Genome-Wide Analysis of the Histone Modification Gene (HM) Family and Expression Investigation during Anther Development in Rice (Oryza sativa L.)
Abstract
:1. Introduction
2. Results
2.1. Characterization of the HM Gene Family in the Rice Genome
2.2. Chromosome Distribution of OsHM Members in the Rice Genome
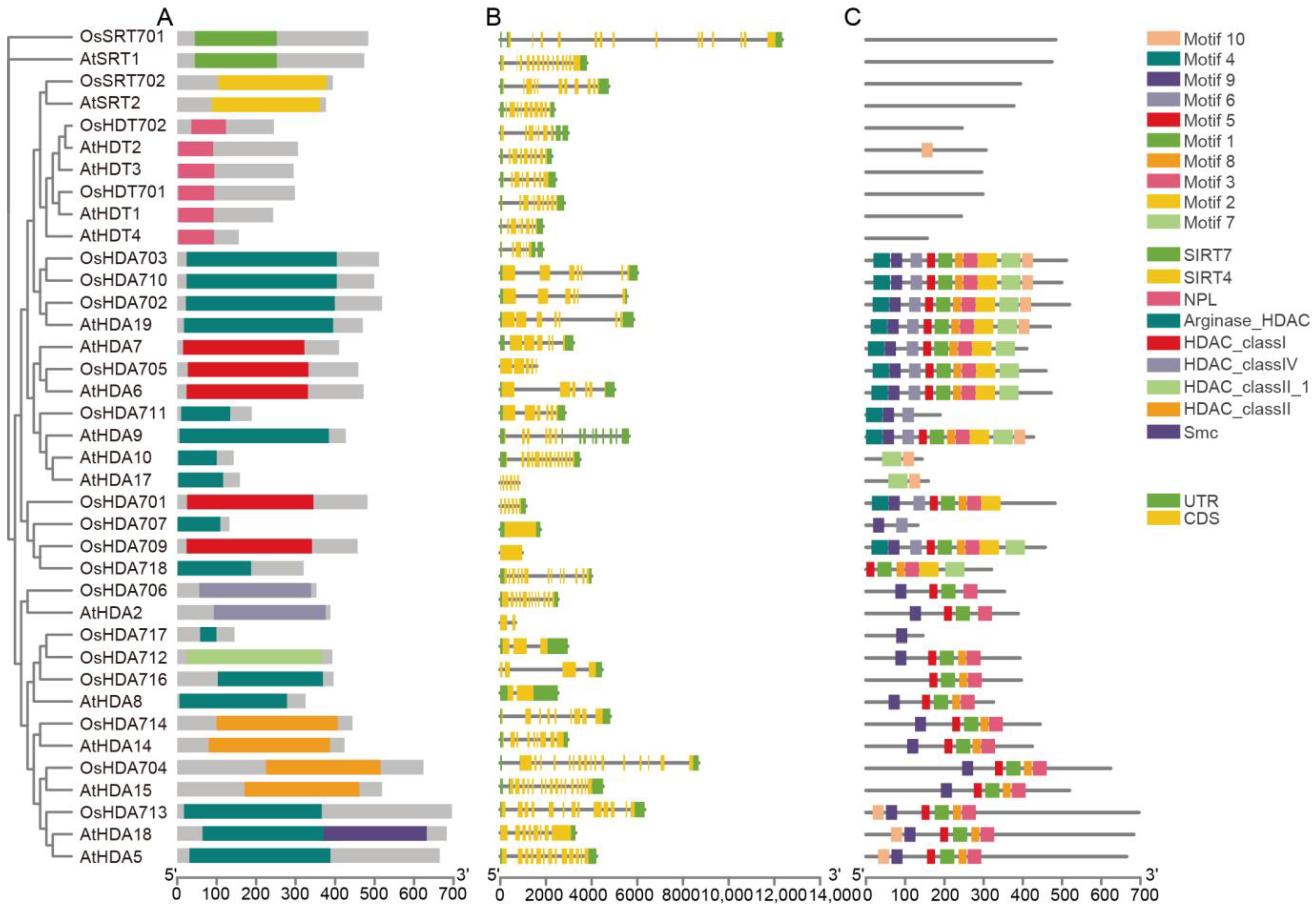
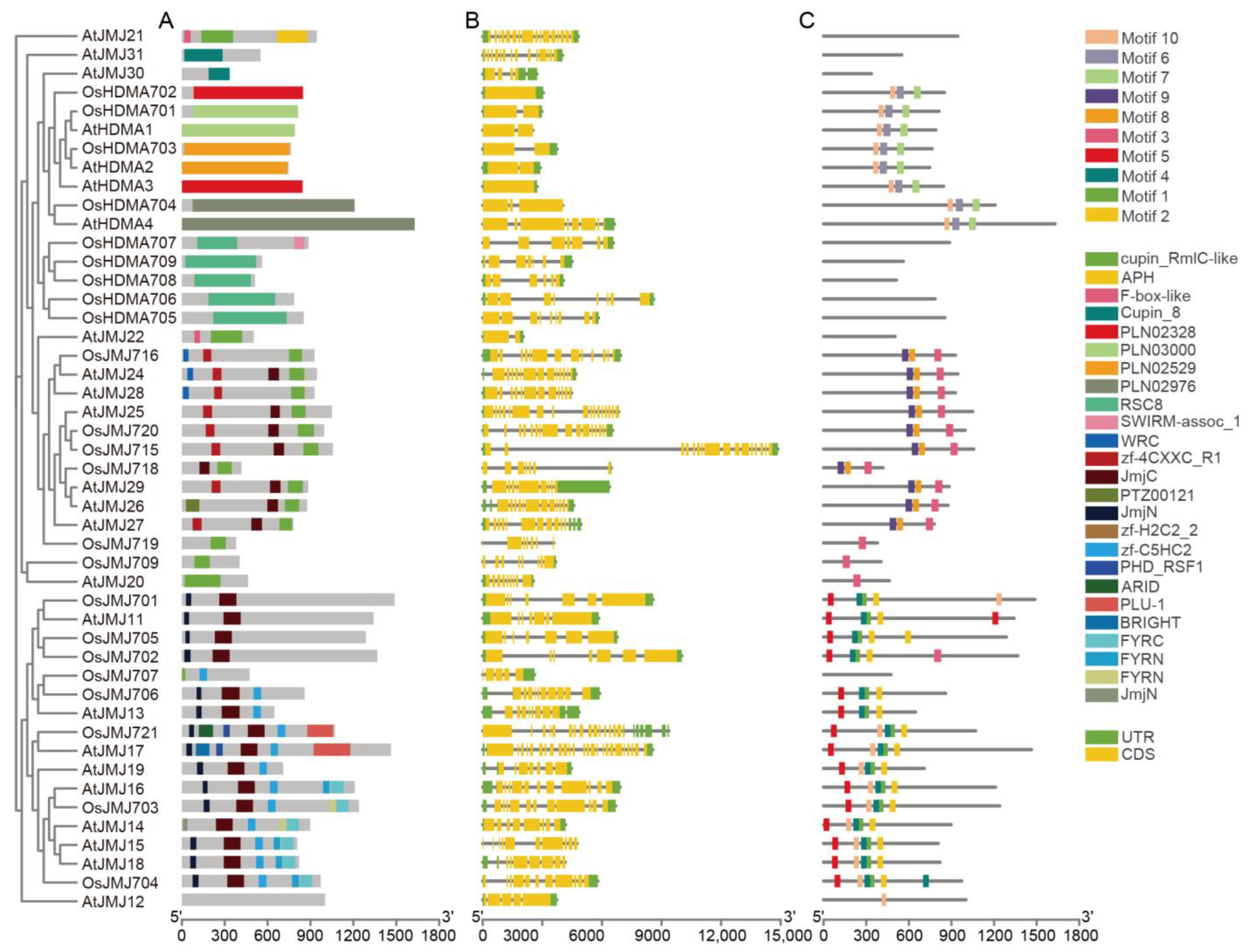
2.3. Gene Structure, Conserved Motifs, and Phylogenetic Analysis of OsHM Gene Family Members
2.4. OsHM Gene Duplication Events in the Rice Genome
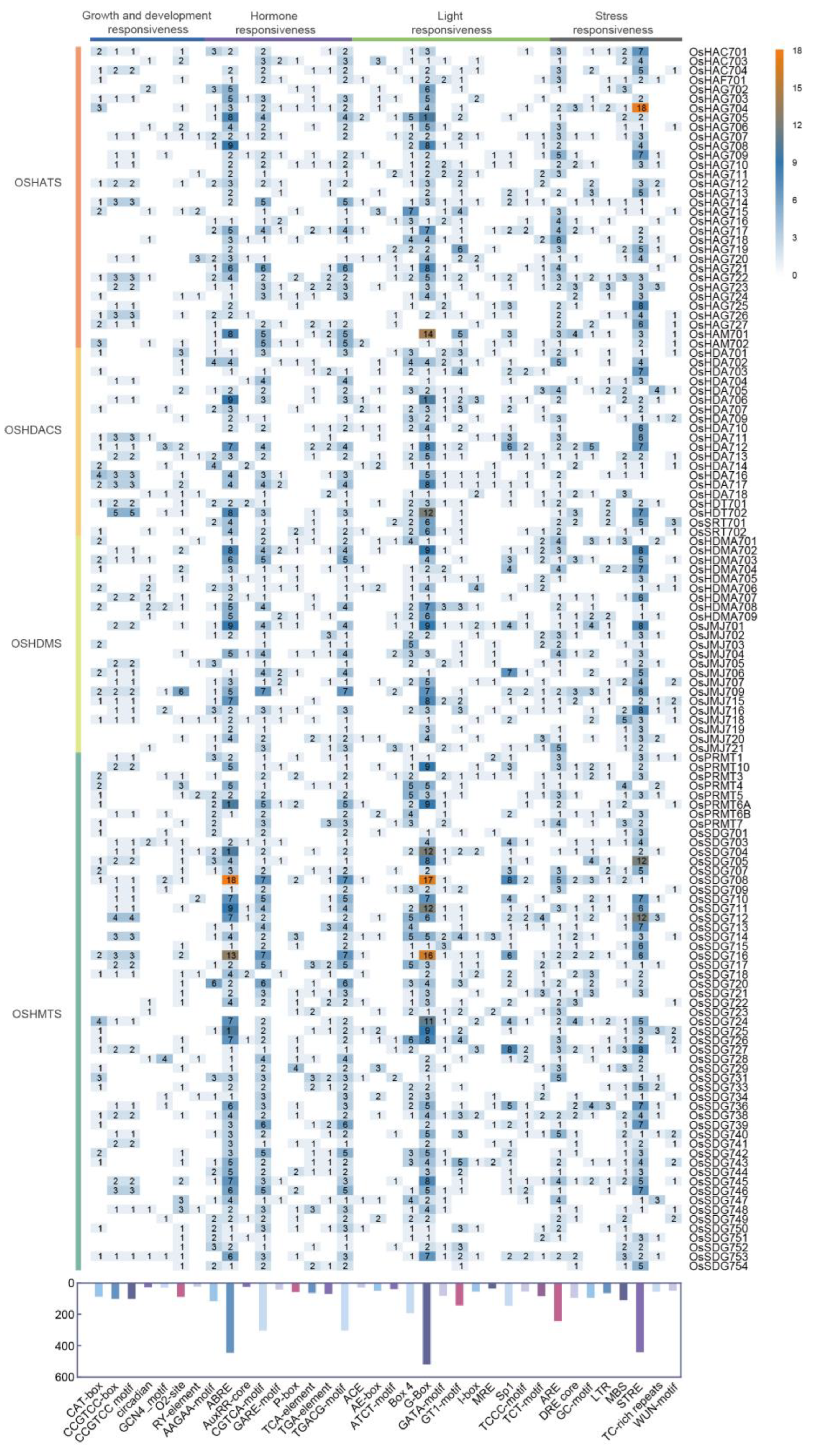
2.5. Cis-Regulatory Elements Analysis of OsHMs
2.6. Tissue-Specific Expression Pattern Analysis of OsHMs in Rice
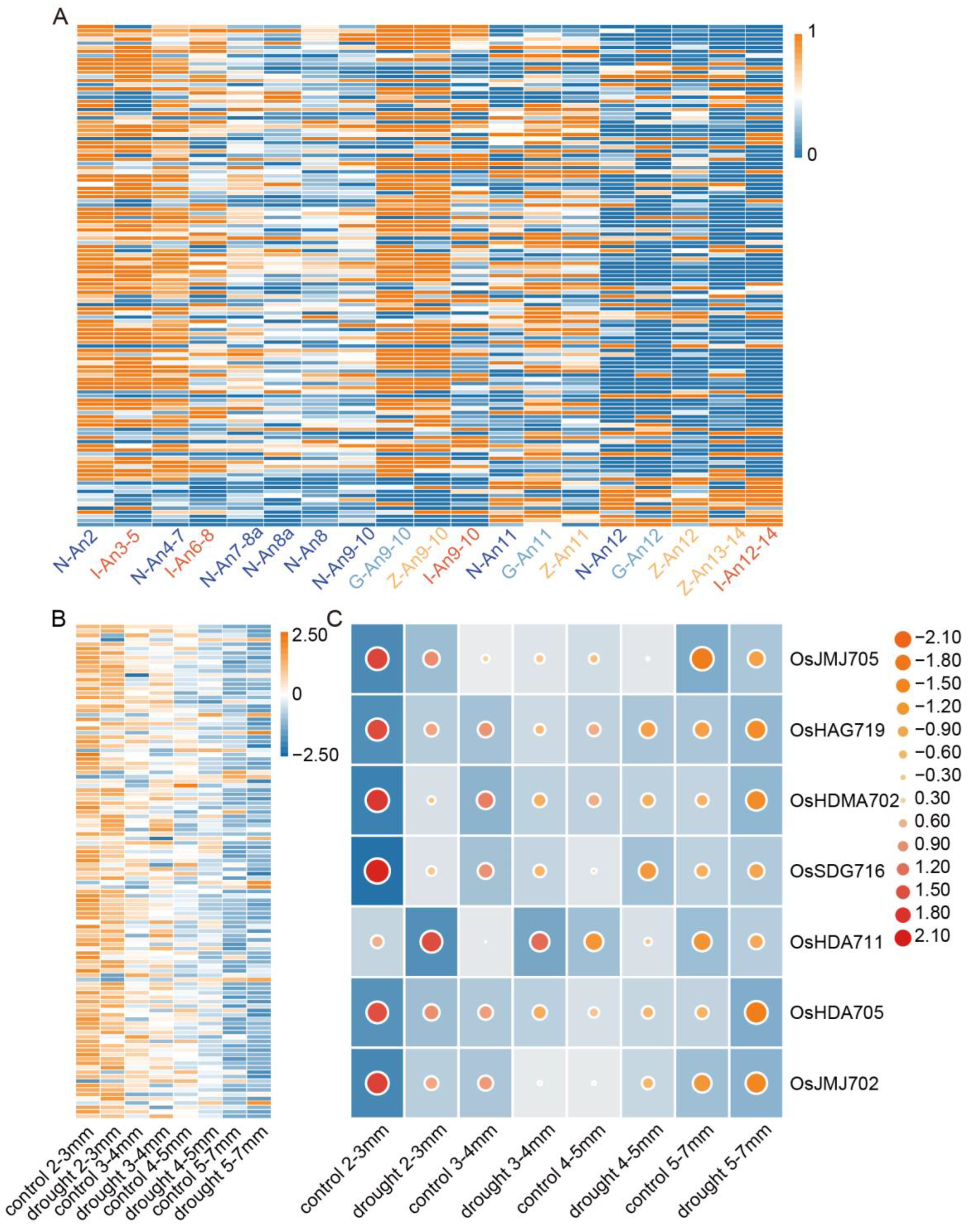
2.7. Abiotic Stress Response Analysis of OsHMs in Rice
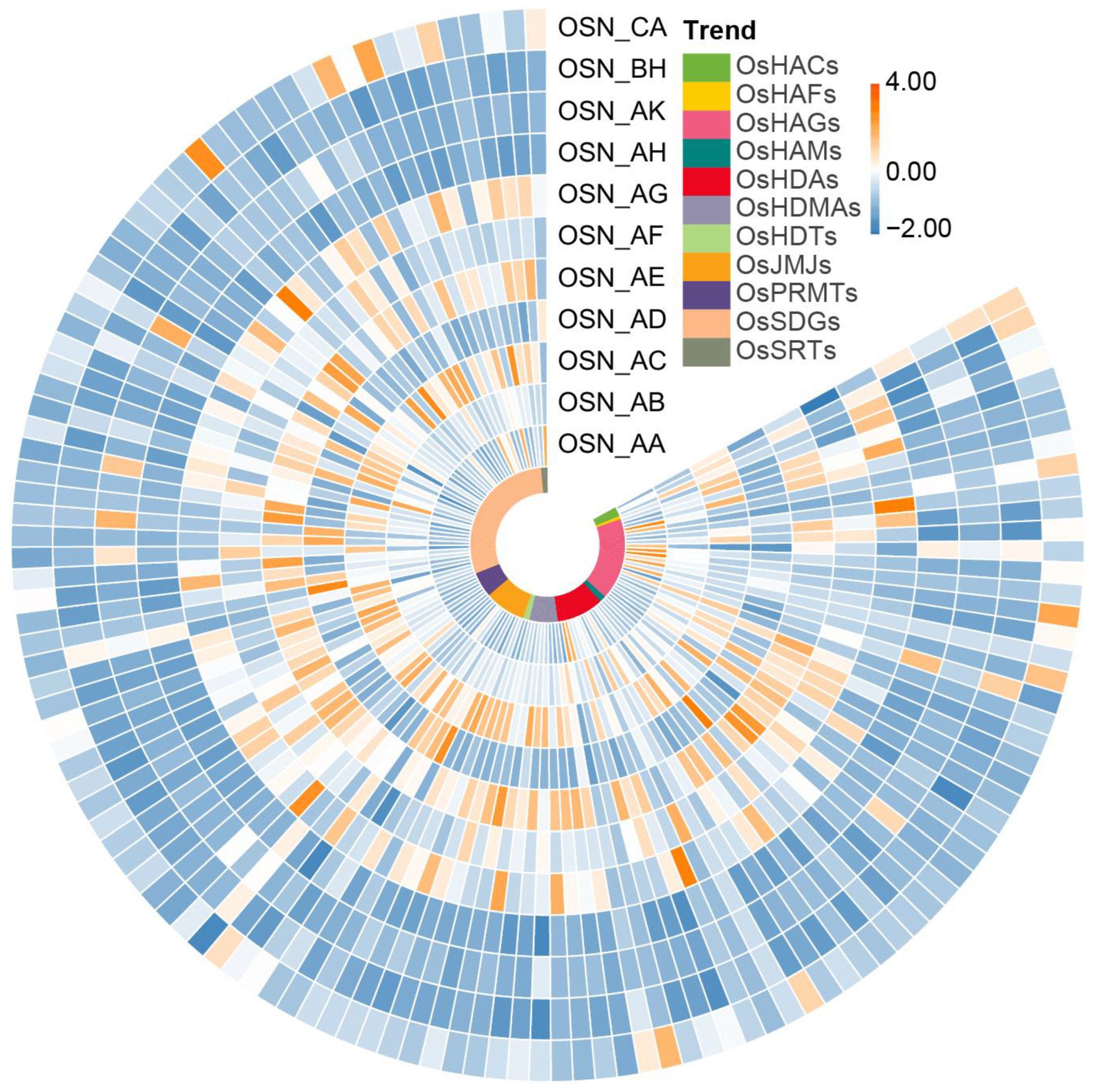
2.8. Expression Pattern Analysis of OsHMs in Rice Anthers Development
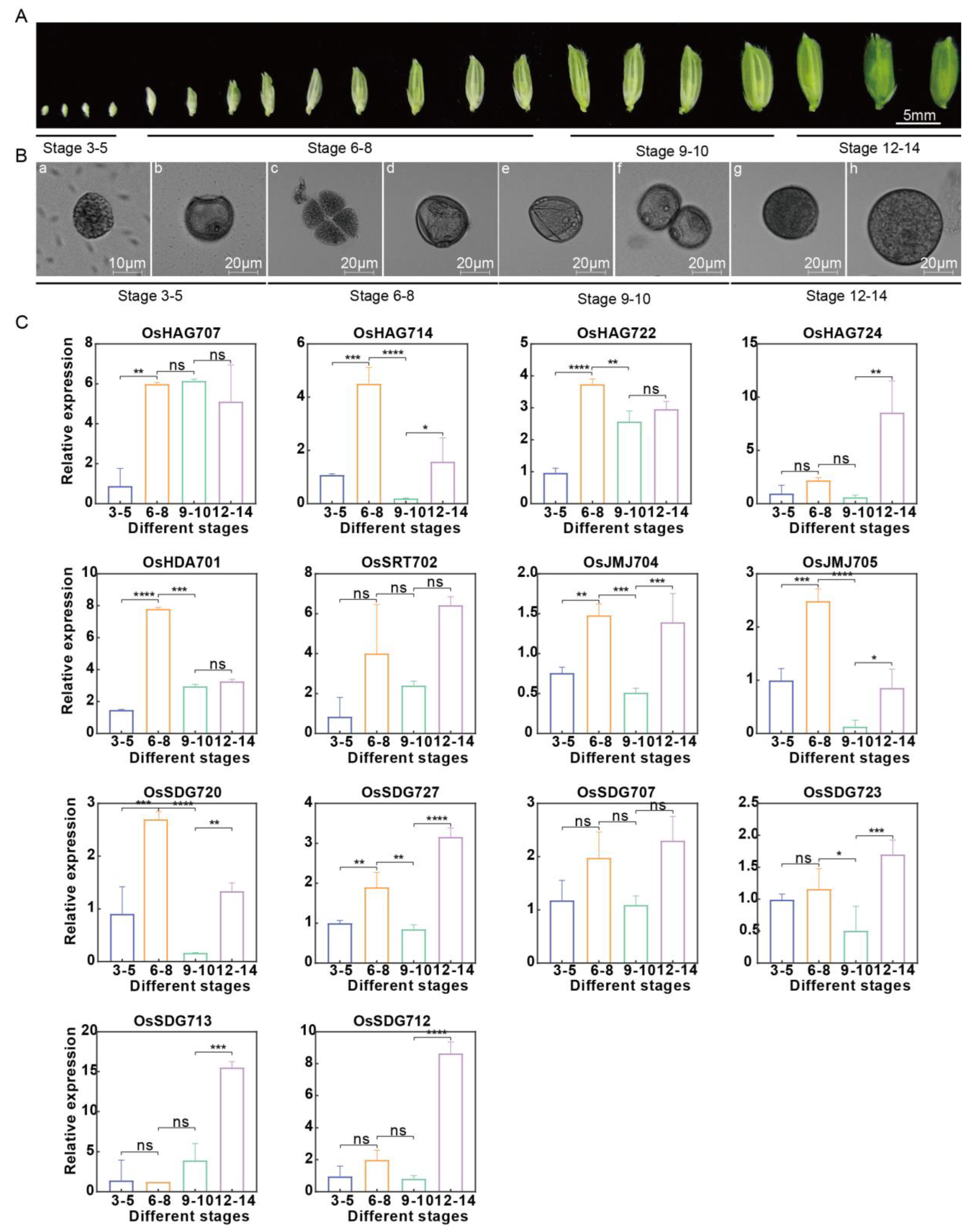
2.9. Expression Analysis of 14 Key Anther Development Genes at Different Anther Development Stages
2.10. Subcellular Localization Analysis of OsSDG713 and OsSDG727
3. Discussion
4. Materials and Methods
4.1. Identification of OsHMs in the Rice Genome
4.2. Gene Structure, Conserved Motif Analysis of OsHMs, and Phylogenetic Relationships of OsHM Genes
4.3. Chromosomal Localization of Rice OsHM Genes
4.4. Cis-Acting Element Analysis
4.5. Expression Analysis of OsHMs in Different Tissue Parts, Different Anther Development Stages, and Different Stress Conditions in Rice
4.6. RT-qPCR Expression Analysis of OsHM Genes
4.7. Plasmid Construction and Subcellular Localization Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Zhang, X. The Epigenetic Landscape of Plants. Science 2008, 320, 489–492. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Cao, X.; Deng, X. Histone Methylation in Epigenetic Regulation and Temperature Responses. Curr. Opin. Plant Biol. 2021, 61, 102001. [Google Scholar] [CrossRef] [PubMed]
- Cheng, K.; Xu, Y.; Yang, C.; Ouellette, L.; Niu, L.; Zhou, X.; Chu, L.; Zhuang, F.; Liu, J.; Wu, H.; et al. Histone Tales: Lysine Methylation, a Protagonist in Arabidopsis Development. J. Exp. Bot. 2019, 71, erz435. [Google Scholar] [CrossRef]
- Zhang, H.; Lang, Z.; Zhu, J.-K. Dynamics and Function of DNA Methylation in Plants. Nat. Rev. Mol. Cell Bio. 2018, 19, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Liu, Y.; Liang, Y.; Zhou, D.; Li, S.; Lin, S.; Dong, H.; Huang, L. The Function of Histone Lysine Methylation Related SET Domain Group Proteins in Plants. Protein Sci. 2020, 29, 1120–1137. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Wang, X.; Su, L.; Zhai, J.; Cao, S.; Zhang, D.; Liu, C.; Bi, Y.; Qian, Q.; Cheng, Z.; et al. SDG714, a Histone H3K9 Methyltransferase, Is Involved in Tos17 DNA Methylation and Transposition in Rice. Plant Cell 2007, 19, 9–22. [Google Scholar] [CrossRef]
- Song, T.; Zhang, Q.; Wang, H.; Han, J.; Xu, Z.; Yan, S.; Zhu, Z. OsJMJ703, a Rice Histone Demethylase Gene, Plays Key Roles in Plant Development and Responds to Drought Stress. Plant Physiol. Biochem. 2018, 132, 183–188. [Google Scholar] [CrossRef]
- Jia, J.; Luo, Y.; Wu, Z.; Ji, Y.; Liu, S.; Shu, J.; Chen, B.; Liu, J. OsJMJ718, a Histone Demethylase Gene, Positively Regulates Seed Germination in Rice. Plant J. 2024, 118, 191–202. [Google Scholar] [CrossRef]
- Kumar, V.; Thakur, J.K.; Prasad, M. Histone Acetylation Dynamics Regulating Plant Development and Stress Responses. Cell Mol. Life Sci. 2021, 78, 4467–4486. [Google Scholar] [CrossRef]
- Tian, L.; Fong, M.P.; Wang, J.J.; Wei, N.E.; Jiang, H.; Doerge, R.W.; Chen, Z.J. Reversible Histone Acetylation and Deacetylation Mediate Genome-Wide, Promoter-Dependent and Locus-Specific Changes in Gene Expression During Plant Development. Genetics 2005, 169, 337–345. [Google Scholar] [CrossRef]
- Jiang, J.; Ding, A.B.; Liu, F.; Zhong, X. Linking Signaling Pathways to Histone Acetylation Dynamics in Plants. J. Exp. Bot. 2020, 71, 5179–5190. [Google Scholar] [CrossRef]
- Li, S.; He, X.; Gao, Y.; Zhou, C.; Chiang, V.L.; Li, W. Histone Acetylation Changes in Plant Response to Drought Stress. Genes 2021, 12, 1409. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Baek, D.; Cha, J.-Y.; Liao, X.; Kang, S.-H.; McClung, C.R.; Lee, S.Y.; Yun, D.-J.; Kim, W.-Y. HOS15 Interacts with the Histone Deacetylase HDA9 and the Evening Complex to Epigenetically Regulate the Floral Activator GIGANTEA. Plant Cell 2019, 31, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Chen, X.; Chen, H.; Wu, K.; Huang, S. Histone Deacetylases HDA6 and HDA9 Coordinately Regulate Valve Cell Elongation through Affecting Auxin Signaling in Arabidopsis. Biochem. Biophys. Res. Commun. 2019, 508, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; He, J.; Velanis, C.N.; Zhu, Y.; He, Y.; Tang, K.; Zhu, M.; Graser, L.; De Leau, E.; Wang, X.; et al. A Domesticated Harbinger Transposase Forms a Complex with HDA6 and Promotes Histone H3 Deacetylation at Genes but Not TEs in Arabidopsis. J. Integr. Plant Biol. 2021, 63, 1462–1474. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhang, J.; Zhang, W.; Wu, K.; Zheng, F.; Tian, L.; Liu, X.; Duan, J. Expression and Functional Analysis of the Plant-Specific Histone Deacetylase HDT701 in Rice. Front. Plant Sci. 2015, 5, 764. [Google Scholar] [CrossRef]
- Sun, Y.; Xie, Z.; Jin, L.; Qin, T.; Zhan, C.; Huang, J. Histone Deacetylase OsHDA716 Represses Rice Chilling Tolerance by Deacetylating OsbZIP46 to Reduce Its Transactivation Function and Protein Stability. Plant Cell 2024, 36, 1913–1936. [Google Scholar] [CrossRef]
- Dresselhaus, T.; Sprunck, S.; Wessel, G.M. Fertilization Mechanisms in Flowering Plants. Curr. Biol. 2016, 26, R125–R139. [Google Scholar] [CrossRef]
- Ashapkin, V.V.; Kutueva, L.I.; Aleksandrushkina, N.I.; Vanyushin, B.F. Epigenetic Regulation of Plant Gametophyte Development. Int. J. Mol. Sci. 2019, 20, 3051. [Google Scholar] [CrossRef]
- Guo, S.; Sun, B.; Looi, L.-S.; Xu, Y.; Gan, E.-S.; Huang, J.; Ito, T. Co-Ordination of Flower Development Through Epigenetic Regulation in Two Model Species: Rice and Arabidopsis. Plant Cell Physiol. 2015, 56, 830–842. [Google Scholar] [CrossRef]
- Gan, E.-S.; Huang, J.; Ito, T. Functional Roles of Histone Modification, Chromatin Remodeling and MicroRNAs in Arabidopsis Flower Development. Int. Rev. Cell Mol. Biol. 2013, 305, 115–161. [Google Scholar] [CrossRef]
- Thorstensen, T.; Grini, P.; Marjara, I.; Alm, V.; Erdal, S.; Aasland, R.; Aalen, R. The Arabidopsis SET-Domain Protein ASHR3 Is Involved in Stamen Development and Interacts with the bHLH Transcription Factor ABORTED MICROSPORES (AMS). Plant Mol. Biol. 2008, 66, 47–59. [Google Scholar] [CrossRef] [PubMed]
- Grini, P.E.; Thorstensen, T.; Alm, V.; Vizcay-Barrena, G.; Windju, S.S.; Jørstad, T.S.; Wilson, Z.A.; Aalen, R.B. The ASH1 HOMOLOG 2 (ASHH2) Histone H3 Methyltransferase Is Required for Ovule and Anther Development in Arabidopsis. PLoS ONE 2009, 4, e7817. [Google Scholar] [CrossRef] [PubMed]
- Jacob, Y.; Feng, S.; LeBlanc, C.A.; Bernatavichute, Y.V.; Stroud, H.; Cokus, S.; Johnson, L.M.; Pellegrini, M.; Jacobsen, S.E.; Michaels, S.D. ATXR5 and ATXR6 Are Novel H3K27 Monomethyltransferases Required for Chromatin Structure and Gene Silencing. Nat. Struct. Mol. Biol. 2009, 16, 763–768. [Google Scholar] [CrossRef]
- Ma, Z.; Castillo-González, C.; Wang, Z.; Sun, D.; Hu, X.; Shen, X.; Potok, M.E.; Zhang, X. Arabidopsis Serrate Coordinates Histone Methyltransferases ATXR5/6 and RNA Processing Factor RDR6 to Regulate Transposon Expression. Dev. Cell 2018, 45, 769–784.e6. [Google Scholar] [CrossRef]
- Zhou, L.; Yarra, R.; Jin, L.; Yang, Y.; Cao, H.; Zhao, Z. Identification and Expression Analysis of Histone Modification Gene (HM) Family during Somatic Embryogenesis of Oil Palm. BMC Genom. 2022, 23, 11. [Google Scholar] [CrossRef] [PubMed]
- Fan, S.; Wang, J.; Lei, C.; Gao, C.; Yang, Y.; Li, Y.; An, N.; Zhang, D.; Han, M. Identification and Characterization of Histone Modification Gene Family Reveal Their Critical Responses to Flower Induction in Apple. BMC Plant Biol. 2018, 18, 173. [Google Scholar] [CrossRef]
- Aiese Cigliano, R.; Sanseverino, W.; Cremona, G.; Ercolano, M.R.; Conicella, C.; Consiglio, F.M. Genome-Wide Analysis of Histone Modifiers in Tomato: Gaining an Insight into Their Developmental Roles. BMC Genom. 2013, 14, 57. [Google Scholar] [CrossRef]
- Xu, J.; Xu, H.; Liu, Y.; Wang, X.; Xu, Q.; Deng, X. Genome-Wide Identification of Sweet Orange (Citrus Sinensis) Histone Modification Gene Families and Their Expression Analysis during the Fruit Development and Fruit-Blue Mold Infection Process. Front. Plant Sci. 2015, 6, 607. [Google Scholar] [CrossRef]
- Wang, L.; Ahmad, B.; Liang, C.; Shi, X.; Sun, R.; Zhang, S.; Du, G. Bioinformatics and Expression Analysis of Histone Modification Genes in Grapevine Predict Their Involvement in Seed Development, Powdery Mildew Resistance, and Hormonal Signaling. BMC Plant Biol. 2020, 20, 412. [Google Scholar] [CrossRef]
- Peng, M.; Ying, P.; Liu, X.; Li, C.; Xia, R.; Li, J.; Zhao, M. Genome-Wide Identification of Histone Modifiers and Their Expression Patterns during Fruit Abscission in Litchi. Front. Plant Sci. 2017, 8, 639. [Google Scholar] [CrossRef] [PubMed]
- Marand, A.P.; Eveland, A.L.; Kaufmann, K.; Springer, N.M. Cis-Regulatory Elements in Plant Development, Adaptation, and Evolution. Annu. Rev. Plant Biol. 2023, 74, 111–137. [Google Scholar] [CrossRef] [PubMed]
- Wittkopp, P.J.; Kalay, G. Cis-Regulatory Elements: Molecular Mechanisms and Evolutionary Processes Underlying Divergence. Nat. Rev. Genet. 2012, 13, 59–69. [Google Scholar] [CrossRef]
- Jamil, A.; Riaz, S.; Ashraf, M.; Foolad, M.R. Gene Expression Profiling of Plants under Salt Stress. Crit. Rev. Plant Sci. 2011, 30, 435–458. [Google Scholar] [CrossRef]
- Wang, D.; Pan, Y.; Zhao, X.; Zhu, L.; Fu, B.; Li, Z. Genome-Wide Temporal-Spatial Gene Expression Profiling of Drought Responsiveness in Rice. BMC Genom. 2011, 12, 149. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.; Yang, H.; Wei, Z.; Ma, H.; Ge, X. Rice Male Development under Drought Stress: Phenotypic Changes and Stage-Dependent Transcriptomic Reprogramming. Mol. Plant 2013, 6, 1630–1645. [Google Scholar] [CrossRef]
- Zhang, D.; Wilson, Z.A. Stamen Specification and Anther Development in Rice. Chin. Sci. Bull. 2009, 54, 2342–2353. [Google Scholar] [CrossRef]
- Zhang, D.; Luo, X.; Zhu, L. Cytological Analysis and Genetic Control of Rice Anther Development. J. Genet. Genom. 2011, 38, 379–390. [Google Scholar] [CrossRef]
- Deveshwar, P.; Bovill, W.D.; Sharma, R.; Able, J.A.; Kapoor, S. Analysis of Anther Transcriptomes to Identify Genes Contributing to Meiosis and Male Gametophyte Development in Rice. BMC Plant Biol. 2011, 11, 78. [Google Scholar] [CrossRef]
- Berr, A.; Shafiq, S.; Shen, W.-H. Histone Modifications in Transcriptional Activation during Plant Development. Biochim. Et Biophys. Acta (BBA)-Gene Regul. Mech. 2011, 1809, 567–576. [Google Scholar] [CrossRef]
- Magadum, S.; Banerjee, U.; Murugan, P.; Gangapur, D.; Ravikesavan, R. Gene Duplication as a Major Force in Evolution. J. Genet. 2013, 92, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Guo, F.; Qi, P.; Huang, Y.; Xie, Y.; Xu, L.; Han, N.; Xu, L.; Bian, H. OsHDA710-Mediated Histone Deacetylation Regulates Callus Formation of Rice Mature Embryo. Plant Cell Physiol. 2020, 61, 1646–1660. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Torres-Garcia, J.; Latrasse, D.; Benhamed, M.; Schilderink, S.; Zhou, W.; Kulikova, O.; Hirt, H.; Bisseling, T. Plant-Specific Histone Deacetylases HDT1/2 Regulate GIBBERELLIN 2-OXIDASE2 Expression to Control Arabidopsis Root Meristem Cell Number. Plant Cell 2017, 29, 2183–2196. [Google Scholar] [CrossRef] [PubMed]
- Colville, A.; Alhattab, R.; Hu, M.; Labbé, H.; Xing, T.; Miki, B. Role of HD2 Genes in Seed Germination and Early Seedling Growth in Arabidopsis. Plant Cell Rep. 2011, 30, 1969–1979. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.; Liu, X.; Singh, P.; Cui, Y.; Zimmerli, L.; Wu, K. Chromatin Modifications and Remodeling in Plant Abiotic Stress Responses. Biochim. Et Biophys. Acta (BBA)-Gene Regul. Mech. 2012, 1819, 129–136. [Google Scholar] [CrossRef]
- Feng, C.; Cai, X.; Su, Y.; Li, L.; Chen, S.; He, X. Arabidopsis RPD3-like Histone Deacetylases Form Multiple Complexes Involved in Stress Response. J. Genet. Genom. 2021, 48, 369–383. [Google Scholar] [CrossRef]
- Thorstensen, T.; Fischer, A.; Sandvik, S.V.; Johnsen, S.S.; Grini, P.E.; Reuter, G.; Aalen, R.B. The Arabidopsis SUVR4 Protein Is a Nucleolar Histone Methyltransferase with Preference for Monomethylated H3K9. Nucleic Acids Res. 2006, 34, 5461–5470. [Google Scholar] [CrossRef]
- Cartagena, J.A.; Matsunaga, S.; Seki, M.; Kurihara, D.; Yokoyama, M.; Shinozaki, K.; Fujimoto, S.; Azumi, Y.; Uchiyama, S.; Fukui, K. The Arabidopsis SDG4 Contributes to the Regulation of Pollen Tube Growth by Methylation of Histone H3 Lysines 4 and 36 in Mature Pollen. Dev. Biol. 2008, 315, 355–368. [Google Scholar] [CrossRef]
- Berr, A.; McCallum, E.J.; Ménard, R.; Meyer, D.; Fuchs, J.; Dong, A.; Shen, W.-H. Arabidopsis SET DOMAIN GROUP2 Is Required for H3K4 Trimethylation and Is Crucial for Both Sporophyte and Gametophyte Development. Plant Cell 2010, 22, 3232–3248. [Google Scholar] [CrossRef]
- Wei, F.; Tang, D.; Li, Z.; Kashif, M.H.; Khan, A.; Lu, H.; Jia, R.; Chen, P. Molecular Cloning and Subcellular Localization of Six HDACs and Their Roles in Response to Salt and Drought Stress in Kenaf (Hibiscus Cannabinus L.). Biol. Res. 2019, 52, 20. [Google Scholar] [CrossRef]
- Blanc, G.; Wolfe, K.H. Widespread Paleopolyploidy in Model Plant Species Inferred from Age Distributions of Duplicate Genes [W]. Plant Cell 2004, 16, 1667–1678. [Google Scholar] [CrossRef] [PubMed]






| Species | HAC | HAG | HAF | HAM | HDMA | JMJ | PRMT | SDG | HDT | HDA | SRT |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Rice | 3 | 26 | 1 | 2 | 9 | 14 | 8 | 47 | 2 | 16 | 2 |
| Arabidopsis | 5 | 3 | 2 | 2 | 4 | 20 | 7 | 41 | 4 | 12 | 2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, Y.; Liu, J.; Cheng, L.; Xu, D.; Liu, S.; Hu, H.; Ling, Y.; Yang, R.; Zhang, Y. Genome-Wide Analysis of the Histone Modification Gene (HM) Family and Expression Investigation during Anther Development in Rice (Oryza sativa L.). Plants 2024, 13, 2496. https://doi.org/10.3390/plants13172496
Huang Y, Liu J, Cheng L, Xu D, Liu S, Hu H, Ling Y, Yang R, Zhang Y. Genome-Wide Analysis of the Histone Modification Gene (HM) Family and Expression Investigation during Anther Development in Rice (Oryza sativa L.). Plants. 2024; 13(17):2496. https://doi.org/10.3390/plants13172496
Chicago/Turabian StyleHuang, Yongxiang, Jiawei Liu, Long Cheng, Duo Xu, Sijia Liu, Hanqiao Hu, Yu Ling, Rongchao Yang, and Yueqin Zhang. 2024. "Genome-Wide Analysis of the Histone Modification Gene (HM) Family and Expression Investigation during Anther Development in Rice (Oryza sativa L.)" Plants 13, no. 17: 2496. https://doi.org/10.3390/plants13172496






