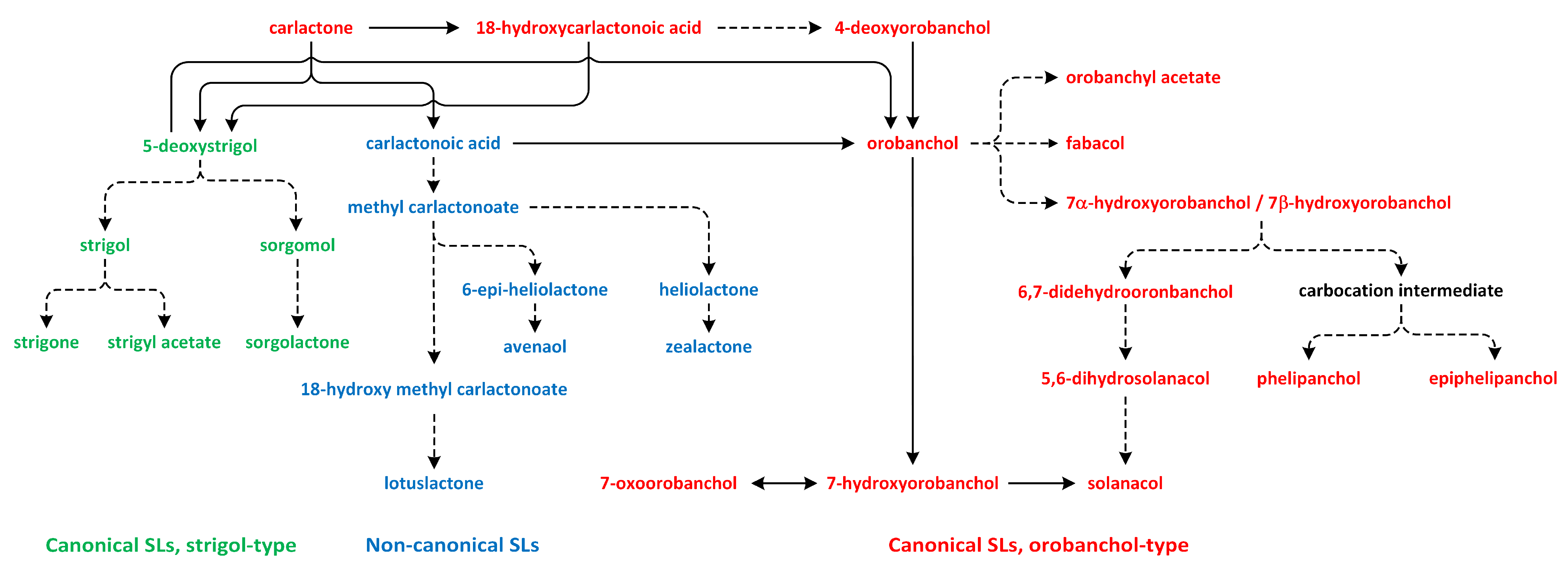
Figure 13.
The biosynthetic pathways of different types of strigolactones (phytohormones are green, blue, and red).
The oxidation of CL results in the formation of carlactonoic acid (CLA) (Figure 13). This reaction is catalyzed by the More Axillary Growth 1 (MAX1) enzyme, which is a member of the CYP711A subfamily. The structure and biochemical properties of MAX1 have been well characterized. It is a conserved protein that possesses many homologs in a number of plant species. CLA can be treated as the precursor of strigol and orobanchol-type SL, i.e., 5-deoxystrigol (5DS) and 4-deoxyorobanchol (4DO). Enzymes identified in plants, such as Os900/OsCYP711A2 in rice (Oryza sativa), VuCYP722C in cowpea (Vigna unguiculata), SlCYP722C in tomato (Solanum lycopersicum), and GaCYP722C in cotton (Gossypium arboreum), take part in the synthesis of organic compounds 4-deoxyorobanchol, orobanchol, and 5-deoxystrigol, respectively, from CLA [189,190,192,193,194].
In the next reaction, CLA is transformed to methyl carlactonoate (MeCLA) by an unknown enzyme [189]. Next, Lateral Branching Oxidoreductase (LBO) converts methyl carlactonoate (MeCLA) into hydroxymethyl carlactonoate (1′-OH-MeCLA) and CLA [195]. LBO can produce CLA via direct demethylation of MeCLA or indirect spontaneous reversion of 1′-OH-MeCLA to CLA [196]. MeCLA is considered to be a precursor used for the synthesis of non-canonical SLs. Feeding experiments showed that MeCLA can be converted into heliolactone in sunflower and tobacco [193,194]. The enzyme responsible for the production of heliolactone has not yet been identified to confirm this theory, and it is also important to study the possibility that MeCLA is an important intermediate for the synthesis of other non-canonical SLs (e.g., avenaol and zealactone) (Figure 13) [196]. The enzymes that lay downstream and upstream of CL may be important for the production of various compounds that belong to the SL family. Carotenoid isomerase, CCD7, and CCD8 are able to transform all-trans-β-carotene into CL and convert 3-hydroxy-carlactone (3-OH-CL) through zeaxanthin [191]. Hydroxy-carlactone derivatives are the main SL in A. thaliana [195]. However, their roles in the regulation of plant growth and development have not yet been confirmed [196]. These non-canonical SLs are characterized by lower stability in relation to canonical SLs. Thus, studies of the biological functions of non-canonical SLs are difficult to carry out, and the results are incomplete [197].
12. Jasmonates
Jasmonates are plant hormones that originate from oxylipins involved in plant devel-opment and stress responses [198]. Their major representatives are the isomers of jasmonic acid (JA): (+)-7-iso-JA and (−)-JA. JA can be metabolized into methyl jasmonate (MeJA), which is a volatile phytohormone. Jasmonates have previously been defined as JA and its diverse metabolites derived from various reactions created via esterification, methylation, sulfation, glycosylation, conjugation, de-carboxylation, hydroxylation, and carboxylation [199]. Recently, an alternative and probably more ancient pathway was proposed. In this route, JA originates from dinor-OPDA (2,3-dinor-12-oxo-10,15(Z)-phytodienoic acid [dn-OPDA]) [200]. Thus, 12-oxo-10,15(Z)-phytodienoic acid (OPDA), dn-OPDA, and their derivatives are members of JAs family. JA and MeJA are the best characterized group of jasmonates in plants, and they are regarded as one of the major phytohormones that regulate both defense responses and the development process [201].
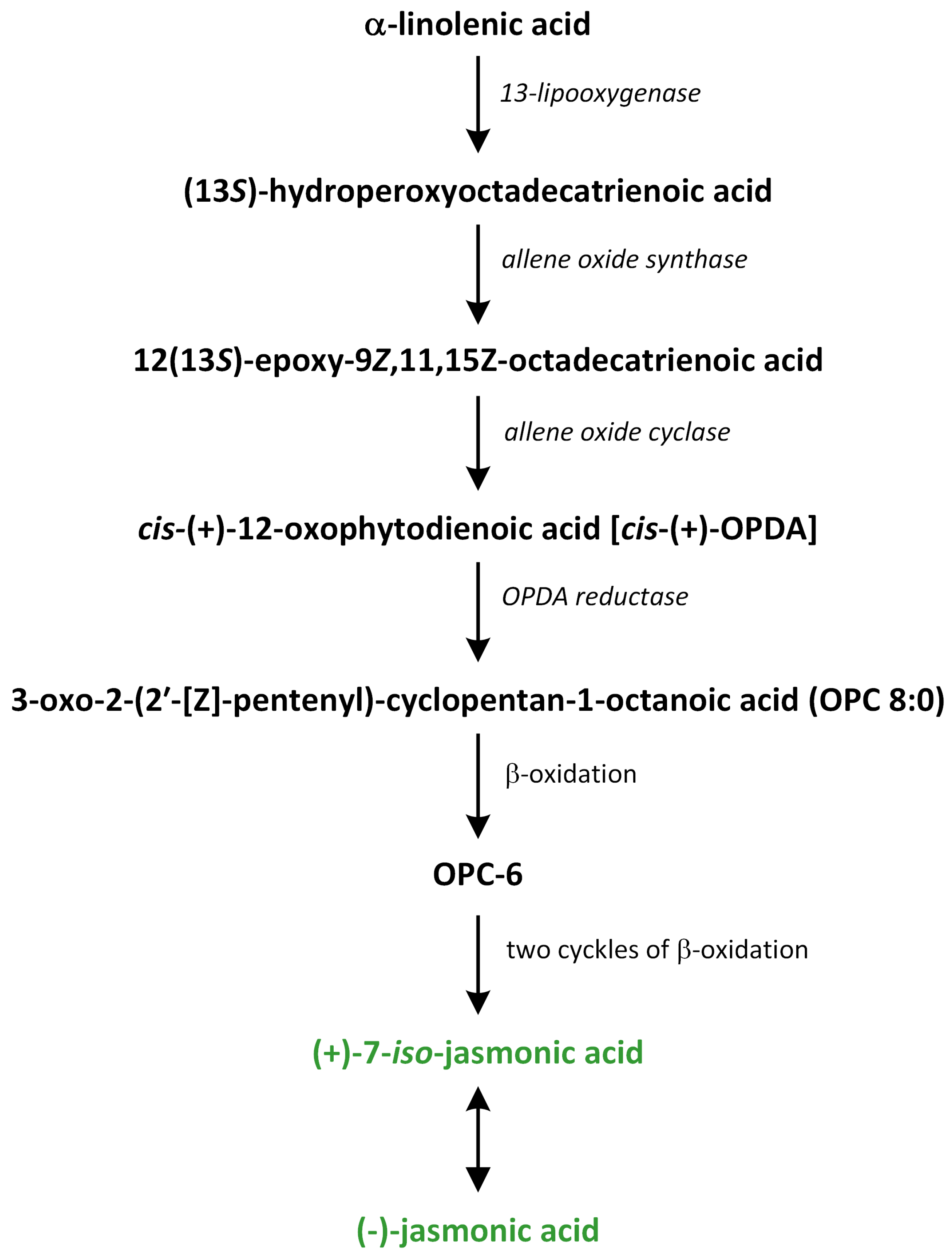
Jasmonate biosynthesis pathways have been extensively investigated in dicotyledonous plants, such as A. thaliana, tobacco, and tomato (Figure 14).
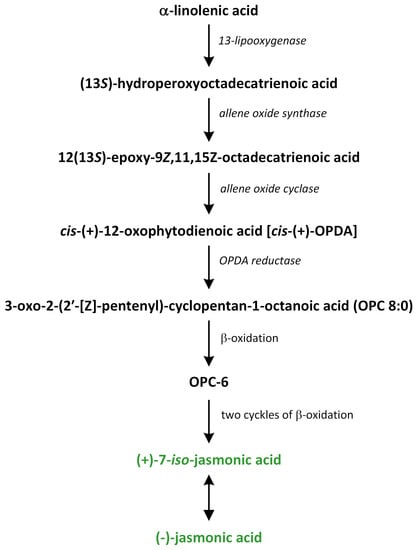
Figure 14.
The biosynthetic pathways of jasmonic acid (phytohormones are green).
In monocotyledonous species, only a few biosynthetic enzymes of JA have been identified and characterized [202,203,204]. JA biosynthesis is initiated via the release of the polyunsaturated fatty acid α-linolenic acid 18:3 (α-LeA) from galactolipids, which are present in chloroplast membranes. This reaction of hydrolysis is catalyzed by phospholipase A1 (PLA1), which is characterized by specificity of sn-1 in the structure of fatty acids. Thus, α-LeA oxygenation is the first step in JA biosynthesis. The flower-specific protein DAD1, which is a PLA1, is required for the formation of JA [205,206].
The formation of JA occurs via one of the seven different branches of lipoxygenase (LOX) pathway [199]. Enzymes LOXs (lipoxygenases) are non-heme-iron-containing dioxygenases that produce hydroperoxides of fatty acid from polyunsaturated substrates. These enzymes can be sub-divided into 9-LOX and 13-LOX types, depending on the carbon atom to which molecular oxygen is added. In A. thaliana, the 13-LOX enzymes, such as AtLOX2, 3, 4, and 6, are pre-sent to the chloroplast. On the other hand, 9-LOX members (e.g., AtLOX1 and AtLOX5) are probably localized in the cytosol, although their functions are poorly understood [207,208]. The distribution and transport of α-LeA between the organelles, such as chloroplast and the cytosol, regulate the metabolic flow between 13-LOX- and 9-LOX-derived oxylipins [202].
In the chloroplast, (13S)-hydroperoxy octadecatrienoic acid (13-HPOT) is formed after oxygenation of α-LeA by the enzyme 13-LOX (Figure 14). This compound can play a role as a substrate of other enzymes, such as cytochrome P450 enzymes that belong to the CYP74 family, hydroperoxide lyase, allene oxide synthase (AOS), divinyl ether synthase, and epoxyalcohol synthase [209]. The next enzyme involved in the octadecanoid pathway is responsible for the biosynthesis of JA is AOS, which prefers to use 9- or 13-hydroperoxide derivatives as substrates. Thus, like LOX, this enzyme is characterized by 9- and 13-positional specificity. AOS is responsible for the formation of unstable allylic epoxides that can be spontaneously hydrolyzed in the presence of molecules of water in 13-hydroxy-12-oxo-octadecadienoic (α-ketols) and 9-hydroxy-12-oxo-octadecadienoic acids (γ-ketols). Moreover, the cyclization to a racemic mixture of cis-(+) and cis-(−) enantiomers is observed [210]. The cis-(+) isomer of oxylipin OPDA is produced in this reaction. This reaction requires the neighborhood of AOC. The stereochemistry of the pentenyl side chain and the carboxylic acid side chain at positions 3 and 7, respectively, of the pentanone ring in the JA structure is fixed in the step catalyzed via AOC and conserved in later reactions. In the cytosol, α-LeA can be oxygenated by enzyme 9-LOX and, subsequently, by enzyme 9-AOS to create two products: α-ketols and/or γ-ketols. On the other hand, α-LeA can also be oxygenated by α-dioxygenase (α-DOX) [211].
OPDA is known as a biosynthetic precursor of the phytohormone JA. It is also in-volved in various biological processes, which range from plant defence and stress responses to the regulation of growth and development [212]. OPDA is produced in the chloroplast through the 13-LOX/AOS pathway and then transported to peroxisomes. Recently, a plastid inner envelope-localized protein, known as OPDAT1, has been found in Populus trichocarpa. This observation suggested that this protein may participate in the export of OPDA from the chloroplast to other organelles [213]. OPDA can enter the peroxisome via diffusion of the protonated form of OPDA along its gradient. The cyclopentenone ring of OPDA after transportation into the peroxisome is reduced in the reaction catalyzed by the flavin-dependent oxidoreductase OPDA reductase 3 (OPR3). 3-oxo-2-(20-[Z]-pentenyl)-cyclopentane-1-octanoic acid (OPC-8) is produced during this step [214]. In the next reaction, OPC-8 is activated via esterification to CoA in the biochemical reaction catalyzed by OPC-8 coenzyme A ligase1, which is a member of the acyl-activating family of enzymes [215]. OPC-8 is then transformed into JA through three rounds of sequentially catalyzed peroxisomal β-oxidation reactions by acyl-CoA oxidase, which is a multifunctional protein, and L-3-ketoacyl-CoA thiolase [202]. During these reactions, six carbons are eliminated from the carboxy-terminal carbon chain. Therefore, the pentenyl side chain of JA is shortened in reactions that involve the ß-oxidation mechanism of the fatty acid [206].
Recently, an alternative pathway for JA synthesis has been proposed. Studies of the opr3-3 mutant in which OPR3 activity is completely depleted showed that OPDA and hexadecatrienoic acid-derived dn-OPDA are metabolized to create tetranor-OPDA and 4,5-didehydro-JA. Next, this compound may be reduced to JA by OPR2 after being transported into the cytosol. This process suggests that the OPR2-dependent and OPR3-independent JA biosynthetic pathways may be very ancient in vascular plants and activated under different processes or the influence of certain environmental conditions [200,216,217].
13. Conclusions
In this review, the biosynthesis routes of all known plant phytohormones have been explored. Understanding these intricate biosynthetic pathways of plant hormones will provide valuable insights regarding their biological functions.
The biosynthesis of PAs begins with putrescine that leading to the production of other PAs. The ethylene synthesis pathway is more straightforward, with methionine being a precursor. Salicylic acid biosynthesis is believed to occur via two routes: the isochorismate synthase pathway and the phenylalanine ammonia-lyase pathway. Auxin synthesis in plants occurs via two main pathways: the Trp-dependent and independent routes. Four Trp-dependent IAA routes have been proposed: the IAM, IPA, TAM, and IAOX routes. Similarly, melatonin biosynthesis starts with Trp, which is synthesized de novo via the shikimate pathway. Two potential pathways of ABA biosynthesis have been identified in plants. The first pathway is direct, and the second pathway is indirect; in these pathways, ABA is derived from C15 farnesyl pyrophosphate and a C40 carotenoid, respectively. Brassinosteroid biosynthesis occurs through three pathways that lead to C27-, C28-, and C29-type BRs, with early steps happening via the MVA or MEP pathways. Cytokinins can be synthesized de novo as low-activity nucleotide mono-, di-, and tri-phosphates. Alternatively, cytokinins can be released from tRNA, creating nucleotide mono-phosphates, with the interconversions of nucleotides, nucleosides, and free bases catalyzed by enzymes involved in adenine metabolism. Gibberellins are formed from GGPP through IPP, which is the C-5 building block of all terpenoid/isoprenoid compounds, including BRs, CKs, ABA, and SLs. Carlactone is the main precursor of SL production in plants. Jasmonic acid synthesis begins with the release of polyunsaturated fatty acid α-linolenic acid from galactolipids present in chloroplast membranes.
The emergence of various phytohormone groups, along with their unique biosynthetic pathways, could be linked to their distinct chemical structures, biosynthetic path-ways, physicochemical properties, plant evolutionary histories, and roles in plants’ adaptation to diverse environmental conditions or developmental programs. Precursors or intermediates involved in one phytohormone pathway may also participate in the biosynthesis of another hormone, thus linking the pathways (Figure S1).
Supplementary Materials
The following supporting information can be downloaded via the following link: https://www.mdpi.com/article/10.3390/metabo13080884/s1, Figure S1: Network of plant hormones’ biosynthetic pathways (all-in-one).
Author Contributions
Conceptualization, A.B.; methodology, A.B.; writing—original draft preparation, A.P.-N.; writing—review and editing, A.B. and A.P.-N.; visualization, A.B.; supervision, A.B. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Acknowledgments
We express our sincere gratitude to the Reviewers and Academic Editors who dedicated their time and valuable insights to the process of reviewing our manuscript. A.B. thanks Adam Bajguz for his consultation support.
Conflicts of Interest
The authors declare no conflict of interest.
Abbreviations
| 1′-OH-MeCLA | 1′-hydroxymethyl carlactonoate |
| 3-DT | 3-dehydroteasterone |
| 4DO | 4-deoxyorobanchol |
| 5DS | 5-deoxystrigol |
| 6-deoxo-3-DT | 3-dehydro-6-deoxoteasterone |
| 6-deoxoCT | 6-deoxocathasterone |
| ABA | Abscisic acid |
| ACC | 1-aminocyclopropane-1-carboxylic acid |
| ACO | ACC oxidase |
| ACS | ACC synthase |
| ADC | Aginine decarboxylase |
| Agm | Agmatine |
| AO1 | Aldehyde oxidase 1 |
| AOS | Allene oxide synthase |
| Arg | Arginine |
| ASMT | N-acetylserotonin O-methyltransferase |
| BL | Brassinolide |
| BR | Brassinosteroid |
| CCD7 | Carotenoid removal dioxygenase 7 |
| CK | Cytokinin |
| CL | Carlactone |
| CLA | Carlactonoic acid |
| CN | Campestanol |
| COMT | Caffeic acid O-methyltransferase |
| CPP | Ent-copalyl diphosphate |
| CPS | Ent-copalyl diphosphate synthase |
| CR | Cholesterol |
| CS | Castasterone |
| CT | Cathasterone |
| cZ | Cis-zeatin |
| DAD1 | Defective in Anther Dehiscence 1 |
| dcSAM | Decarboxylated S-adenosyl-L-methionine |
| DL | Dolicholide |
| DMAPP | Dimethylallyl pyrophosphate |
| dn-OPDA | 2,3-dinor-12-oxo-10,15(Z)-phytodienoic acid |
| DS | Dolichosterone |
| DOXP | 1-deoxy-D-xylulose-5-phosphate |
| DXS | DOXP synthase |
| FPP | Farnesyl pyrohosphate |
| GA | Gibberellin |
| GGPP | Geranylgeranyl pyrophosphate |
| GGPPS | Geranylgeranyl pyrophosphate synthase |
| GPP | Geranyl pyrophosphate |
| HMBDP | E-4-hydroxy-3-methyl-but-2-enyl diphosphate |
| IAA | Indole-3-acetic acid |
| IAM | Indole-3-acetamide |
| IAN | Indole-3-acetonitrile |
| IAOX | Indole-3-acetaldoxime |
| ICS | Isochorismate synthase |
| ICS-Glu | Isochorismate-9-glutamate |
| iP | N6-∆2-isopentenyl adenine |
| IPA | Indole-3-pyruvic acid |
| IPP | Isopentenyl diphosphate |
| JA | Jasmonic acid |
| KAO | Ent-kaurenoic acid oxidase |
| KO | Ent-kaurene oxidase |
| KS | Ent-kaurene synthase |
| LBO | Lateral Branching Oxidoreductase |
| MAX1 | More Axillary Growth 1 |
| MeJA | Methyl jasmonate |
| MEP | Methylerythritol phosphate |
| MTA | 5′-methylthioadenosine |
| MVA | Mevalonate |
| NCPA | N-carbamoyl putrescine |
| ODC | Ornithine decarboxylase |
| OPC-8 | 3-oxo-2-(20-[Z]-pentenyl)-cyclopentane-1-octanoic acid |
| OPDA | 12-oxo-Z-phytodienoic acid |
| OPR3 | Oxidoreductase OPDA reductase 3 |
| Orn | Ornithine |
| PA | Polyamine |
| PAL | Phenylalanine ammonia-lyase |
| PLP | Pyridoxal-5′-phosphate |
| Put | Putrescine |
| SA | Salicylic acid |
| SAM | S-adenosyl-L-methionine |
| SAMDC | S-adenosyl-L-methionine decarboxylases |
| SL | Strigolactone |
| SMO1 | C-4 sterol methyl oxidase 1 |
| SNAT | Serotonin N-acetyltransferase |
| Spd | Spermidine |
| Spm | Spermine |
| SPMS | Spermine synthase |
| T5H | Tryptamine 5-hydroxylase |
| TAM | Tryptamine |
| TDC | Tryptophan decarboxylase |
| TPH | Tryptophan hydroxylase |
| Trp | Tryptophan |
| TY | Typhasterol |
| tZ | Trans-zeatin |
| α-LeA | α-linolenic acid 18:3 |
References
- Majumdar, R.; Barchi, B.; Turlapati, S.A.; Gagne, M.; Minocha, R.; Long, S.; Minocha, S.C. Glutamate, ornithine, arginine, proline, and polyamine metabolic interactions: The pathway is regulated at the post-transcriptional level. Front. Plant Sci. 2016, 7, 78. [Google Scholar] [CrossRef] [PubMed]
- Li, M.X.; Zhu, Y.C.; Li, S.S.; Zhang, W.; Yin, C.X.; Lin, Y.J. Regulation of phytohormones on the growth and development of plant root hair. Front. Plant Sci. 2022, 13, 865302. [Google Scholar] [CrossRef] [PubMed]
- Song, X.-F.; Ren, S.-C.; Liu, C.-M. Peptide hormones. In Hormone Metabolism and Signaling in Plants; Li, J., Li, C., Smith, S.M., Eds.; Academic Press: London, UK, 2017; pp. 361–404. [Google Scholar] [CrossRef]
- Royek, S.; Bayer, M.; Pfannstiel, J.; Pleiss, J.; Ingram, G.; Stintzi, A.; Schaller, A. Processing of a plant peptide hormone precursor facilitated by posttranslational tyrosine sulfation. Proc. Natl. Acad. Sci. USA 2022, 119, e2201195119. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, Y.-C.; Yamada, M. The roles of peptide hormones and their receptors during plant root development. Genes 2021, 12, 22. [Google Scholar] [CrossRef] [PubMed]
- Dodueva, I.; Lebedeva, M.; Lutova, L. Dialog between kingdoms: Enemies, allies and peptide phytohormones. Plants 2021, 10, 2243. [Google Scholar] [CrossRef] [PubMed]
- Davies, P.J. (Ed.) Plant Hormones. Biosynthesis, Signal Transduction, Action! Kluwer Academic Publisher: Dordrecht, The Netherlands, 2004. [Google Scholar]
- Li, J.; Li, C.; Smith, S.M. (Eds.) Hormone Metabolism and Signaling in Plants; Academic Press: London, UK, 2017. [Google Scholar]
- Benkő, P.; Gémes, K.; Fehér, A. Polyamine oxidase-generated reactive oxygen species in plant development and adaptation: The polyamine oxidase-NADPH oxidase nexus. Antioxidants 2022, 11, 2488. [Google Scholar] [CrossRef]
- Chen, D.; Shao, Q.; Yin, L.; Younis, A.; Zheng, B. Polyamine function in plants: Metabolism, regulation on development, and roles in abiotic stress responses. Front. Plant Sci. 2019, 9, 1945. [Google Scholar] [CrossRef]
- González-Hernández, A.I.; Scalschi, L.; Vicedo, B.; Marcos-Barbero, E.L.; Morcuende, R.; Camañes, G. Putrescine: A key metabolite involved in plant development, tolerance and resistance responses to stress. Int. J. Mol. Sci. 2022, 23, 2971. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Alhaithloul, H.A.S.; Parvin, K.; Bhuyan, M.H.M.B.; Tanveer, M.; Mohsin, S.M.; Nahar, K.; Soliman, M.H.; Mahmud, J.A.; Fujita, M. Polyamine action under metal/metalloid stress: Regulation of biosynthesis, metabolism, and molecular interactions. Int. J. Mol. Sci. 2019, 20, 3215. [Google Scholar] [CrossRef]
- Docimo, T.; Reichelt, M.; Schneider, B.; Kai, M.; Kunert, G.; Gershenzon, J.; D’Auria, J.C. The first step in the biosynthesis of cocaine in Erythroxylum coca: The characterization of arginine and ornithine decarboxylases. Plant Mol. Biol. 2012, 78, 599–615. [Google Scholar] [CrossRef]
- Perez-Amador, M.A.; Leon, J.; Green, P.J.; Carbonell, J. Induction of the arginine decarboxylase ADC2 gene provides evidence for the involvement of polyamines in the wound response in Arabidopsis. Plant Physiol. 2002, 130, 1454–1463. [Google Scholar] [CrossRef] [PubMed]
- Patel, J.; Ariyaratne, M.; Ahmed, S.; Ge, L.; Phuntumart, V.; Kalinoski, A.; Morris, P.F. Dual functioning of plant arginases provides a third route for putrescine synthesis. Plant Sci. 2017, 262, 62–73. [Google Scholar] [CrossRef] [PubMed]
- Dewey, R.E.; Xie, J. Molecular genetics of alkaloid biosynthesis in Nicotiana tabacum. Phytochemistry 2013, 94, 10–27. [Google Scholar] [CrossRef] [PubMed]
- Hanfrey, C.; Sommer, S.; Mayer, M.J.; Burtin, D.; Michael, A.J. Arabidopsis polyamine biosynthesis: Absence of ornithine decarboxylase and the mechanism of arginine decarboxylase activity. Plant J. 2001, 27, 551–560. [Google Scholar] [CrossRef]
- Ebeed, H.T. Genome-wide analysis of polyamine biosynthesis genes in wheat reveals gene expression specificity and involvement of STRE and MYB-elements in regulating polyamines under drought. BMC Genom. 2022, 23, 734. [Google Scholar] [CrossRef] [PubMed]
- Fuell, C.; Elliott, K.A.; Hanfrey, C.C.; Franceschetti, M.; Michael, A.J. Polyamine biosynthetic diversity in plants and algae. Plant Physiol. Biochem. 2010, 48, 513–520. [Google Scholar] [CrossRef]
- Ramazan, S.; Nazir, I.; Yousuf, W.; John, R. Environmental stress tolerance in maize (Zea mays): Role of polyamine metabolism. Funct. Plant Biol. 2023, 50, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Wang, X.; Zhao, F.; Zhang, X.; Li, W.; Huang, J.; Pei, X.; Ren, X.; Liu, Y.; He, K.; et al. Roles of S-adenosylmethionine and its derivatives in salt tolerance of cotton. Int. J. Mol. Sci. 2023, 24, 9517. [Google Scholar] [CrossRef]
- Basu, S.; Roychoudhury, A.; Sengupta, D.N. Identification of trans-acting factors regulating SamDC expression in Oryza sativa. Biochem. Biophys. Res. Comm. 2014, 445, 398–403. [Google Scholar] [CrossRef]
- Ge, C.; Cui, X.; Wang, Y.; Hu, Y.; Fu, Z.; Zhang, D.; Cheng, Z.; Li, J. BUD2, encoding an S-adenosylmethionine decarboxylase, is required for Arabidopsis growth and development. Cell Res. 2006, 16, 446–456. [Google Scholar] [CrossRef]
- Capell, T.; Bassie, L.; Christou, P. Modulation of the polyamine biosynthetic pathway in transgenic rice confers tolerance to drought stress. Proc. Natl. Acad. Sci. USA 2004, 101, 9909–9914. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira, L.F.; Elbl, P.; Navarro, B.V.; Macedo, A.F.; dos Santos, A.L.W.; Floh, E.I.S.; Cooke, J. Elucidation of the polyamine biosynthesis pathway during Brazilian pine (Araucaria angustifolia) seed development. Tree Physiol. 2017, 37, 116–130. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, R.K.; Fatima, T.; Handa, A.K.; Mattoo, A.K. Polyamines and their biosynthesis/catabolism genes are differentially modulated in response to heat versus cold stress in tomato leaves (Solanum lycopersicum L.). Cells 2020, 9, 1749. [Google Scholar] [CrossRef] [PubMed]
- Lasanajak, Y.; Minocha, R.; Minocha, S.C.; Goyal, R.; Fatima, T.; Handa, A.K.; Mattoo, A.K. Enhanced flux of substrates into polyamine biosynthesis but not ethylene in tomato fruit engineered with yeast S-adenosylmethionine decarboxylase gene. Amino Acids 2013, 46, 729–742. [Google Scholar] [CrossRef]
- Takács, Z.; Poór, P.; Tari, I. Interaction between polyamines and ethylene in the response to salicylic acid under normal photoperiod and prolonged darkness. Plant Physiol. Biochem. 2021, 167, 470–480. [Google Scholar] [CrossRef]
- Pattyn, J.; Vaughan-Hirsch, J.; Van de Poel, B. The regulation of ethylene biosynthesis: A complex multilevel control circuitry. New Phytol. 2021, 229, 770–782. [Google Scholar] [CrossRef]
- Wang, K.L.C.; Li, H.; Ecker, J.R. Ethylene biosynthesis and signaling networks. Plant Cell 2002, 14, S131–S151. [Google Scholar] [CrossRef]
- Hayashi, T.; Teruya, T.; Chaleckis, R.; Morigasaki, S.; Yanagida, M. S-adenosylmethionine synthetase is required for cell growth, maintenance of G0 phase, and termination of quiescence in fission yeast. iScience 2018, 5, 38–51. [Google Scholar] [CrossRef]
- Houben, M.; Van de Poel, B. 1-aminocyclopropane-1-carboxylic acid oxidase (ACO): The enzyme that makes the plant hormone ethylene. Front. Plant Sci. 2019, 10, 695. [Google Scholar] [CrossRef]
- Lieberman, M.; Kunishi, A. Stimulation of ethylene production in apple tissue slices by methionine. Plant Physiol. 1966, 41, 376–382. [Google Scholar] [CrossRef]
- Boller, T.; Herner, R.C.; Kende, H. Assay for and enzymatic formation of an ethylene precursor, 1-aminocyclopropane-1-carboxylic acid. Planta 1979, 145, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.Y.; Chen, Y.C.; Kieber, J.J.; Yoon, G.M. Regulation of the turnover of ACC synthases by phytohormones and heterodimerization in Arabidopsis. Plant J. 2017, 91, 491–504. [Google Scholar] [CrossRef] [PubMed]
- Yamagami, T.; Tsuchisaka, A.; Yamada, K.; Haddon, W.F.; Harden, L.A.; Theologis, A. Biochemical diversity among the 1-amino-cyclopropane-1-carboxylate synthase isozymes encoded by the Arabidopsis gene family. J. Biol. Chem. 2003, 278, 49102–49112. [Google Scholar] [CrossRef] [PubMed]
- Barry, C.S.; Llop-Tous, M.I.; Grierson, D. The regulation of 1-aminocyclopropane-1-carboxylic acid synthase gene expression during the transition from system-1 to system-2 ethylene synthesis in tomato. Plant Physiol. 2000, 123, 979–986. [Google Scholar] [CrossRef]
- Polko, J.K.; Kieber, J.J. 1-aminocyclopropane 1-carboxylic acid and its emerging role as an ethylene-independent growth regulator. Front. Plant Sci. 2019, 10, 1602. [Google Scholar] [CrossRef]
- Hatzfeld, Y.; Maruyama, A.; Schmidt, A.; Noji, M.; Ishizawa, K.; Saito, K. b-cyanoalanine synthase is a mitochondrial cysteine synthase-like protein in spinach and Arabidopsis. Plant Physiol. 2000, 123, 1163–1171. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Han, X.; Feng, D.; Yuan, D.; Huang, L.-J. Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: Do we understand what they are whispering? Int. J. Mol. Sci. 2019, 20, 671. [Google Scholar] [CrossRef]
- Staswick, P.E.; Tiryaki, I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell 2004, 16, 2117–2127. [Google Scholar] [CrossRef]
- Mishra, A.K.; Baek, K.-H. Salicylic acid biosynthesis and metabolism: A divergent pathway for plants and bacteria. Biomolecules 2021, 11, 705. [Google Scholar] [CrossRef]
- Arif, Y.; Sami, F.; Siddiqui, H.; Bajguz, A.; Hayat, S. Salicylic acid in relation to other phytohormones in plant: A study towards physiology and signal transduction under challenging environment. Environ. Exp. Bot. 2020, 175, 104040. [Google Scholar] [CrossRef]
- Larkindale, J.; Knight, M.R. Protection against heat stress-induced oxidative damage in Arabidopsis involves calcium, abscisic acid, ethylene, and salicylic acid. Plant Physiol. 2002, 128, 682–695. [Google Scholar] [CrossRef] [PubMed]
- Rekhter, D.; Lüdke, D.; Ding, Y.; Feussner, K.; Zienkiewicz, K.; Lipka, V.; Wiermer, M.; Zhang, Y.; Feussner, I. Isochorismate-derived biosynthesis of the plant stress hormone salicylic acid. Science 2019, 365, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Averesch, N.J.H.; Krömer, J.O. Metabolic engineering of the shikimate pathway for production of aromatics and derived compounds-present and future strain construction strategies. Front. Bioeng. Biotechnol. 2018, 6, 32. [Google Scholar] [CrossRef] [PubMed]
- Yokoo, S.; Inoue, S.; Suzuki, N.; Amakawa, N.; Matsui, H.; Nakagami, H.; Takahashi, A.; Arai, R.; Katou, S. Comparative analysis of plant isochorismate synthases reveals structural mechanisms underlying their distinct biochemical properties. Biosci. Rep. 2018, 38, BSR20171457. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, M.; Zeier, J. N-hydroxypipecolic acid and salicylic acid: A metabolic duo for systemic acquired resistance. Curr. Opin. Plant Biol. 2019, 50, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Torrens-Spence, M.P.; Bobokalonova, A.; Carballo, V.; Glinkerman, C.M.; Pluskal, T.; Shen, A.; Weng, J.-K. PBS3 and EPS1 complete salicylic acid biosynthesis from isochorismate in Arabidopsis. Mol. Plant 2019, 12, 1577–1586. [Google Scholar] [CrossRef]
- Chen, Z.; Zheng, Z.; Huang, J.; Lai, Z.; Fan, B. Biosynthesis of salicylic acid in plants. Plant Signal. Behav. 2009, 4, 493–496. [Google Scholar] [CrossRef]
- Dixon, R.A.; Paiva, N.L. Stress-induced phenylpropanoid metabolism. Plant Cell 1995, 7, 1085–1097. [Google Scholar] [CrossRef]
- Lefevere, H.; Bauters, L.; Gheysen, G. Salicylic acid biosynthesis in plants. Front. Plant Sci. 2020, 11, 338. [Google Scholar] [CrossRef]
- Reichert, A.I.; He, X.-Z.; Dixon, R.A. Phenylalanine ammonia-lyase (PAL) from tobacco (Nicotiana tabacum): Characterization of the four tobacco PAL genes and active heterotetrameric enzymes. Biochem. J. 2009, 424, 233–242. [Google Scholar] [CrossRef]
- Klämbt, H.D. Conversion in plants of benzoic acid to salicylic acid and its βd-glucoside. Nature 1962, 196, 491. [Google Scholar] [CrossRef]
- el-Basyouni, S.Z.; Chen, D.; Ibrahim, R.K.; Neish, A.C.; Towers, G.H.N. The biosynthesis of hydroxybenzoic acids in higher plants. Phytochemistry 1964, 3, 485–492. [Google Scholar] [CrossRef]
- Korver, R.A.; Koevoets, I.T.; Testerink, C. Out of shape during stress: A key role for auxin. Trends Plant Sci. 2018, 23, 783–793. [Google Scholar] [CrossRef]
- Song, X.; Xiong, Y.; Kong, X.; Huang, G. Roles of auxin response factors in rice development and stress responses. Plant Cell Environ. 2022, 46, 1075–1086. [Google Scholar] [CrossRef]
- Ljung, K.; Hull, A.K.; Celenza, J.; Yamada, M.; Estelle, M.; Normanly, J.; Sandberg, G. Sites and regulation of auxin biosynthesis in Arabidopsis roots. Plant Cell 2005, 17, 1090–1104. [Google Scholar] [CrossRef]
- Ljung, K.; Hull, A.K.; Kowalczyk, M.; Marchant, A.; Celenza, J.; Cohen, J.D.; Sandberg, G. Biosynthesis, conjugation, catabolism and homeostasis of indole-3-acetic acid in Arabidopsis thaliana. In Auxin Molecular Biology; Springer: Dordrecht, The Netherlands, 2002; pp. 249–272. [Google Scholar] [CrossRef]
- Stepanova, A.N.; Robertson-Hoyt, J.; Yun, J.; Benavente, L.M.; Xie, D.-Y.; Doležal, K.; Schlereth, A.; Jürgens, G.; Alonso, J.M. TAA1-mediated auxin biosynthesis is essential for hormone crosstalk and plant development. Cell 2008, 133, 177–191. [Google Scholar] [CrossRef] [PubMed]
- Mano, Y.; Nemoto, K.; Suzuki, M.; Seki, H.; Fujii, I.; Muranaka, T. The AMI1 gene family: Indole-3-acetamide hydrolase functions in auxin biosynthesis in plants. J. Exp. Bot. 2010, 61, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y. Auxin biosynthesis and its role in plant development. Annu. Rev. Plant Biol. 2010, 61, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Normanly, J. Approaching cellular and molecular resolution of auxin biosynthesis and metabolism. Cold Spring Harb. Perspect. Biol. 2010, 2, a001594. [Google Scholar] [CrossRef]
- Luo, P.; Di, D.-W. Precise regulation of the TAA1/TAR-YUCCA auxin biosynthesis pathway in plants. Int. J. Mol. Sci. 2023, 24, 8514. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin biosynthesis: A simple two-step pathway converts tryptophan to indole-3-acetic acid in plants. Mol. Plant 2012, 5, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Mano, Y.; Nemoto, K. The pathway of auxin biosynthesis in plants. J. Exp. Bot. 2012, 63, 2853–2872. [Google Scholar] [CrossRef]
- Galili, G.; Amir, R.; Fernie, A.R. The regulation of essential amino acid synthesis and accumulation in plants. Annu. Rev. Plant Biol. 2016, 67, 153–178. [Google Scholar] [CrossRef] [PubMed]
- Radwanski, E.R.; Last, R.L. Tryptophan biosynthesis and metabolism: Biochemical and molecular genetics. Plant Cell 1995, 7, 921–934. [Google Scholar] [CrossRef] [PubMed]
- Pollmann, S.; Neu, D.; Weiler, E.W. Molecular cloning and characterization of an amidase from Arabidopsis thaliana capable of converting indole-3-acetamide into the plant growth hormone, indole-3-acetic acid. Phytochemistry 2003, 62, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Pollmann, S.; Düchting, P.; Weiler, E.W. Tryptophan-dependent indole-3-acetic acid biosynthesis by ‘IAA-synthase’ proceeds via indole-3-acetamide. Phytochemistry 2009, 70, 523–531. [Google Scholar] [CrossRef]
- Romano, C.P.; Robson, P.R.H.; Smith, H.; Estelle, M.; Klee, H. Transgene-mediated auxin overproduction in Arabidopsis: Hypocotyl elongation phenotype and interactions with the hy6-1 hypocotyl elongation and axr1 auxin-resistant mutants. Plant Mol. Biol. 1995, 27, 1071–1083. [Google Scholar] [CrossRef]
- Tao, Y.; Ferrer, J.-L.; Ljung, K.; Pojer, F.; Hong, F.; Long, J.A.; Li, L.; Moreno, J.E.; Bowman, M.E.; Ivans, L.J.; et al. Rapid synthesis of auxin via a new tryptophan-dependent pathway is required for shade avoidance in plants. Cell 2008, 133, 164–176. [Google Scholar] [CrossRef]
- Songstad, D.D.; De Luca, V.; Brisson, N.; Kurz, W.G.; Nessler, C.L. High levels of tryptamine accumulation in transgenic tobacco expressing tryptophan decarboxylase. Plant Physiol. 1990, 94, 1410–1413. [Google Scholar] [CrossRef]
- Kang, S.; Kang, K.; Lee, K.; Back, K. Characterization of rice tryptophan decarboxylases and their direct involvement in serotonin biosynthesis in transgenic rice. Planta 2007, 227, 263–272. [Google Scholar] [CrossRef]
- Fujiwara, T.; Maisonneuve, S.; Isshiki, M.; Mizutani, M.; Chen, L.; Wong, H.L.; Kawasaki, T.; Shimamoto, K. Sekiguchi lesion gene encodes a cytochrome P450 monooxygenase that catalyzes conversion of tryptamine to serotonin in rice. J. Biol. Chem. 2010, 285, 11308–11313. [Google Scholar] [CrossRef]
- Zhang, D.; Song, Y.H.; Dai, R.; Lee, T.G.; Kim, J. Aldoxime metabolism is linked to phenylpropanoid production in Camelina sativa. Front. Plant Sci. 2020, 11, 17. [Google Scholar] [CrossRef]
- Kim, J.I.; Zhang, X.; Pascuzzi, P.E.; Liu, C.J.; Chapple, C. Glucosinolate and phenylpropanoid biosynthesis are linked by proteasome-dependent degradation of PAL. New Phytol. 2019, 225, 154–168. [Google Scholar] [CrossRef] [PubMed]
- Aoi, Y.; Tanaka, K.; Cook, S.D.; Hayashi, K.-I.; Kasahara, H. GH3 auxin-amido synthetases alter the ratio of indole-3-acetic acid and phenylacetic acid in Arabidopsis. Plant Cell Physiol. 2020, 61, 596–605. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin biosynthesis. In The Arabidopsis Book; American Society of Plant Biologists: Rockville, MD, USA, 2014; Volume 12, p. e0173. [Google Scholar] [CrossRef]
- Lee, H.Y.; Byeon, Y.; Back, K. Melatonin as a signal molecule triggering defense responses against pathogen attack in Arabidopsis and tobacco. J. Pineal Res. 2014, 57, 262–268. [Google Scholar] [CrossRef] [PubMed]
- Mannino, G.; Pernici, C.; Serio, G.; Gentile, C.; Bertea, C.M. Melatonin and phytomelatonin: Chemistry, biosynthesis, metabolism, distribution and bioactivity in plants and animals—An overview. Int. J. Mol. Sci. 2021, 22, 9996. [Google Scholar] [CrossRef]
- Kanwar, M.K.; Yu, J.Q.; Zhou, J. Phytomelatonin: Recent advances and future prospects. J. Pineal Res. 2018, 65, e12526. [Google Scholar] [CrossRef]
- Mir, A.R.; Faizan, M.; Bajguz, A.; Sami, F.; Siddiqui, H.; Hayat, S. Occurrence and biosynthesis of melatonin and its exogenous effect on plants. Acta Soc. Bot. Pol. 2020, 89, 8922. [Google Scholar] [CrossRef]
- Back, K. Melatonin metabolism, signaling and possible roles in plants. Plant J. 2021, 105, 376–391. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.F.; Hu, Q.; Zhang, X.; Jiang, J.F.; Zhang, Y.; Zhang, Z.X. Melatonin biosynthesis and signal transduction in plants in response to environmental conditions. J. Exp. Bot. 2022, 73, 5818–5827. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.X.; Reiter, R.J. An evolutionary view of melatonin synthesis and metabolism related to its biological functions in plants. J. Exp. Bot. 2020, 71, 4677–4689. [Google Scholar] [CrossRef] [PubMed]
- Byeon, Y.; Lee, H.Y.; Back, K. Chloroplastic and cytoplasmic overexpression of sheep serotonin N-acetyltransferase in transgenic rice plants is associated with low melatonin production despite high enzyme activity. J. Pineal Res. 2015, 58, 461–469. [Google Scholar] [CrossRef] [PubMed]
- Back, K.; Tan, D.-X.; Reiter, R.J. Melatonin biosynthesis in plants: Multiple pathways catalyze tryptophan to melatonin in the cytoplasm or chloroplasts. J. Pineal Res. 2016, 61, 426–437. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.; Lee, K.; Park, S.; Byeon, Y.; Back, K. Molecular cloning of rice serotonin N-acetyltransferase, the penultimate gene in plant melatonin biosynthesis. J. Pineal Res. 2013, 55, 7–13. [Google Scholar] [CrossRef]
- Park, S.; Byeon, Y.; Back, K. Functional analyses of three ASMT gene family members in rice plants. J. Pineal Res. 2013, 55, 409–415. [Google Scholar] [CrossRef]
- Zhao, D.; Yao, Z.; Zhang, J.; Zhang, R.; Mou, Z.; Zhang, X.; Li, Z.; Feng, X.; Chen, S.; Reiter, R.J. Melatonin synthesis genes N-acetylserotonin methyltransferases evolved into caffeic acid O-methyltransferases and both assisted in plant terrestrialization. J. Pineal Res. 2021, 71, e12737. [Google Scholar] [CrossRef]
- Xiong, L.; Zhu, J.-K. Regulation of abscisic acid biosynthesis. Plant Physiol. 2003, 133, 29–36. [Google Scholar] [CrossRef]
- Cardoso, A.A.; Gori, A.; Da-Silva, C.J.; Brunetti, C. Abscisic acid biosynthesis and signaling in plants: Key targets to improve water use efficiency and drought tolerance. Appl. Sci. 2020, 10, 6322. [Google Scholar] [CrossRef]
- Schwartz, S.H.; Qin, X.; Zeevaart, J.A.D. Elucidation of the indirect pathway of abscisic acid biosynthesis by mutants, genes, and enzymes. Plant Physiol. 2003, 131, 1591–1601. [Google Scholar] [CrossRef]
- Finkelstein, R. Abscisic acid synthesis and response. In The Arabidopsis Book; American Society of Plant Biologists: Rockville, MD, USA, 2013; Volume 11, p. e0166. [Google Scholar] [CrossRef]
- Seo, M.; Koshiba, T. Complex regulation of ABA biosynthesis in plants. Trends Plant Sci. 2002, 7, 41–48. [Google Scholar] [CrossRef]
- Nambara, E.; Suzuki, M.; Abrams, S.; McCarty, D.R.; Kamiya, Y.; McCourt, P. A screen for genes that function in abscisic acid signaling in Arabidopsis thaliana. Genetics 2002, 161, 1247–1255. [Google Scholar] [CrossRef]
- Dong, T.; Park, Y.; Hwang, I. Abscisic acid: Biosynthesis, inactivation, homoeostasis and signalling. Essays Biochem. 2015, 58, 29–48. [Google Scholar] [CrossRef]
- Nambara, E.; Marion-Poll, A. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 2005, 56, 165–185. [Google Scholar] [CrossRef]
- Laule, O.; Fürholz, A.; Chang, H.-S.; Zhu, T.; Wang, X.; Heifetz, P.B.; Gruissem, W.; Lange, M. Crosstalk between cytosolic and plastidial pathways of isoprenoid biosynthesis in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2003, 100, 6866–6871. [Google Scholar] [CrossRef]
- Frebort, I.; Kowalska, M.; Hluska, T.; Frebortova, J.; Galuszka, P. Evolution of cytokinin biosynthesis and degradation. J. Exp. Bot. 2011, 62, 2431–2452. [Google Scholar] [CrossRef] [PubMed]
- Sakakibara, H.; Kasahara, H.; Ueda, N.; Kojima, M.; Takei, K.; Hishiyama, S.; Asami, T.; Okada, K.; Kamiya, Y.; Yamaya, T.; et al. Agrobacterium tumefaciens increases cytokinin production in plastids by modifying the biosynthetic pathway in the host plant. Proc. Natl. Acad. Sci. USA 2005, 102, 9972–9977. [Google Scholar] [CrossRef] [PubMed]
- Milborrow, B.V. The pathway of biosynthesis of abscisic acid in vascular plants: A review of the present state of knowledge of ABA biosynthesis. J. Exp. Bot. 2001, 52, 1145–1164. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Hu, Q.; Yan, J.; Sun, K.; Liang, Y.; Jia, M.; Meng, X.; Fang, S.; Wang, Y.; Jing, Y.; et al. ζ-Carotene isomerase suppresses tillering in rice through the coordinated biosynthesis of strigolactone and abscisic acid. Mol. Plant 2020, 13, 1784–1801. [Google Scholar] [CrossRef]
- Bittner, F.; Oreb, M.; Mendel, R.R. ABA3 Is a molybdenum cofactor sulfurase required for activation of aldehyde oxidase and xanthine dehydrogenase in Arabidopsis thaliana. J. Biol. Chem. 2001, 276, 40381–40384. [Google Scholar] [CrossRef]
- Lee, H.S.; Milborrow, B.V. Endogenous biosynthetic precursors of (+)-abscisic acid. V. Inhibition by tungstate and its removal by cinchonine shows that xanthoxal is oxidised by a molybdo-aldehyde oxidase. Funct. Plant Biol. 1997, 24, 727. [Google Scholar] [CrossRef][Green Version]
- Rock, C.D.; Heath, T.G.; Gage, D.A.; Zeevaart, J.A. Abscisic alcohol is an intermediate in abscisic acid biosynthesis in a shunt pathway from abscisic aldehyde. Plant Physiol. 1991, 97, 670–676. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ahanger, M.A.; Ashraf, M.; Bajguz, A.; Ahmad, P. Brassinosteroids regulate growth in plants under stressful environments and crosstalk with other potential phytohormones. J. Plant Growth Regul. 2018, 37, 1007–1024. [Google Scholar] [CrossRef]
- Bajguz, A.; Hayat, S. Effects of brassinosteroids on the plant responses to environmental stresses. Plant Physiol. Biochem. 2009, 47, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Hayat, S.; Mori, M.; Fariduddin, Q.; Bajguz, A.; Ahmad, A. Physiological role of brassinosteroids: An update. Indian J. Plant Physiol. 2010, 15, 99–109. [Google Scholar]
- Rajewska, I.; Talarek, M.; Bajguz, A. Brassinosteroids and response of plants to heavy metals action. Front. Plant Sci. 2016, 7, 629. [Google Scholar] [CrossRef]
- Wei, Z.; Li, J. Regulation of brassinosteroid homeostasis in higher plants. Front. Plant Sci. 2020, 11, 583622. [Google Scholar] [CrossRef]
- Wei, Z.Y.; Li, J. Brassinosteroids regulate root growth, development, and symbiosis. Mol. Plant 2016, 9, 86–100. [Google Scholar] [CrossRef]
- Yao, T.; Xie, R.; Zhou, C.; Wu, X.; Li, D. Roles of brassinosteroids signaling in biotic and abiotic stresses. J. Agr. Food Chem. 2023, 71, 7947–7960. [Google Scholar] [CrossRef]
- Bajguz, A. Metabolism of brassinosteroids in plants. Plant Physiol. Biochem. 2007, 45, 95–107. [Google Scholar] [CrossRef]
- Bajguz, A.; Chmur, M.; Gruszka, D. Comprehensive overview of the brassinosteroid biosynthesis pathways: Substrates, products, inhibitors, and connections. Front. Plant Sci. 2020, 11, 1034. [Google Scholar] [CrossRef]
- Manghwar, H.; Hussain, A.; Ali, Q.; Liu, F. Brassinosteroids (BRs) role in plant development and coping with different stresses. Int. J. Mol. Sci. 2022, 23, 1012. [Google Scholar] [CrossRef] [PubMed]
- Zullo, M.A.T.; Bajguz, A. The brassinosteroids family—Structural diversity of natural compounds and their precursors. In Brassinosteroids: Plant Growth and Development; Hayat, S., Yusuf, M., Bhardwaj, R., Bajguz, A., Eds.; Springer: Singapore, 2019; pp. 1–44. [Google Scholar] [CrossRef]
- Bajguz, A.; Tretyn, A. The chemical characteristic and distribution of brassinosteroids in plants. Phytochemistry 2003, 62, 1027–1046. [Google Scholar] [CrossRef] [PubMed]
- Roh, J.; Moon, J.; Youn, J.H.; Seo, C.; Park, Y.J.; Kim, S.K. Establishment of biosynthetic pathways to generate castasterone as the biologically active brassinosteroid in Brachypodium distachyon. J. Agr. Food Chem. 2020, 68, 3912–3923. [Google Scholar] [CrossRef]
- Rozhon, W.; Akter, S.; Fernandez, A.; Poppenberger, B. Inhibitors of brassinosteroid biosynthesis and signal transduction. Molecules 2019, 24, 4372. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Wei, Z.; Li, J.; Wang, X. Brassinosteroids. In Hormone Metabolism and Signaling in Plants; Li, J., Li, C., Smith, S.M., Eds.; Academic Press: London, UK, 2017; pp. 291–326. [Google Scholar] [CrossRef]
- Ohnishi, T.; Szatmari, A.-M.; Watanabe, B.; Fujita, S.; Bancos, S.; Koncz, C.; Lafos, M.; Shibata, K.; Yokota, T.; Sakata, K.; et al. C-23 hydroxylation by Arabidopsis CYP90C1 and CYP90D1 reveals a novel shortcut in brassinosteroid biosynthesis. Plant Cell 2006, 18, 3275–3288. [Google Scholar] [CrossRef] [PubMed]
- Joo, S.-H.; Jang, M.-S.; Kim, M.K.; Lee, J.-E.; Kim, S.-K. Biosynthetic relationship between C₂₈-brassinosteroids and C₂₉-brassinosteroids in rice (Oryza sativa) seedlings. Phytochemistry 2015, 111, 84–90. [Google Scholar] [CrossRef]
- Kim, S.; Moon, J.; Roh, J.; Kim, S.-K. Castasterone can be biosynthesized from 28-homodolichosterone in Arabidopsis thaliana. J. Plant Biol. 2018, 61, 330–335. [Google Scholar] [CrossRef]
- Lee, S.-C.; Hwang, J.-Y.; Joo, S.-H.; Son, S.-H.; Youn, J.-H.; Kim, S.-K. Biosynthesis and metabolism of dolichosterone in Arabidopsis thaliana. Bull. Korean Chem. Soc. 2010, 31, 3475–3478. [Google Scholar] [CrossRef]
- Lee, S.-C.; Joo, S.-H.; Son, S.-H.; Youn, J.-H.; Kim, S.-K. Metabolism of 28-homodolichosterone in Phaseolus vulgaris. Bull. Korean Chem. Soc. 2011, 32, 403–404. [Google Scholar] [CrossRef]
- Roh, J.; Yeom, H.S.; Jang, H.; Kim, S.; Youn, J.H.; Kim, S.-K. Identification and biosynthesis of C-24 ethylidene brassinosteroids in Arabidopsis thaliana. J. Plant Biol. 2017, 60, 533–538. [Google Scholar] [CrossRef]
- Sonawane, P.D.; Pollier, J.; Panda, S.; Szymanski, J.; Massalha, H.; Yona, M.; Unger, T.; Malitsky, S.; Arendt, P.; Pauwels, L.; et al. Plant cholesterol biosynthetic pathway overlaps with phytosterol metabolism. Nat. Plants 2016, 3, 16205. [Google Scholar] [CrossRef] [PubMed]
- Xin, P.; Yan, J.; Li, B.; Fang, S.; Fan, J.; Tian, H.; Shi, Y.; Tian, W.; Yan, C.; Chu, J. A comprehensive and effective mass spectrometry-based screening strategy for discovery and identification of new brassinosteroids from rice tissues. Front. Plant Sci. 2016, 7, 1786. [Google Scholar] [CrossRef] [PubMed]
- Fujiyama, K.; Hino, T.; Kanadani, M.; Watanabe, B.; Lee, H.J.; Mizutani, M.; Nagano, S. Structural insights into a key step of brassinosteroid biosynthesis and its inhibition. Nat. Plants 2019, 5, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, T. Recent advances in brassinosteroid biosynthetic pathway: Insight into novel brassinosteroid shortcut pathway. J. Pestic. Sci. 2018, 43, 159–167. [Google Scholar] [CrossRef]
- Joo, S.-H.; Kim, T.-W.; Son, S.-H.; Lee, W.S.; Yokota, T.; Kim, S.-K. Biosynthesis of a cholesterol-derived brassinosteroid, 28-norcastasterone, in Arabidopsis thaliana. J. Exp. Bot. 2012, 63, 1823–1833. [Google Scholar] [CrossRef]
- Zhao, B.; Li, J. Regulation of brassinosteroid biosynthesis and inactivation. J. Integr. Plant Biol. 2012, 54, 746–759. [Google Scholar] [CrossRef]
- Chung, Y.; Choe, S. The regulation of brassinosteroid biosynthesis in Arabidopsis. Crit. Rev. Plant Sci. 2013, 32, 396–410. [Google Scholar] [CrossRef]
- Kakimoto, T. Biosynthesis of cytokinins. J. Plant Res. 2003, 116, 233–239. [Google Scholar] [CrossRef]
- Mishra, D.C.; Arora, D.; Budhlakoti, N.; Solanke, A.U.; Mithra, S.V.A.C.; Kumar, A.; Pandey, P.S.; Srivastava, S.; Kumar, S.; Farooqi, M.S.; et al. Identification of potential cytokinin responsive key genes in rice treated with trans-zeatin through systems biology approach. Front. Genet. 2022, 12, 780599. [Google Scholar] [CrossRef]
- Raspor, M.; Motyka, V.; Ninković, S.; Dobrev, P.I.; Malbeck, J.; Ćosić, T.; Cingel, A.; Savić, J.; Tadić, V.; Dragićević, I.Č. Endogenous levels of cytokinins, indole-3-acetic acid and abscisic acid in in vitro grown potato: A contribution to potato hormonomics. Sci. Rep. 2020, 10, 3437. [Google Scholar] [CrossRef]
- Žižková, E.; Kubeš, M.; Dobrev, P.I.; Přibyl, P.; Šimura, J.; Zahajská, L.; Záveská Drábková, L.; Novák, O.; Motyka, V. Control of cytokinin and auxin homeostasis in cyanobacteria and algae. Ann. Bot. 2017, 119, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Emery, R.J.; Ma, Q.; Atkins, C.A. The forms and sources of cytokinins in developing white lupine seeds and fruits. Plant Physiol. 2000, 123, 1593–1604. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Zhang, J.; Wang, Z.; Zhu, Q.; Wang, W. Hormonal changes in the grains of rice subjected to water stress during grain filling. Plant Physiol. 2001, 127, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Hrtyan, M.; Šliková, E.; Hejátko, J.; Růžička, K. RNA processing in auxin and cytokinin pathways. J. Exp. Bot. 2015, 66, 4897–4912. [Google Scholar] [CrossRef]
- Kakimoto, T. Identification of plant cytokinin biosynthetic enzymes as dimethylallyl diphosphate:ATP/ADP isopentenyltransferases. Plant Cell Physiol. 2001, 42, 677–685. [Google Scholar] [CrossRef]
- Astot, C.; Dolezal, K.; Nordström, A.; Wang, Q.; Kunkel, T.; Moritz, T.; Chua, N.H.; Sandberg, G. An alternative cytokinin biosynthesis pathway. Proc. Natl. Acad. Sci. USA 2000, 97, 14778–14783. [Google Scholar] [CrossRef]
- Krall, L.; Raschke, M.; Zenk, M.H.; Baron, C. The Tzs protein from Agrobacterium tumefaciens C58 produces zeatin riboside 5′-phosphate from 4-hydroxy-3-methyl-2-(E)-butenyl diphosphate and AMP. FEBS Lett. 2002, 527, 315–318. [Google Scholar] [CrossRef]
- Takei, K.; Ueda, N.; Aoki, K.; Kuromori, T.; Hirayama, T.; Shinozaki, K.; Yamaya, T.; Sakakibara, H. AtIPT3 is a key determinant of nitrate-dependent cytokinin biosynthesis in Arabidopsis. Plant Cell Physiol. 2004, 45, 1053–1062. [Google Scholar] [CrossRef]
- Kasahara, H.; Takei, K.; Ueda, N.; Hishiyama, S.; Yamaya, T.; Kamiya, Y.; Yamaguchi, S.; Sakakibara, H. Distinct isoprenoid origins of cis- and trans-zeatin biosyntheses in Arabidopsis. J. Biol. Chem. 2004, 279, 14049–14054. [Google Scholar] [CrossRef]
- Abe, I.; Tanaka, H.; Abe, T.; Noguchi, H. Enzymatic formation of unnatural cytokinin analogs by adenylate isopentenyltransferase from mulberry. Biochem. Biophys. Res. Comm. 2007, 355, 795–800. [Google Scholar] [CrossRef]
- Sakano, Y.; Okada, Y.; Matsunaga, A.; Suwama, T.; Kaneko, T.; Ito, K.; Noguchi, H.; Abe, I. Molecular cloning, expression, and characterization of adenylate isopentenyltransferase from hop (Humulus lupulus L.). Phytochemistry 2004, 65, 2439–2446. [Google Scholar] [CrossRef]
- Hwang, I.; Sakakibara, H. Cytokinin biosynthesis and perception. Physiol. Plant. 2006, 126, 528–538. [Google Scholar] [CrossRef]
- Chu, H.-M.; Ko, T.-P.; Wang, A.H.J. Crystal structure and substrate specificity of plant adenylate isopentenyltransferase from Humulus lupulus: Distinctive binding affinity for purine and pyrimidine nucleotides. Nucleic Acids Res. 2010, 38, 1738–1748. [Google Scholar] [CrossRef][Green Version]
- Zhou, C.; Huang, R.H. Crystallographic snapshots of eukaryotic dimethylallyltransferase acting on tRNA: Insight into tRNA recognition and reaction mechanism. Proc. Natl. Acad. Sci. USA 2008, 105, 16142–16147. [Google Scholar] [CrossRef]
- Pimenta Lange, M.J.; Lange, T. Gibberellin biosynthesis and the regulation of plant development. Plant Biol. 2006, 8, 281–290. [Google Scholar] [CrossRef]
- Gupta, R.; Chakrabarty, S.K. Gibberellic acid in plant: Still a mystery unresolved. Plant Signal. Behav. 2013, 8, e25504. [Google Scholar] [CrossRef] [PubMed]
- Sponsel, V.M.; Hedden, P. Gibberellin biosynthesis and inactivation. In Plant Hormones: Biosynthesis, Signal transduction, Action! Davies, P.J., Ed.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2004; pp. 63–94. [Google Scholar] [CrossRef]
- Takahashi, N.; Kitamura, H.; Kawarada, A.; Seta, Y.; Takai, M.; Tamura, S.; Sumiki, Y. Biochemical studies on “Bakanae” fungus. Bull. Agricult. Chem. Soc. Jpn. 1955, 19, 267–281. [Google Scholar] [CrossRef]
- Hedden, P. The current status of research on gibberellin biosynthesis. Plant Cell Physiol. 2020, 61, 1832–1849. [Google Scholar] [CrossRef] [PubMed]
- Hedden, P.; Sponsel, V. A century of gibberellin research. J. Plant Growth Regul. 2015, 34, 740–760. [Google Scholar] [CrossRef] [PubMed]
- Salazar-Cerezo, S.; Martínez-Montiel, N.; García-Sánchez, J.; Pérez-y-Terrón, R.; Martínez-Contreras, R.D. Gibberellin biosynthesis and metabolism: A convergent route for plants, fungi and bacteria. Microbiol. Res. 2018, 208, 85–98. [Google Scholar] [CrossRef]
- Flügge, U.I.; Gao, W. Transport of isoprenoid intermediates across chloroplast envelope membranes. Plant Biol. 2005, 7, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Bräutigam, A.; Weber, A.P.M. Proteomic analysis of the proplastid envelope membrane provides novel insights into small molecule and protein transport across proplastid membranes. Mol. Plant 2009, 2, 1247–1261. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Sola, M.Á.; Coman, D.; Beck, G.; Barja, M.V.; Colinas, M.; Graf, A.; Welsch, R.; Rütimann, P.; Bühlmann, P.; Bigler, L.; et al. Arabidopsis GERANYLGERANYL DIPHOSPHATE SYNTHASE11 is a hub isozyme required for the production of most photosynthesis-related isoprenoids. New Phytol. 2015, 209, 252–264. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Wang, C.-Y.; Gutensohn, M.; Jiang, L.; Zhang, P.; Zhang, D.; Dudareva, N.; Lu, S. A recruiting protein of geranylgeranyl diphosphate synthase controls metabolic flux toward chlorophyll biosynthesis in rice. Proc. Natl. Acad. Sci. USA 2017, 114, 6866–6871. [Google Scholar] [CrossRef]
- Lemke, C.; Potter, K.C.; Schulte, S.; Peters, R.J. Conserved bases for the initial cyclase in gibberellin biosynthesis: From bacteria to plants. Biochem. J. 2019, 476, 2607–2621. [Google Scholar] [CrossRef]
- Prisic, S.; Peters, R.J. Synergistic substrate inhibition of ent-copalyl diphosphate synthase: A potential feed-forward inhibition mechanism limiting gibberellin metabolism. Plant Physiol. 2007, 144, 445–454. [Google Scholar] [CrossRef]
- Zi, J.; Mafu, S.; Peters, R.J. To gibberellins and beyond! Surveying the evolution of (di)terpenoid metabolism. Annu. Rev. Plant Biol. 2014, 65, 259–286. [Google Scholar] [CrossRef]
- Liu, W.; Feng, X.; Zheng, Y.; Huang, C.-H.; Nakano, C.; Hoshino, T.; Bogue, S.; Ko, T.-P.; Chen, C.-C.; Cui, Y.; et al. Structure, function and inhibition of ent-kaurene synthase from Bradyrhizobium japonicum. Sci. Rep. 2014, 4, 6214. [Google Scholar] [CrossRef]
- Shimane, M.; Ueno, Y.; Morisaki, K.; Oogami, S.; Natsume, M.; Hayashi, K.-i.; Nozaki, H.; Kawaide, H. Molecular evolution of the substrate specificity of ent-kaurene synthases to adapt to gibberellin biosynthesis in land plants. Biochem. J. 2014, 462, 539–546. [Google Scholar] [CrossRef]
- Ding, Y.; Murphy, K.M.; Poretsky, E.; Mafu, S.; Yang, B.; Char, S.N.; Christensen, S.A.; Saldivar, E.; Wu, M.; Wang, Q.; et al. Multiple genes recruited from hormone pathways partition maize diterpenoid defences. Nat. Plants 2019, 5, 1043–1056. [Google Scholar] [CrossRef]
- Helliwell, C.A.; Chandler, P.M.; Poole, A.; Dennis, E.S.; Peacock, W.J. The CYP88A cytochrome P450, ent-kaurenoic acid oxidase, catalyzes three steps of the gibberellin biosynthesis pathway. Proc. Natl. Acad. Sci. USA 2001, 98, 2065–2070. [Google Scholar] [CrossRef] [PubMed]
- Nagel, R.; Turrini, P.C.G.; Nett, R.S.; Leach, J.E.; Verdier, V.; Van Sluys, M.-A.; Peters, R.J. An operon for production of bioactive gibberellin A4 phytohormone with wide distribution in the bacterial rice leaf streak pathogen Xanthomonas oryzae pv. oryzicola. New Phytol. 2017, 214, 1260–1266. [Google Scholar] [CrossRef]
- Morrone, D.; Chambers, J.; Lowry, L.; Kim, G.; Anterola, A.; Bender, K.; Peters, R.J. Gibberellin biosynthesis in bacteria: Separate ent-copalyl diphosphate and ent-kaurene synthases in Bradyrhizobium japonicum. FEBS Lett. 2009, 583, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Magome, H.; Nomura, T.; Hanada, A.; Takeda-Kamiya, N.; Ohnishi, T.; Shinma, Y.; Katsumata, T.; Kawaide, H.; Kamiya, Y.; Yamaguchi, S. CYP714B1 and CYP714B2 encode gibberellin 13-oxidases that reduce gibberellin activity in rice. Proc. Natl. Acad. Sci. USA 2013, 110, 1947–1952. [Google Scholar] [CrossRef]
- Nomura, T.; Magome, H.; Hanada, A.; Takeda-Kamiya, N.; Mander, L.N.; Kamiya, Y.; Yamaguchi, S. Functional analysis of Arabidopsis CYP714A1 and CYP714A2 reveals that they are distinct gibberellin modification enzymes. Plant Cell Physiol. 2013, 54, 1837–1851. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Chen, Q.; Xin, P.; Yuan, J.; Ma, Y.; Wang, X.; Xu, M.; Chu, J.; Peters, R.J.; Wang, G. CYP72A enzymes catalyse 13-hydrolyzation of gibberellins. Nat. Plants 2019, 5, 1057–1065. [Google Scholar] [CrossRef]
- Pimenta Lange, M.J.; Liebrandt, A.; Arnold, L.; Chmielewska, S.-M.; Felsberger, A.; Freier, E.; Heuer, M.; Zur, D.; Lange, T. Functional characterization of gibberellin oxidases from cucumber, Cucumis sativus L. Phytochemistry 2013, 90, 62–69. [Google Scholar] [CrossRef]
- Fleet, C.M.; Yamaguchi, S.; Hanada, A.; Kawaide, H.; David, C.J.; Kamiya, Y.; Sun, T.-P. Overexpression of AtCPS and AtKS in Arabidopsis confers increased ent-kaurene production but no increase in bioactive gibberellins. Plant Physiol. 2003, 132, 830–839. [Google Scholar] [CrossRef]
- Wang, S.; Wang, Y. Harnessing hormone gibberellin knowledge for plant height regulation. Plant Cell Rep. 2022, 41, 1945–1953. [Google Scholar] [CrossRef]
- Wu, F.; Gao, Y.; Yang, W.; Sui, N.; Zhu, J. Biological functions of strigolactones and their crosstalk with other phytohormones. Front. Plant Sci. 2022, 13, 821563. [Google Scholar] [CrossRef]
- Mashiguchi, K.; Seto, Y.; Yamaguchi, S. Strigolactone biosynthesis, transport and perception. Plant J. 2021, 105, 335–350. [Google Scholar] [CrossRef]
- Soliman, S.; Wang, Y.; Han, Z.H.; Pervaiz, T.; El-kereamy, A. Strigolactones in plants and their interaction with the ecological microbiome in response to abiotic stress. Plants 2022, 11, 3499. [Google Scholar] [CrossRef]
- Xie, X.N.; Mori, N.; Yoneyama, K.; Nomura, T.; Uchida, K.; Yoneyama, K.; Akiyama, K. Lotuslactone, a non-canonical strigolactone from Lotus japonicus. Phytochemistry 2019, 157, 200–205. [Google Scholar] [CrossRef]
- Ruiz-Lozano, J.M.; Aroca, R.; Zamarreno, A.M.; Molina, S.; Andreo-Jimenez, B.; Porcel, R.; Garcia-Mina, J.M.; Ruyter-Spira, C.; Lopez-Raez, J.A. Arbuscular mycorrhizal symbiosis induces strigolactone biosynthesis under drought and improves drought tolerance in lettuce and tomato. Plant Cell Environ. 2016, 39, 441–452. [Google Scholar] [CrossRef]
- Matusova, R.; Rani, K.; Verstappen, F.W.A.; Franssen, M.C.R.; Beale, M.H.; Bouwmeester, H.J. The strigolactone germination stimulants of the plant-parasitic Striga and Orobanche spp. are derived from the carotenoid pathway. Plant Physiol. 2005, 139, 920–934. [Google Scholar] [CrossRef] [PubMed]
- Alder, A.; Jamil, M.; Marzorati, M.; Bruno, M.; Vermathen, M.; Bigler, P.; Ghisla, S.; Bouwmeester, H.; Beyer, P.; Al-Babili, S. The path from β-carotene to carlactone, a strigolactone-like plant hormone. Science 2012, 335, 1348–1351. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.-T.E.; Wang, J.Y.; Jamil, M.; Braguy, J.; Al-Babili, S. 9-cis-β-Apo-10ʹ-carotenal is the precursor of strigolactones in planta. Planta 2022, 256, 88. [Google Scholar] [CrossRef] [PubMed]
- Yoda, A.; Xie, X.; Yoneyama, K.; Miura, K.; McErlean, C.S.P.; Nomura, T. A stereoselective strigolactone biosynthesis catalyzed by a 2-oxoglutarate-dependent dioxygenase in Sorghum. Plant Cell Physiol. 2023; in press. [Google Scholar] [CrossRef]
- Waters, M.T.; Gutjahr, C.; Bennett, T.; Nelson, D.C. Strigolactone signaling and evolution. Annu. Rev. Plant Biol. 2017, 68, 291–322. [Google Scholar] [CrossRef]
- Abe, S.; Sado, A.; Tanaka, K.; Kisugi, T.; Asami, K.; Ota, S.; Kim, H.I.; Yoneyama, K.; Xie, X.; Ohnishi, T.; et al. Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro. Proc. Natl. Acad. Sci. USA 2014, 111, 18084–18089. [Google Scholar] [CrossRef]
- Seto, Y.; Sado, A.; Asami, K.; Hanada, A.; Umehara, M.; Akiyama, K.; Yamaguchi, S. Carlactone is an endogenous biosynthetic precursor for strigolactones. Proc. Natl. Acad. Sci. USA 2014, 111, 1640–1645. [Google Scholar] [CrossRef]
- Baz, L.; Mori, N.; Mi, J.N.; Jamil, M.; Kountche, B.A.; Guo, X.J.; Balakrishna, A.; Jia, K.P.; Vermathen, M.; Akiyama, K.; et al. 3-Hydroxycarlactone, a novel product of the strigolactone biosynthesis core pathway. Mol. Plant 2018, 11, 1312–1314. [Google Scholar] [CrossRef] [PubMed]
- Vinde, M.H.; Cao, D.; Chesterfield, R.J.; Yoneyama, K.; Gumulya, Y.; Thomson, R.E.S.; Matila, T.; Ebert, B.E.; Beveridge, C.A.; Vickers, C.E.; et al. Ancestral sequence reconstruction of the CYP711 family reveals functional divergence in strigolactone biosynthetic enzymes associated with gene duplication events in monocot grasses. New Phytol. 2022, 235, 1900–1912. [Google Scholar] [CrossRef]
- Wakabayashi, T.; Moriyama, D.; Miyamoto, A.; Okamura, H.; Shiotani, N.; Shimizu, N.; Mizutani, M.; Takikawa, H.; Sugimoto, Y. Identification of novel canonical strigolactones produced by tomato. Front. Plant Sci. 2022, 13, 1064378. [Google Scholar] [CrossRef]
- Wakabayashi, T.; Shinde, H.; Shiotani, N.; Yamamoto, S.; Mizutani, M.; Takikawa, H.; Sugimoto, Y. Conversion of methyl carlactonoate to heliolactone in sunflower. Nat. Prod. Res. 2022, 36, 2215–2222. [Google Scholar] [CrossRef]
- Yoneyama, K.; Akiyama, K.; Brewer, P.B.; Mori, N.; Kawano-Kawada, M.; Haruta, S.; Nishiwaki, H.; Yamauchi, S.; Xie, X.; Umehara, M.; et al. Hydroxyl carlactone derivatives are predominant strigolactones in Arabidopsis. Plant Direct 2020, 4, e00219. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, K.; Brewer, P.B. Strigolactones, how are they synthesized to regulate plant growth and development? Curr. Opin. Plant Biol. 2021, 63, 102072. [Google Scholar] [CrossRef] [PubMed]
- Yoneyama, K. Recent progress in the chemistry and biochemistry of strigolactones. J. Pestic. Sci. 2020, 45, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Katsir, L.; Chung, H.S.; Koo, A.J.K.; Howe, G.A. Jasmonate signaling: A conserved mechanism of hormone sensing. Curr. Opin. Plant Biol. 2008, 11, 428–435. [Google Scholar] [CrossRef]
- Wasternack, C.; Strnad, M. Jasmonates: News on occurrence, biosynthesis, metabolism and action of an ancient group of signaling compounds. Int. J. Mol. Sci. 2018, 19, 2539. [Google Scholar] [CrossRef]
- Chini, A.; Monte, I.; Zamarreño, A.M.; Hamberg, M.; Lassueur, S.; Reymond, P.; Weiss, S.; Stintzi, A.; Schaller, A.; Porzel, A.; et al. An OPR3-independent pathway uses 4,5-didehydrojasmonate for jasmonate synthesis. Nat. Chem. Biol. 2018, 14, 171–178. [Google Scholar] [CrossRef]
- Wasternack, C.; Forner, S.; Strnad, M.; Hause, B. Jasmonates in flower and seed development. Biochimie 2013, 95, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Yu, G.; Cao, C.; Liu, P. Metabolism, signaling, and transport of jasmonates. Plant Commun. 2021, 2, 100231. [Google Scholar] [CrossRef] [PubMed]
- Shan, X.; Yan, J.; Xie, D. Comparison of phytohormone signaling mechanisms. Curr. Opin. Plant Biol. 2012, 15, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Tani, T.; Sobajima, H.; Okada, K.; Chujo, T.; Arimura, S.-i.; Tsutsumi, N.; Nishimura, M.; Seto, H.; Nojiri, H.; Yamane, H. Identification of the OsOPR7 gene encoding 12-oxophytodienoate reductase involved in the biosynthesis of jasmonic acid in rice. Planta 2007, 227, 517–526. [Google Scholar] [CrossRef]
- Ishiguro, S.; Kawai-Oda, A.; Ueda, J.; Nishida, I.; Okada, K. The DEFECTIVE IN ANTHER DEHISCENCE1 gene encodes a novel phospholipase a1 catalyzing the initial step of jasmonic acid biosynthesis, which synchronizes pollen maturation, anther dehiscence, and flower opening in Arabidopsis. Plant Cell 2001, 13, 2191. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C.; Song, S. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transciption. J. Exp. Bot. 2017, 68, 1303–1321. [Google Scholar] [CrossRef] [PubMed]
- Chauvin, A.; Caldelari, D.; Wolfender, J.L.; Farmer, E.E. Four 13-lipoxygenases contribute to rapid jasmonate synthesis in wounded Arabidopsis thaliana leaves: A role for lipoxygenase 6 in responses to long-distance wound signals. New Phytol. 2012, 197, 566–575. [Google Scholar] [CrossRef]
- Singh, P.; Arif, Y.; Miszczuk, E.; Bajguz, A.; Hayat, S. Specific roles of lipoxygenases in development and responses to stress in plants. Plants 2022, 11, 979. [Google Scholar] [CrossRef]
- Stolterfoht, H.; Rinnofner, C.; Winkler, M.; Pichler, H. Recombinant lipoxygenases and hydroperoxide lyases for the synthesis of green leaf volatiles. J. Agr. Food Chem. 2019, 67, 13367–13392. [Google Scholar] [CrossRef]
- Farmer, E.E.; Goossens, A. Jasmonates: What ALLENE OXIDE SYNTHASE does for plants. J. Exp. Bot. 2019, 70, 3373–3378. [Google Scholar] [CrossRef]
- Christensen, S.A.; Nemchenko, A.; Park, Y.-S.; Borrego, E.; Huang, P.-C.; Schmelz, E.A.; Kunze, S.; Feussner, I.; Yalpani, N.; Meeley, R.; et al. The novel monocot-specific 9-lipoxygenase ZmLOX12 Is required to mount an effective jasmonate-mediated defense against Fusarium verticillioides in maize. Mol. Plant-Microbe Interact. 2014, 27, 1263–1276. [Google Scholar] [CrossRef] [PubMed]
- Jimenez Aleman, G.H.; Thirumalaikumar, V.P.; Jander, G.; Fernie, A.R.; Skirycz, A. OPDA, more than just a jasmonate precursor. Phytochemistry 2022, 204, 113432. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Li, N.; Song, Q.; Li, X.; Meng, H.; Luo, K. OPDAT1, a plastid envelope protein involved in 12-oxo-phytodienoic acid export for jasmonic acid biosynthesis in Populus. Tree Physiol. 2021, 41, 1714–1728. [Google Scholar] [CrossRef] [PubMed]
- Sanders, P.M.; Lee, P.Y.; Biesgen, C.; Boone, J.D.; Beals, T.P.; Weiler, E.W.; Goldberg, R.B. The Arabidopsis DELAYED DEHISCENCE1 gene encodes an enzyme in the jasmonic acid synthesis pathway. Plant Cell 2000, 12, 1041. [Google Scholar] [CrossRef] [PubMed]
- Koo, A.J.K.; Chung, H.S.; Kobayashi, Y.; Howe, G.A. Identification of a peroxisomal acyl-activating enzyme involved in the biosynthesis of jasmonic acid in Arabidopsis. J. Biol. Chem. 2006, 281, 33511–33520. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. A bypass in jasmonate biosynthesis—The OPR3-independent formation. Trends Plant Sci. 2018, 23, 276–279. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. The missing link in jasmonic acid biosynthesis. Nat. Plants 2019, 5, 776–777. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).