Genome-Wide Association Study Identifies Quantitative Trait Loci and Candidate Genes Involved in Deep-Sowing Tolerance in Maize (Zea mays L.)
Abstract
:1. Introduction
2. Results
2.1. Evaluation of Deep-Sowing Tolerance-Related Traits
2.2. Linkage Disequilibrium and Structure Analysis
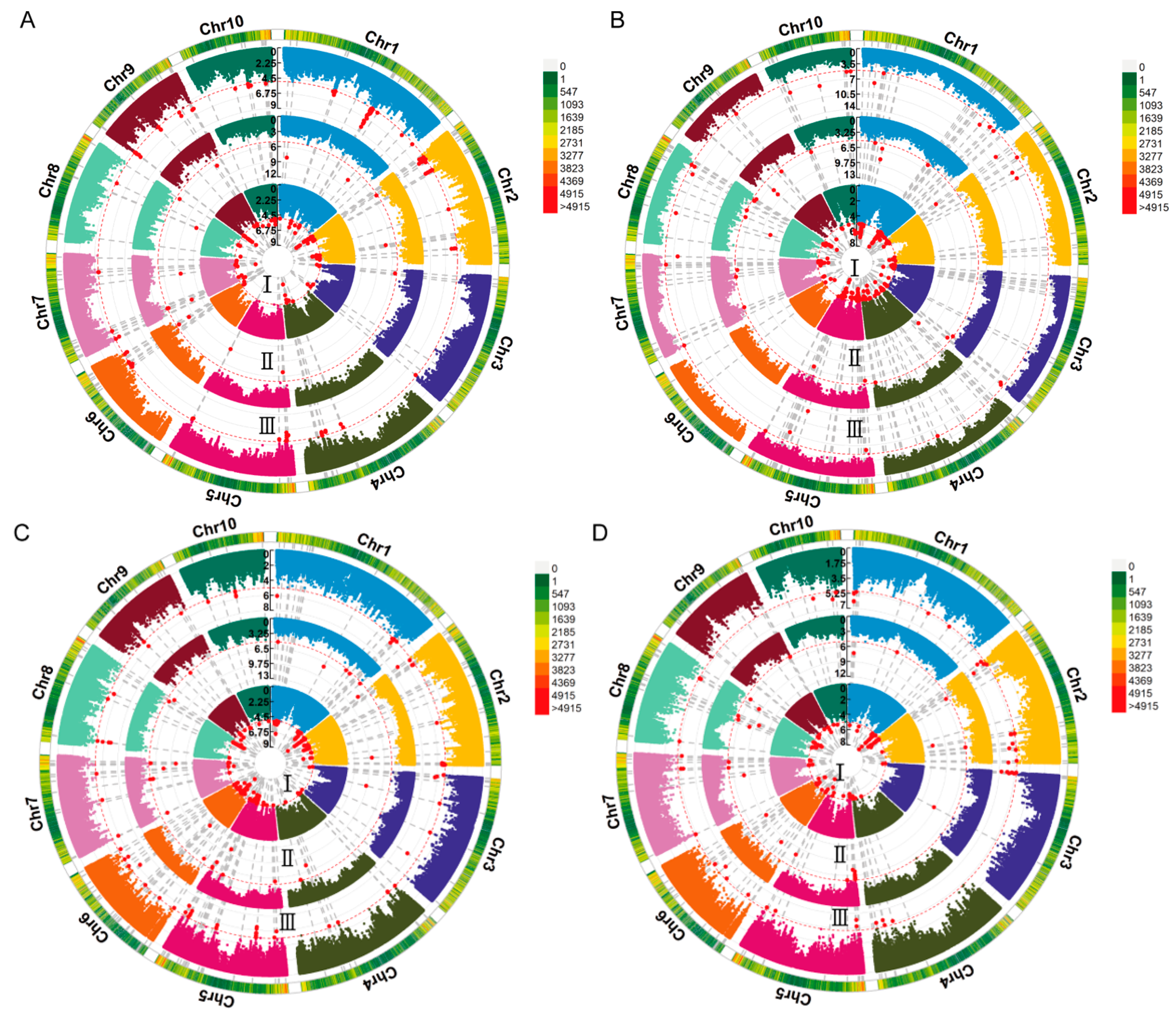
2.3. GWAS of Deep-Sowing Tolerance-Related Traits in Maize
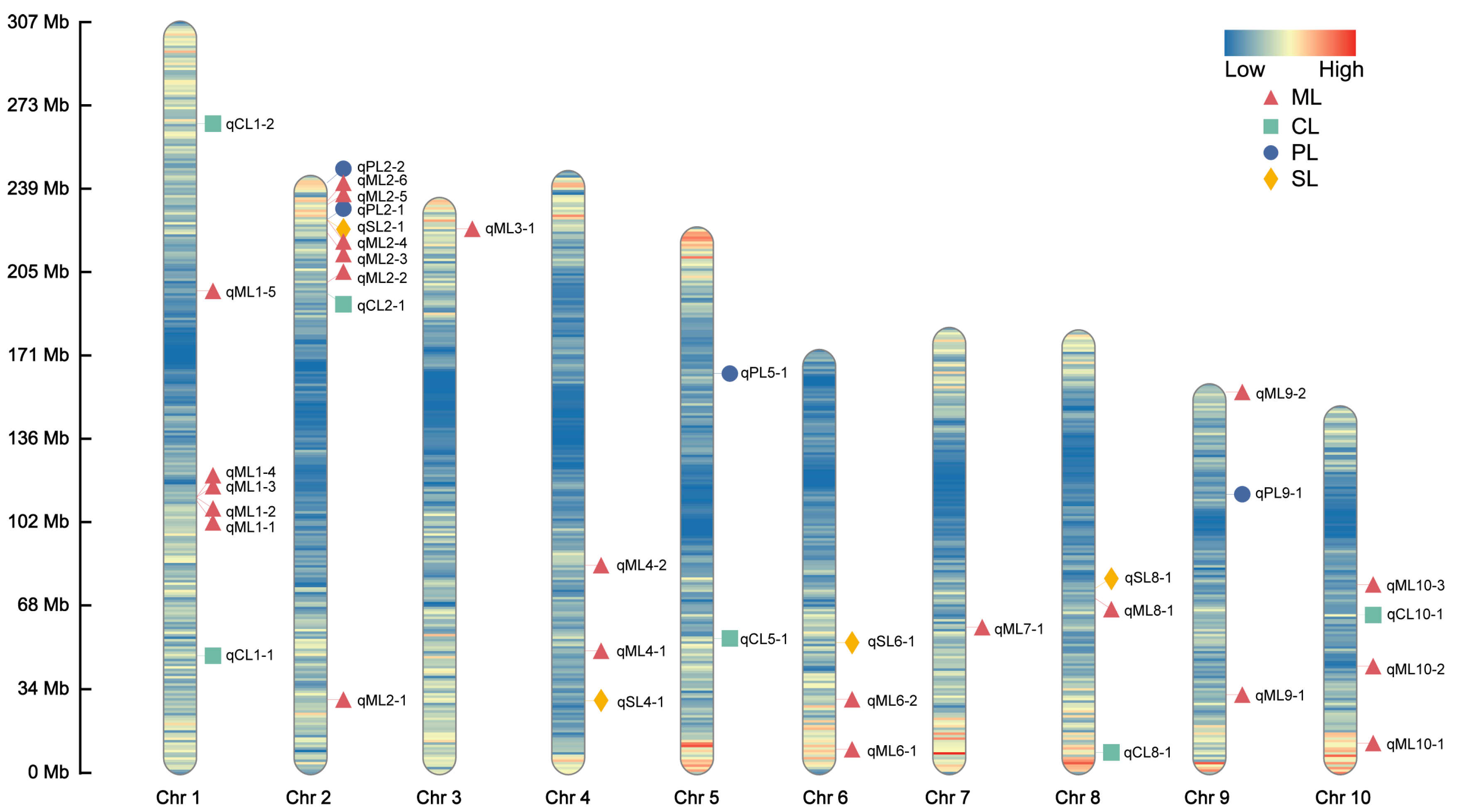
2.4. Candidate Genes Co-Localized by ML, PL, and SL
2.5. Functional Verification of Candidate Gene ZmGCP2
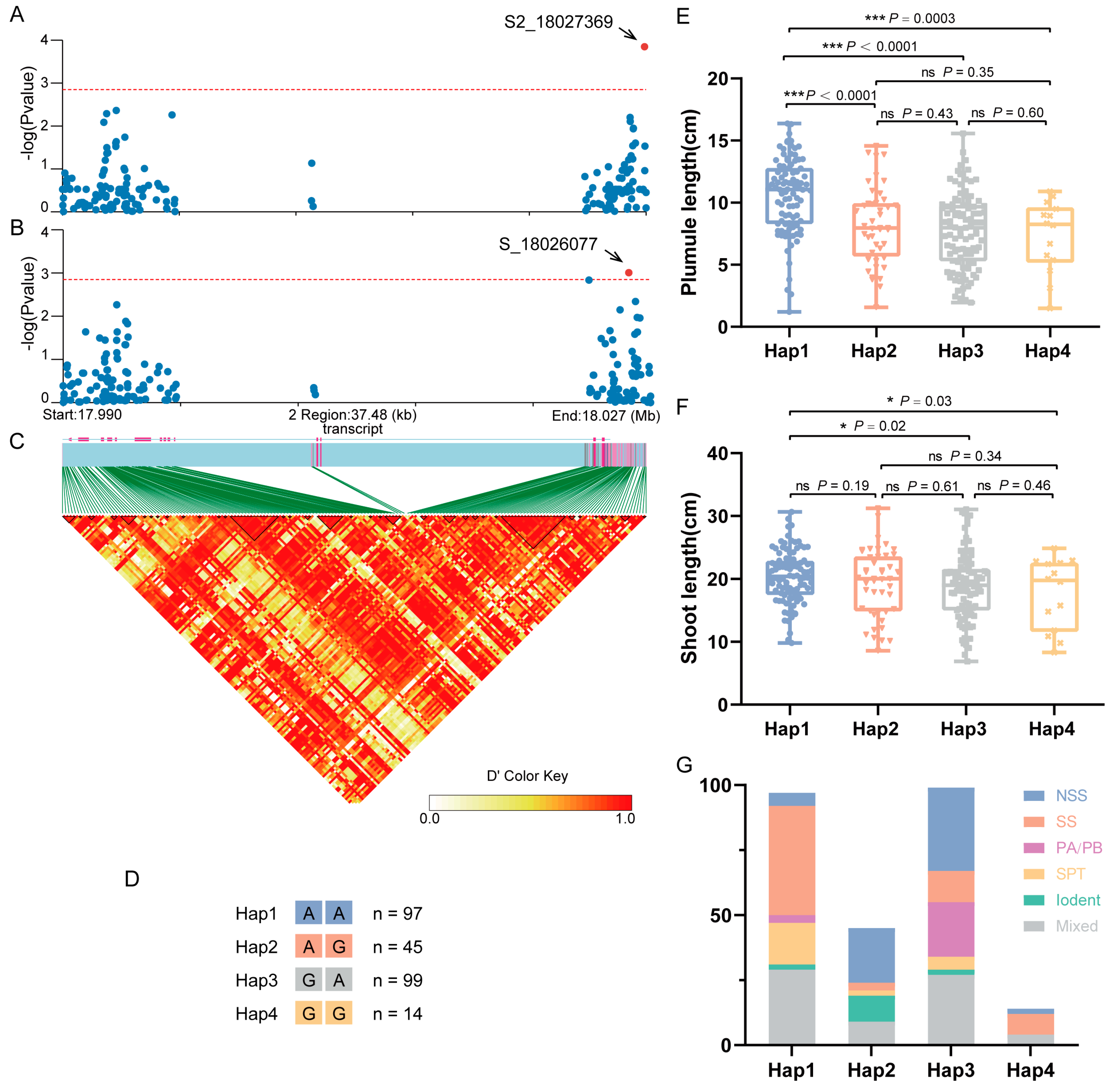
2.6. Natural Variation in ZmGCP2
3. Discussion
3.1. GWAS Is Effective for Dissecting the Genetic Basis of Maize Deep-Sowing Tolerance
3.2. Comparison of Co-Localized QTLs with Previously Reported QTLs
3.3. Functional Analysis of Candidate Gene ZmGCP2
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Phenotyping and Statistical Analysis
4.3. SNP Genotyping
4.4. LD and Structure Analysis
4.5. GWAS
4.6. Gene Expression Patterns Analysis
4.7. Candidate Gene Association Study
4.8. Cell Observation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Iodent | Iowa Experiment Station Reid Yellow Dent |
| NSS | Non-Stiff Stalk |
| PA | Group A germplasm derived from modern U.S. hybrids |
| PB | Group B germplasm derived from modern U.S. hybrids |
| SPT | Si Ping Tou |
| SS | Stiff Stalk |
References
- Hubert, B.; Rosegrant, M.; van Boekel, M.A.; Ortiz, R. The Future of Food: Scenarios for 2050. Crop Sci. 2010, 50, 33–50. [Google Scholar] [CrossRef]
- Wang, Y.Z.; Liu, Z.Q.; Wang, G.Q.; Li, Y. Investigation on Relationship between High Yield and Population’s Uniformity in Maize. J. Maize Sci. 2000, 8, 43–45. [Google Scholar]
- Feng, P.; Wang, B.; Harrison, M.T.; Wang, J.; Liu, K.; Huang, M.; Liu, D.L.; Yu, Q.; Hu, K. Soil Properties Resulting in Superior Maize Yields upon Climate Warming. Agron. Sustain. Dev. 2022, 42, 85. [Google Scholar] [CrossRef]
- Fahad, S.; Bajwa, A.A.; Nazir, U.; Anjum, S.A.; Farooq, A.; Zohaib, A.; Sadia, S.; Nasim, W.; Adkins, S.; Saud, S.; et al. Crop Production under Drought and Heat Stress: Plant Responses and Management Options. Front. Plant Sci. 2017, 8, 1147. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Jiang, H.; Zhao, G.; Ren, J. Gene Cloning of ZmMYB59 Transcription Factor in Maize and Its Expression during Seed Germination in Response to Deep-sowing and Exogenous Hormones. Plant Breed. 2017, 136, 834–844. [Google Scholar] [CrossRef]
- Grzybowski, M.; Adamczyk, J.; Jończyk, M.; Sobkowiak, A.; Szczepanik, J.; Frankiewicz, K.; Fronk, J.; Sowiński, P. Increased Photosensitivity at Early Growth as a Possible Mechanism of Maize Adaptation to Cold Springs. J. Exp. Bot. 2019, 70, 2887–2904. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.; Zhao, X.Q.; Niu, Y.N.; Chen, X.Y.; Ren, X.W. Brassinosteroid Affects the Elongation of Mesocotyl and Coleoptile in Zea mays L. by Regulating the Network of Circadian Rhythm under Severe Stress. Russ. J. Plant Physiol. 2022, 69, 90. [Google Scholar] [CrossRef]
- Leng, B.; Li, M.; Mu, C.; Yan, Z.; Yao, G.; Kong, X.; Ma, C.; Zhang, F.; Liu, X. Molecular Mechanism of Gibberellins in Mesocotyl Elongation Response to Deep-Sowing Stress in Sweet Maize. Curr. Issues Mol. Biol. 2022, 45, 197–211. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Ma, Y.; Liu, Y.; Lyle, D.; Li, D.; Wang, P.; Xu, J.; Zhen, S.; Lu, J.; Peng, Y.; et al. Dissecting the Genetic Basis of Maize Deep-sowing Tolerance by Combining Association Mapping and Gene Expression Analysis. J. Integr. Agric. 2022, 21, 1266–1277. [Google Scholar] [CrossRef]
- Zhong, G.; Bai, Y.; Meng, J. Application and Development of New Maize Varieties with Drought Resistance, Deep-sowing Tolerance and Higher Yield. J. Maize Sci. 1998, 6, 29–31. [Google Scholar]
- Troyer, A.F. The Location of Genes Governing Long First Internode of Corn. Genetics 1997, 145, 1149–1154. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.W.; Wang, J.H. Effect of Gibberellin and Uniconazole on Mesocotyl Elongation of Dark-Grown Maize under Different Seeding Depths. Plant Prod. Sci. 2015, 11, 423–429. [Google Scholar] [CrossRef]
- Hoshikawa, K. Underground Organs of the Seedlings and the Systematics of Gramineae. Bot. Gaz. 1969, 130, 192–203. [Google Scholar] [CrossRef]
- Dungan, G.H. Response of Corn to Extremely Deep Planting. Agron. J. 1950, 42, 256–257. [Google Scholar] [CrossRef]
- Zhao, G.W.; Ma, P.; Wang, J.H.; Wang, G.Y. Identification of Deep-seeding Tolerance in Different Maize Inbred Lines and Their Physiological Response to Deep-seeding Condition. J. Maize Sci. 2009, 17, 9–13. [Google Scholar]
- Zhao, X.Q.; Niu, Y.N.; Hossain, Z.; Shi, J.; Mao, T.T.; Bai, X.D. Integrated QTL Mapping, Meta-Analysis, and RNA-Sequencing Reveal Candidate Genes for Maize Deep-Sowing Tolerance. Int. J. Mol. Sci. 2023, 24, 6770. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Ma, P.; Zhao, Z.; Zhao, G.; Tian, B.; Wang, J.; Wang, G. Mapping QTL Controlling Maize Deep-seeding Tolerance-related Traits and Confirmation of a Major QTL for Mesocotyl Length. Theor. Appl. Genet. 2012, 124, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Copeland, L.O.; McDonald, M.B. Seed Vigor and Vigor Testing; Copeland, L.O., McDonald, M.B., Eds.; Springer: Berlin/Heidelberg, Germany, 2001; pp. 165–191. ISBN 978-1-4615-1619-4. [Google Scholar]
- Lee, H.; Sasaki, K.; Kang, J.; Sato, T.; Song, W.Y.; Ahn, S.N. Mesocotyl Elongation is Essential for Seedling Emergence under Deep-Seeding Condition in Rice. Rice 2017, 10, 32. [Google Scholar] [CrossRef]
- Lee, H.; Sasaki, K.; Higashitani, A.; Ahn, S.N.; Sato, T. Mapping and Characterization of Quantitative Trait Loci for Mesocotyl Elongation in Rice (Oryza sativa L.). Rice 2012, 5, 13. [Google Scholar] [CrossRef]
- Wu, J.; Feng, F.; Lian, X.; Teng, X.; Wei, H.; Yu, H.; Xie, W.; Yan, M.; Fan, P.; Li, Y.; et al. Genome-wide Association Study (GWAS) of Mesocotyl Elongation based on Re-sequencing Approach in Rice. BMC Plant Biol. 2015, 15, 218. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, W.; Jiang, C.; Wang, X.; Xiong, H.; Todorovska, E.G.; Yin, Z.; Chen, Y.; Wang, X.; Xie, J.; et al. Genetic Architecture and Candidate Genes for Deep-Sowing Tolerance in Rice Revealed by Non-syn GWAS. Front. Plant Sci. 2018, 9, 332. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Q.; Ma, B.; Lu, X.; Huang, Y.H.; He, S.J.; Yang, C.; Yin, C.C.; Zhao, H.; Zhou, Y.; Zhang, W.K.; et al. Ethylene-Inhibited Jasmonic Acid Biosynthesis Promotes Mesocotyl/Coleoptile Elongation of Etiolated Rice Seedlings. Plant Cell 2017, 29, 1053–1072. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; Wang, T.; Wang, L.; Li, X.; Jia, Y.; Liu, C.; Huang, X.; Xie, W.; Wang, X. Natural Selection of a GSK3 Determines Rice Mesocotyl Domestication by Coordinating Strigolactone and Brassinosteroid Signaling. Nat. Commun. 2018, 9, 2523. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Shao, G.; Jiao, G.; Sheng, Z.; Xie, L.; Hu, S.; Tang, S.; Wei, X.; Hu, P. Targeted Mutagenesis of POLYAMINE OXIDASE 5 that Negatively Regulates Mesocotyl Elongation Enables the Generation of Direct-seeding Rice with Improved Grain Yield. Mol. Plant 2021, 14, 344–351. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Zhang, L.; Wang, J.; Li, C.; Zeng, X.; Xie, S.; Zhang, Y.; Liu, S.; Hu, S.; Wang, J.; et al. Quantitative Trait Locus Analysis for Deep-Sowing Germination Ability in the Maize IBM Syn10 DH Population. Front. Plant Sci. 2017, 8, 813. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; He, J.; Ye, H.; Ding, M.; Xu, F.; Wu, R.; Zhao, F.; Zhao, G. Transcriptome Analysis Revealed the Key Genes and Pathways Involved in Seed Germination of Maize Tolerant to Deep-Sowing. Plants 2022, 11, 359. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Warburton, M.; Crouch, J. Association Mapping for Enhancing Maize (Zea mays L.) Genetic Improvement. Crop Sci. 2011, 51, 433–449. [Google Scholar] [CrossRef]
- Yang, Q.; Zhang, D.; Xu, M. A Sequential Quantitative Trait Locus Fine-Mapping Strategy Using Recombinant-Derived Progeny. J. Integr. Plant Biol. 2012, 54, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Peng, Z.; Yang, X.; Wang, W.; Fu, J.; Wang, J.; Han, Y.; Chai, Y.; Gou, T.; Yang, N.; et al. Genome-wide Association Study Dissects the Genetic Architecture of Oil Biosynthesis in Maize Kernels. Nat. Genet. 2013, 45, 43–50. [Google Scholar] [CrossRef]
- Zhang, H.; Zhang, J.; Xu, Q.; Wang, D.; Di, H.; Huang, J.; Yang, X.; Wang, Z.; Zhang, L.; Dong, L.; et al. Identification of Candidate Tolerance Genes to Low-temperature During Maize Germination by GWAS and RNA-seq Approaches. BMC Plant Biol. 2020, 20, 333. [Google Scholar] [CrossRef]
- Zhang, Y.; Hu, Y.; Guan, Z.; Liu, P.; He, Y.; Zou, C.; Li, P.; Gao, S.; Peng, H.; Yang, C.; et al. Combined Linkage Mapping and Association Analysis Reveals Genetic Control of Maize Kernel Moisture Content. Physiol. Plant. 2020, 170, 508–518. [Google Scholar] [CrossRef] [PubMed]
- Rafalski, J.A. Association Genetics in Crop Improvement. Curr. Opin. Plant Biol. 2010, 13, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Gore, M.; Buckler, E.S.; Yu, J. Status and Prospects of Association Mapping in Plants. Plant Genome 2008, 1, 5–20. [Google Scholar] [CrossRef]
- Wang, X.; Wang, H.; Liu, S.; Ferjani, A.; Li, J.; Yan, J.; Yang, X.; Qin, F. Genetic variation in ZmVPP1 contributes to drought tolerance in maize seedlings. Nat. Genet. 2016, 48, 1233–1241. [Google Scholar] [CrossRef] [PubMed]
- Wasteneys, G.O.; Yang, Z. New Views on the Plant Cytoskeleton. Plant Physiol. 2004, 136, 3884–3891. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Zhang, X.; Liu, X.; Zhao, Y.; Wang, B.; Zhang, J. MiRNA Alterations are Important Mechanism in Maize Adaptations to Low-phosphate Environments. Plant Sci. 2016, 252, 103–117. [Google Scholar] [CrossRef]
- Teng, L.; Zhang, X.; Wang, R.; Lin, K.; Zeng, M.; Chen, H.; Cao, F. MiRNA Transcriptome Reveals Key MiRNAs and Their Targets Contributing to the Difference in Cd Tolerance of Two Contrasting Maize Genotypes. Ecotoxicol. Environ. Saf. 2023, 256, 114881. [Google Scholar] [CrossRef] [PubMed]
- Qu, Q.; Liu, N.; Su, Q.; Liu, X.; Jia, H.; Liu, Y.; Sun, M.; Cao, Z.; Dong, J. MicroRNAs Involved in the Trans-kingdom Gene Regulation in the Interaction of Maize Kernels and Fusarium verticillioides. Int. J. Biol. Macromol. 2023, 242, 125046. [Google Scholar] [CrossRef]
- Stelpflug, S.C.; Sekhon, R.S.; Vaillancourt, B.; Hirsch, C.N.; Buell, C.R.; de Leon, N.; Kaeppler, S.M. An Expanded Maize Gene Expression Atlas based on RNA Sequencing and its Use to Explore Root Development. Plant Genome 2016, 9, 1–16. [Google Scholar] [CrossRef]
- Guo, Z.; Kuang, Z.; Wang, Y.; Zhao, Y.; Tao, Y.; Cheng, C.; Yang, J.; Lu, X.; Hao, C.; Wang, T.; et al. PmiREN: A comprehensive encyclopedia of plant miRNAs. Nucleic Acids Res. 2020, 48, D1114–D1121. [Google Scholar] [CrossRef]
- Schaefer, E.; Belcram, K.; Uyttewaal, M.; Duroc, Y.; Goussot, M.; Legland, D.; Laruelle, D.; de Tauzia-Moreau, M.L.; Pastuglia, M.; Bouchez, D. The Preprophase Band of Microtubules Controls the Robustness of Division Orientation in Plants. Science 2017, 356, 186–189. [Google Scholar] [CrossRef] [PubMed]
- Roudier, F.; Fernandez, A.G.; Fujita, M.; Himmelspach, R.; Borner, G.H.; Schindelman, G.; Song, S.; Baskin, T.I.; Dupree, P.; Wasteneys, G.O.; et al. COBRA, an Arabidopsis Extracellular Glycosyl-phosphatidyl Inositol-anchored Protein, Specifically Controls Highly Anisotropic Expansion through Its Involvement in Cellulose Microfibril Orientation. Plant Cell 2005, 17, 1749–1763. [Google Scholar] [CrossRef] [PubMed]
- Chan, J. Microtubule and Cellulose Microfibril Orientation during Plant Cell and Organ Growth. J. Microsc. 2012, 247, 23–32. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, M.; Yang, G.; Kawaguchi, K.; Komatsu, S. Expression Analyses of [beta]-tubulin Isotype Genes in Rice. Plant Cell Physiol. 2003, 44, 1202–1207. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Guo, L.; Qu, G.; Xiang, Y.; Zhao, X.; Yuan, H.; Li, T.; Kang, L.; Tang, S.; Tu, B.; et al. Wide grain 4, Encoding an Alpha-tubulin, Regulates Grain Size by Affecting Cell Expansion in Rice. Crop J. 2023, 11, 1931–1936. [Google Scholar] [CrossRef]
- Li, W.; Ge, F.; Qiang, Z.; Zhang, S.; Chen, L.; Wang, X.; Li, J.; Fu, Y. Maize ZmRPH1 Encodes a Microtubule-associated Protein that Controls Plant and Ear Height. Plant Biotechnol. J. 2020, 18, 1345–1347. [Google Scholar] [CrossRef] [PubMed]
- Yan, P.; Du, Q.; Chen, H.; Guo, Z.; Wang, Z.; Tang, J.; Li, W.X. Biofortification of Iron Content by Regulating a NAC Transcription Factor in Maize. Science 2023, 382, 1159–1165. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.L.; Gao, Z.Y.; Shang, L.G.; Yang, C.H.; Ruan, B.P.; Zeng, D.L.; Guo, L.B.; Zhao, F.J.; Huang, C.F.; Qian, Q. Natural Variation in the Promoter of OsHMA3 Contributes to Differential Grain Cadmium Accumulation between Indica and Japonica Rice. J. Integr. Plant Biol. 2020, 62, 314–329. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Liu, J.; Ren, W.; Yang, Q.; Chai, Z.; Chen, R.; Wang, L.; Zhao, J.; Lang, Z.; Wang, H.; et al. Gene-Indexed Mutations in Maize. Mol. Plant 2018, 11, 496–504. [Google Scholar] [CrossRef]
- Abdel-Ghani, A.H.; Sanchez, D.L.; Kumar, B.; Lubberstedt, T. Paper Roll Culture and Assessment of Maize Root Parameters. Bio Protoc. 2016, 6, e1926. [Google Scholar] [CrossRef]
- Raj, A.; Stephens, M.; Pritchard, J.K. fastSTRUCTURE: Variational Inference of Population Structure in Large SNP Data Sets. Genetics 2014, 197, 573–589. [Google Scholar] [CrossRef] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the Number of Clusters of Individuals Using the Software STRUCTURE: A Simulation Study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Lee, T.; Guo, H.; Wang, X.; Kim, C.; Paterson, A.H. SNPhylo: A Pipeline to Construct a Phylogenetic Tree from Huge SNP Data. BMC Genomics 2014, 15, 162. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, H.; Li, L.; Lan, H.; Ren, Z.; Liu, D.E.; Wu, L.; Liu, H.; Jaqueth, J.; Li, B.; et al. Characterizing the Population Structure and Genetic Diversity of Maize Breeding Germplasm in Southwest China using Genome-wide SNP Markers. BMC Genomics 2016, 17, 697. [Google Scholar] [CrossRef]
- Yang, X.; Gao, S.; Xu, S.; Zhang, Z.; Prasanna, B.M.; Li, L.; Li, J.; Yan, J. Characterization of a Global Germplasm Collection and Its Potential Utilization for Analysis of Complex Quantitative Traits in Maize. Mol. Breed. 2011, 28, 511–526. [Google Scholar] [CrossRef]
- Wang, B.; Hou, M.; Shi, J.; Ku, L.; Song, W.; Li, C.; Ning, Q.; Li, X.; Li, C.; Zhao, B.; et al. De Novo Genome Assembly and Analyses of 12 Founder Inbred Lines Provide Insights into Maize Heterosis. Nat. Genet. 2023, 55, 312–323. [Google Scholar] [CrossRef]
- Zhang, C.; Dong, S.; Xu, J.; He, W.M.; Yang, T.L. PopLDdecay: A Fast and Effective Tool for Linkage Disequilibrium Decay Analysis Based on Variant Call Format Files. Bioinformatics 2019, 35, 1786–1788. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Zhang, H.; Tang, Z.; Xu, J.; Yin, D.; Zhang, Z.; Yuan, X.; Zhu, M.; Zhao, S.; Li, X.; et al. rMVP: A Memory-efficient, Visualization-enhanced, and Parallel-accelerated Tool for Genome-wide Association Study. Genom. Proteom. Bioinform. 2021, 19, 619–628. [Google Scholar] [CrossRef]
- Bradbury, P.J.; Zhang, Z.; Kroon, D.E.; Casstevens, T.M.; Ramdoss, Y.; Buckler, E.S. TASSEL: Software for Association Mapping of Complex Traits in Diverse Samples. Bioinformatics 2007, 23, 2633–2635. [Google Scholar] [CrossRef]


| Env. | Traits | Range (cm) | Mean (cm) | SD | CV% | Skewness | Kurtosis |
|---|---|---|---|---|---|---|---|
| 20ZC | ML | 3.25–21.74 | 10.47 | 3.40 | 31.67% | 0.53 | 0.16 |
| CL | 1.25–7.78 | 4.59 | 1.09 | 24.26% | −0.13 | 0.33 | |
| PL | 1.10–16.35 | 9.16 | 3.49 | 38.14% | −0.07 | −0.63 | |
| SL | 5.13–33.37 | 19.44 | 5.14 | 26.44% | −0.17 | 0.04 | |
| PRL | 4.85–36.85 | 23.95 | 6.53 | 27.27% | −0.48 | −0.28 | |
| 20YN | ML | 3.00–24.25 | 10.74 | 3.59 | 33.43% | 0.66 | 0.74 |
| CL | 1.40–8.02 | 4.51 | 1.08 | 23.88% | 0.15 | 0.67 | |
| PL | 1.4–17.38 | 8.84 | 3.43 | 38.80% | −0.04 | −0.66 | |
| SL | 6.03–33.83 | 19.48 | 5.45 | 27.96% | −0.10 | −0.31 | |
| PRL | 3.20–40.10 | 24.48 | 7.38 | 30.14% | −0.48 | 0.10 | |
| 21CZ | ML | 2.70–22.03 | 10.79 | 3.72 | 34.47% | 0.40 | −0.08 |
| CL | 1.52–7.57 | 4.44 | 1.03 | 23.19% | −0.17 | 0.24 | |
| PL | 1.20–17.75 | 8.97 | 3.66 | 40.73% | −0.10 | −0.55 | |
| SL | 4.70–31.68 | 19.30 | 5.35 | 27.74% | −0.30 | −0.22 | |
| PRL | 2.00–40.00 | 25.38 | 7.38 | 29.07% | −0.71 | 0.05 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, J.; Liu, Z.; Liu, Y.; Fan, X.; Gao, L.; Li, Y.; Hu, Y.; Hu, K.; Huang, Y. Genome-Wide Association Study Identifies Quantitative Trait Loci and Candidate Genes Involved in Deep-Sowing Tolerance in Maize (Zea mays L.). Plants 2024, 13, 1533. https://doi.org/10.3390/plants13111533
Yang J, Liu Z, Liu Y, Fan X, Gao L, Li Y, Hu Y, Hu K, Huang Y. Genome-Wide Association Study Identifies Quantitative Trait Loci and Candidate Genes Involved in Deep-Sowing Tolerance in Maize (Zea mays L.). Plants. 2024; 13(11):1533. https://doi.org/10.3390/plants13111533
Chicago/Turabian StyleYang, Jin, Zhou Liu, Yanbo Liu, Xiujun Fan, Lei Gao, Yangping Li, Yufeng Hu, Kun Hu, and Yubi Huang. 2024. "Genome-Wide Association Study Identifies Quantitative Trait Loci and Candidate Genes Involved in Deep-Sowing Tolerance in Maize (Zea mays L.)" Plants 13, no. 11: 1533. https://doi.org/10.3390/plants13111533





