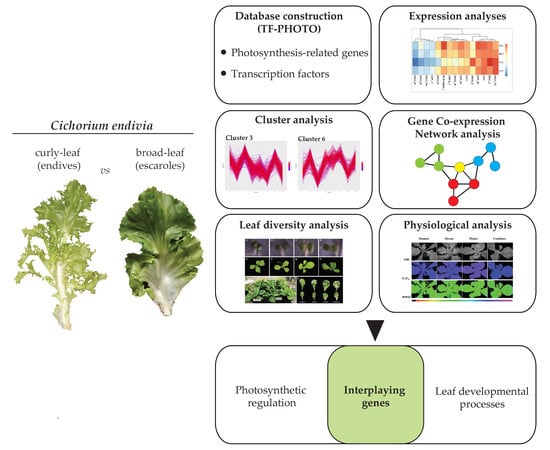
Transcription Factor Networks in Leaves of Cichorium endivia: New Insights into the Relationship between Photosynthesis and Leaf Development
Abstract
:1. Introduction
2. Results
2.1. Construction of TF-PHOTO, a C. Endivia Leaf Database for Transcription Factors and Photosynthesis-Related Genes
2.2. Finding Expression Patterns in Leaves Using K-Means Cluster Analysis
2.3. Analysis of Gene Co-expression Networks of C. endivia Leaves
2.4. Interplay Between Shape and Function: Photosynthetic Parameters in Relation to Morphological Leaf Diversity
2.5. Interplay Between Development and Function: Developmental Genes Related to the Photosynthetic Regulatory Pathway
2.6. TF-PHOTO Genes Differentially Expressed in Broad Versus Curly Leaves
3. Discussion
3.1. The Regulatory Network of Photosynthesis in the Leaves of C. endivia
3.2. Developmental Genes in Cluster Three Linked to Positive Regulation of Photosynthesis
3.3. Developmental Genes in Cluster Six Antagonistic to Photosynthesis
3.4. Transcription Factors that Are Differentially Expressed Between Broad- and Curly-Leaved C. endivia Cultivars
4. Materials and Methods
4.1. Biological Material and Morphological Analysis
4.2. RNA Isolation, Sequencing, and Expression Analysis
4.3. TF-PHOTO Database Construction
4.4. K-Means Cluster Analysis and Gene Co-Expression Network Construction
4.5. Chlorophyll Fluorescence Measurements
4.6. Determination of Chlorophyll and Total Carotenoids Content
4.7. Measurements of Spectral Reflectance and Spectral Reflectance Indices
4.8. Measurement of Lipid Peroxidation Levels
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ren, T.; Weraduwage, S.M.; Sharkey, T.D. Prospects for enhancing leaf photosynthetic capacity by manipulating mesophyll cell morphology. J. Exp. Bot. 2019, 70, 1153–1165. [Google Scholar] [CrossRef]
- Fritz, M.A.; Rosa, S.; Sicard, A. Mechanisms underlying the environmentally induced plasticity of leaf morphology. Front. Genet. 2018, 9, 478. [Google Scholar] [CrossRef] [PubMed]
- Tsukaya, H. A consideration of leaf shape evolution in the context of the primary function of the leaf as a photosynthetic organ. In The Leaf: A Platform for Performing Photosynthesis; Adams, W.W., III, Terashima, I., Eds.; Springer International Publishing: Cham, Switzerland, 2018; pp. 1–26. [Google Scholar]
- Eberhard, S.; Finazzi, G.; Wollman, F.A. The dynamics of photosynthesis. Annu. Rev. Genet. 2008, 42, 463–515. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.H. Reactive oxygen species, oxidative signaling and the regulation of photosynthesis. Env. Exp. Bot. 2018, 154, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Friso, G.; Giacomelli, L.; Ytterberg, A.J.; Peltier, J.B.; Rudella, A.; Sun, Q.; Wijk, K.J. In-Depth analysis of the thylakoid membrane proteome of Arabidopsis thaliana chloroplasts: New proteins, new functions, and a plastid proteome database. Plant Cell 2004, 16, 478–499. [Google Scholar] [CrossRef]
- Martin, W.; Rujan, T.; Richly, E.; Hansen, A.; Cornelsen, S.; Lins, T.; Leister, D.; Stoebe, B.; Hasegawa, M.; Penny, D. Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc. Natl. Acad. Sci. USA 2002, 99, 12246–12251. [Google Scholar] [CrossRef]
- Wang, P.; Hendron, R.W.; Kelly, S. Transcriptional control of photosynthetic capacity: Conservation and divergence from Arabidopsis to rice. New Phytol. 2017, 216, 32–45. [Google Scholar] [CrossRef]
- Smith, H. Phytochromes and light signal perception by plants-an emerging synthesis. Nature 2000, 407, 585–591. [Google Scholar] [CrossRef]
- Schepens, I.; Duek, P.; Fankhauser, C. Phytochrome-mediated light signalling in Arabidopsis. Curr. Opin. Plant Biol. 2004, 7, 564–569. [Google Scholar] [CrossRef]
- Rockwell, N.C.; Su, Y.S.; Lagarias, J.C. Phytochrome structure and signaling mechanisms. Annu. Rev. Plant Biol. 2006, 57, 837–858. [Google Scholar] [CrossRef]
- Quail, P.H. Phytochrome photosensory signalling networks. Nat. Rev. Mol. Cell Biol. 2002, 3, 85–93. [Google Scholar] [CrossRef] [PubMed]
- Franklin, K.A.; Quail, P.H. Phytochrome functions in Arabidopsis development. J. Exp. Bot. 2010, 61, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Yang, E.J.; Yoo, C.Y.; Liu, J.; Wang, H.; Cao, J.; Li, F.W.; Pryer, K.M.; Sun, T.P.; Weigel, D.; Zhou, P.; et al. NCP activates chloroplast transcription by controlling phytochrome-dependent dual nuclear and plastidial switches. Nat. Commun. 2019, 10, 2630. [Google Scholar] [CrossRef] [PubMed]
- Lau, O.S.; Deng, X.W. Plant hormone signaling lightens up: Integrators of light and hormones. Curr. Opin. Plant Biol. 2010, 13, 571–577. [Google Scholar] [CrossRef] [PubMed]
- Toledo-Ortiz, G.; Huq, E.; Rodriguez-Concepcion, M. Direct regulation of phytoene synthase gene expression and carotenoid biosynthesis by phytochrome-interacting factors. Proc. Natl. Acad. Sci. USA 2010, 107, 11626–11631. [Google Scholar] [CrossRef]
- Leivar, P.; Quail, P.H. PIFs: Pivotal components in a cellular signaling hub. Trends Plant Sci. 2011, 16, 19–28. [Google Scholar] [CrossRef]
- Martin, G.; Leivar, P.; Ludevid, D.; Tepperman, J.M.; Quail, P.H.; Monte, E. Phytochrome and retrograde signalling pathways converge to antagonistically regulate a light-induced transcriptional network. Nat. Commun. 2016, 7, 11431. [Google Scholar] [CrossRef]
- Wang, L.; Gu, X.; Xu, D.; Wang, W.; Wang, H.; Zeng, M.; Chang, Z.; Huang, H.; Cui, X. miR396-targeted AtGRF transcription factors are required for coordination of cell division and differentiation during leaf development in Arabidopsis. J. Exp. Bot. 2011, 62, 761–773. [Google Scholar] [CrossRef]
- Gollan, P.J.; Tikkanen, M.; Aro, E.M. Photosynthetic light reactions: Integral to chloroplast retrograde signalling. Curr. Opin. Plant Biol. 2015, 27, 180–191. [Google Scholar] [CrossRef]
- Chi, W.; Feng, P.; Ma, J.; Zhang, L. Metabolites and chloroplast retrograde signaling. Curr. Opin. Plant Biol. 2015, 25, 32–38. [Google Scholar] [CrossRef]
- Agrawal, G.K.; Yamazaki, M.; Kobayashi, M.; Hirochika, R.; Miyao, A.; Hirochika, H. Screening of the rice viviparous mutants generated by endogenous retrotransposon Tos17 insertion. Tagging of a zeaxanthin epoxidase gene and a novel ostatc gene. Plant Physiol. 2001, 125, 1248–1257. [Google Scholar] [CrossRef] [PubMed]
- Nicotra, A.B.; Cosgrove, M.J.; Cowling, A.; Schlichting, C.D.; Jones, C.S. Leaf shape linked to photosynthetic rates and temperature optima in South African Pelargonium species. Oecologia 2008, 154, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Strauss, S.; Lempe, J.; Prusinkiewicz, P.; Tsiantis, M.; Smith, R.S. Phyllotaxis: Is the golden angle optimal for light capture? New Phytol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Testone, G.; Mele, G.; di Giacomo, E.; Tenore, G.C.; Gonnella, M.; Nicolodi, C.; Frugis, G.; Iannelli, M.A.; Arnesi, G.; Schiappa, A.; et al. Transcriptome driven characterization of curly- and smooth-leafed endives reveals molecular differences in the sesquiterpenoid pathway. Hortic. Res. 2019, 6, 1. [Google Scholar] [CrossRef]
- Jin, J.; Zhang, H.; Kong, L.; Gao, G.; Luo, J. PlantTFDB 3.0: A portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, D1182–D1187. [Google Scholar] [CrossRef]
- Meisrimler, C.N.; Pelgrom, A.J.E.; Oud, B.; Out, S.; Van den Ackerveken, G. Multiple downy mildew effectors target the stress-related NAC transcription factor LsNAC069 in lettuce. Plant J. 2019, 99, 1098–1115. [Google Scholar] [CrossRef]
- Reyes-Chin-Wo, S.; Wang, Z.; Yang, X.; Kozik, A.; Arikit, S.; Song, C.; Xia, L.; Froenicke, L.; Lavelle, D.O.; Truco, M.J.; et al. Genome assembly with in vitro proximity ligation data and whole-genome triplication in lettuce. Nat. Commun. 2017, 8, 14953. [Google Scholar] [CrossRef]
- Thimm, O.; Blasing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Kruger, P.; Selbig, J.; Muller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- Bryan, J. Problems in gene clustering based on gene expression data. J. Multivar. Anal. 2004, 90, 44–66. [Google Scholar] [CrossRef]
- Altman, N.S.; Krzywinski, M. Points of significance: Clustering. Nat. Methods 2017, 14, 545–546. [Google Scholar] [CrossRef]
- Huq, E.; Al-Sady, B.; Hudson, M.; Kim, C.; Apel, K.; Quail, P.H. Phytochrome-interacting factor 1 is a critical bHLH regulator of chlorophyll biosynthesis. Science 2004, 305, 1937–1941. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.X.; Thomas, C.E.; Brunak, S. Network biology concepts in complex disease comorbidities. Nat. Rev. Genet. 2016, 17, 615–629. [Google Scholar] [CrossRef] [PubMed]
- Argyros, R.D.; Mathews, D.E.; Chiang, Y.H.; Palmer, C.M.; Thibault, D.M.; Etheridge, N.; Argyros, D.A.; Mason, M.G.; Kieber, J.J.; Schaller, G.E. Type B response regulators of Arabidopsis play key roles in cytokinin signaling and plant development. Plant Cell 2008, 20, 2102–2116. [Google Scholar] [CrossRef] [Green Version]
- De la Riva, E.G.; Olmo, M.; Poorter, H.; Ubera, J.L.; Villar, R. Leaf mass per area (LMA) and its relationship with leaf structure and anatomy in 34 mediterranean woody species along a water availability gradient. PLoS ONE 2016, 11, e0148788. [Google Scholar] [CrossRef] [Green Version]
- Baker, N.R. Chlorophyll fluorescence: A probe of photosynthesis in vivo. Annu. Rev. Plant Biol. 2008, 59, 89–113. [Google Scholar] [CrossRef] [Green Version]
- Harbinson, J.; Genty, B.; Baker, N.R. Relationship between the quantum efficiencies of photosystems I and II in pea leaves. Plant Physiol. 1989, 90, 1029–1034. [Google Scholar] [CrossRef] [Green Version]
- Penuelas, J.; Garbulsky, M.F.; Filella, I. Photochemical reflectance index (PRI) and remote sensing of plant CO2 uptake. New Phytol. 2011, 191, 596–599. [Google Scholar] [CrossRef]
- Schmitz, C.; Ramlov, F.; de Lucena, L.A.F.; Uarrota, V.; Batista, M.B.; Sissini, M.N.; Oliveira, I.; Briani, B.; Martins, C.D.L.; Nunes, J.M.C.; et al. UVR and PAR absorbing compounds of marine brown macroalgae along a latitudinal gradient of the Brazilian coast. J. Photochem. Photobiol. B 2018, 178, 165–174. [Google Scholar] [CrossRef]
- Morales, M.; Munne-Bosch, S. Malondialdehyde: Facts and artifacts. Plant Physiol. 2019, 180, 1246–1250. [Google Scholar] [CrossRef] [Green Version]
- Chan, K.X.; Phua, S.Y.; Crisp, P.; McQuinn, R.; Pogson, B.J. Learning the languages of the chloroplast: Retrograde signaling and beyond. Annu. Rev. Plant Biol. 2016, 67, 25–53. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.; Yamaguchi, S.; Kamiya, Y.; Bae, G.; Chung, W.I.; Choi, G. Light activates the degradation of PIL5 protein to promote seed germination through gibberellin in Arabidopsis. Plant J. 2006, 47, 124–139. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.X.; Srivastava, R.; Howell, S.H. Stress-induced expression of an activated form of AtbZIP17 provides protection from salt stress in Arabidopsis. Plant Cell Env. 2008, 31, 1735–1743. [Google Scholar] [CrossRef] [PubMed]
- Choudhury, F.K.; Rivero, R.M.; Blumwald, E.; Mittler, R. Reactive oxygen species, abiotic stress and stress combination. Plant J. 2017, 90, 856–867. [Google Scholar] [CrossRef]
- Serin, E.A.; Nijveen, H.; Hilhorst, H.W.; Ligterink, W. Learning from co-expression networks: Possibilities and challenges. Front. Plant Sci. 2016, 7, 444. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, S.; Gao, J.; Zhu, X.; Song, Y.; Li, Z.; Ren, G.; Zhou, X.; Kuai, B. ABF2, ABF3, and ABF4 promote ABA-mediated chlorophyll degradation and leaf senescence by transcriptional activation of chlorophyll catabolic genes and senescence-associated genes in Arabidopsis. Mol. Plant 2016, 9, 1272–1285. [Google Scholar] [CrossRef] [Green Version]
- Xu, Z.S.; Chen, M.; Li, L.C.; Ma, Y.Z. Functions and application of the AP2/ERF transcription factor family in crop improvement. J. Integr. Plant Biol. 2011, 53, 570–585. [Google Scholar] [CrossRef]
- Karaba, A.; Dixit, S.; Greco, R.; Aharoni, A.; Trijatmiko, K.R.; Marsch-Martinez, N.; Krishnan, A.; Nataraja, K.N.; Udayakumar, M.; Pereira, A. Improvement of water use efficiency in rice by expression of HARDY, an Arabidopsis drought and salt tolerance gene. Proc. Natl. Acad. Sci. USA 2007, 104, 15270–15275. [Google Scholar] [CrossRef] [Green Version]
- Mawlong, I.; Ali, K.; Srinivasan, R.; Rai, R.D.; Tyagi, A. Functional validation of a drought-responsive AP2/ERF family transcription factor-encoding gene from rice in Arabidopsis. Mol. Breed. 2015, 35, 163. [Google Scholar] [CrossRef]
- Sharabi-Schwager, M.; Lers, A.; Samach, A.; Guy, C.L.; Porat, R. Overexpression of the CBF2 transcriptional activator in Arabidopsis delays leaf senescence and extends plant longevity. J. Exp. Bot. 2010, 61, 261–273. [Google Scholar] [CrossRef] [Green Version]
- Koyama, T.; Nii, H.; Mitsuda, N.; Ohta, M.; Kitajima, S.; Ohme-Takagi, M.; Sato, F. A regulatory cascade involving class II ethylene response factor transcriptional repressors operates in the progression of leaf senescence. Plant Physiol. 2013, 162, 991–1005. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, X.L.; Xia, X.J.; Kuang, S.; Zhang, X.L.; Yin, X.R.; Yu, J.Q.; Chen, K.S. A novel ethylene responsive factor CitERF13 plays a role in photosynthesis regulation. Plant Sci. 2017, 256, 112–119. [Google Scholar] [CrossRef] [PubMed]
- Baldoni, E.; Genga, A.; Cominelli, E. Plant MYB transcription factors: Their role in drought response mechanisms. Int. J. Mol. Sci. 2015, 16, 15811–15851. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chezem, W.R.; Clay, N.K. Regulation of plant secondary metabolism and associated specialized cell development by MYBs and bHLHs. Phytochemistry 2016, 131, 26–43. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Kim, W.-Y.; Ali, Z.; Park, H.J.; Park, S.J.; Cha, J.-Y.; Perez-Hormaeche, J.; Quintero, F.J.; Shin, G.; Kim, M.R.; Qiang, Z.; et al. Release of SOS2 kinase from sequestration with Gigantea determines salt tolerance in Arabidopsis. Nat. Commun. 2013, 4, 1352. [Google Scholar] [CrossRef] [Green Version]
- Jahns, P.; Latowski, D.; Strzalka, K. Mechanism and regulation of the violaxanthin cycle: The role of antenna proteins and membrane lipids. Biochim. Biophys. Acta 2009, 1787, 3–14. [Google Scholar] [CrossRef] [Green Version]
- Hirsch, S.; Oldroyd, G.E. GRAS-domain transcription factors that regulate plant development. Plant Signal Behav. 2009, 4, 698–700. [Google Scholar] [CrossRef] [Green Version]
- Engstrom, E.M.; Andersen, C.M.; Gumulak-Smith, J.; Hu, J.; Orlova, E.; Sozzani, R.; Bowman, J.L. Arabidopsis homologs of the petunia hairy meristem gene are required for maintenance of shoot and root indeterminacy. Plant Physiol. 2011, 155, 735–750. [Google Scholar] [CrossRef] [Green Version]
- Ma, Z.; Hu, X.; Cai, W.; Huang, W.; Zhou, X.; Luo, Q.; Yang, H.; Wang, J.; Huang, J. Arabidopsis miR171-targeted scarecrow-like proteins bind to GT cis-elements and mediate gibberellin-regulated chlorophyll biosynthesis under light conditions. PLoS Genet. 2014, 10, e1004519. [Google Scholar] [CrossRef] [Green Version]
- Vercruyssen, L.; Tognetti, V.B.; Gonzalez, N.; Van Dingenen, J.; De Milde, L.; Bielach, A.; De Rycke, R.; Van Breusegem, F.; Inze, D. GROWTH REGULATING FACTOR5 stimulates Arabidopsis chloroplast division, photosynthesis, and leaf longevity. Plant Physiol. 2015, 167, 817–832. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, Y.; Xu, X.; Gong, Z.; Tang, Y.; Wu, M.; Yan, F.; Zhang, X.; Zhang, Q.; Yang, F.; Hu, X.; et al. Auxin response factor 6A regulates photosynthesis, sugar accumulation, and fruit development in tomato. Hortic. Res. 2019, 6, 85. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Z.; Wang, F.; Hong, Y.; Yao, J.; Ren, Z.; Shi, H.; Zhu, J.K. The flowering repressor SVP confers drought resistance in Arabidopsis by regulating abscisic acid catabolism. Mol. Plant 2018, 11, 1184–1197. [Google Scholar] [CrossRef] [Green Version]
- Danisman, S. TCP transcription factors at the interface between environmental challenges and the plant’s growth responses. Front. Plant Sci. 2016, 7, 1930. [Google Scholar] [CrossRef] [PubMed]
- Herve, C.; Dabos, P.; Bardet, C.; Jauneau, A.; Auriac, M.C.; Ramboer, A.; Lacout, F.; Tremousaygue, D. In vivo interference with AtTCP20 function induces severe plant growth alterations and deregulates the expression of many genes important for development. Plant Physiol. 2009, 149, 1462–1477. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andriankaja, M.E.; Danisman, S.; Mignolet-Spruyt, L.F.; Claeys, H.; Kochanke, I.; Vermeersch, M.; De Milde, L.; De Bodt, S.; Storme, V.; Skirycz, A.; et al. Transcriptional coordination between leaf cell differentiation and chloroplast development established by TCP20 and the subgroup Ib bHLH transcription factors. Plant Mol. Biol. 2014, 85, 233–245. [Google Scholar] [CrossRef]
- Van Es, S.W.; van der Auweraert, E.B.; Silveira, S.R.; Angenent, G.C.; van Dijk, A.D.J.; Immink, R.G.H. Comprehensive phenotyping reveals interactions and functions of Arabidopsis thaliana TCP genes in yield determination. Plant J. 2019, 99, 316–328. [Google Scholar]
- Danisman, S.; van der Wal, F.; Dhondt, S.; Waites, R.; de Folter, S.; Bimbo, A.; van Dijk, A.D.; Muino, J.M.; Cutri, L.; Dornelas, M.C.; et al. Arabidopsis class I and class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically. Plant Physiol. 2012, 159, 1511–1523. [Google Scholar] [CrossRef] [Green Version]
- Baima, S.; Possenti, M.; Matteucci, A.; Wisman, E.; Altamura, M.M.; Ruberti, I.; Morelli, G. The Arabidopsis ATHB-8 HD-zip protein acts as a differentiation-promoting transcription factor of the vascular meristems. Plant Physiol. 2001, 126, 643–655. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.Y.; Shen, A.; Xiong, W.; Sun, Q.L.; Luo, Q.; Song, T.; Li, Z.L.; Luan, W.J. Overexpression of OsHox32 results in pleiotropic effects on plant type architecture and leaf development in rice. Rice 2016, 9, 46. [Google Scholar] [CrossRef] [Green Version]
- Galvez-Valdivieso, G.; Mullineaux, P.M. The role of reactive oxygen species in signalling from chloroplasts to the nucleus. Physiol. Plant 2010, 138, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Strasser, R. Protein quality control in the endoplasmic reticulum of plants. Annu. Rev. Plant Biol. 2018, 69, 147–172. [Google Scholar] [CrossRef] [PubMed]
- Ruckle, M.E.; DeMarco, S.M.; Larkin, R.M. Plastid signals remodel light signaling networks and are essential for efficient chloroplast biogenesis in Arabidopsis. Plant Cell 2007, 19, 3944–3960. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Andriankaja, M.; Dhondt, S.; De Bodt, S.; Vanhaeren, H.; Coppens, F.; De Milde, L.; Muhlenbock, P.; Skirycz, A.; Gonzalez, N.; Beemster, G.T.; et al. Exit from proliferation during leaf development in Arabidopsis thaliana: A not-so-gradual process. Dev. Cell 2012, 22, 64–78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gordon, M.J.; Carmody, M.; Albrecht, V.; Pogson, B. Systemic and local responses to repeated hl stress-induced retrograde signaling in Arabidopsis. Front. Plant Sci. 2012, 3, 303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogel, M.O.; Moore, M.; König, K.; Pecher, P.; Alsharafa, K.; Lee, J.; Dietz, K.J. Fast retrograde signaling in response to high light involves metabolite export, mitogen-activated protein KINASE6, and AP2/ERF transcription factors in Arabidopsis. Plant Cell 2014, 26, 1151. [Google Scholar] [CrossRef] [Green Version]
- Rossel, J.B.; Wilson, P.B.; Hussain, D.; Woo, N.S.; Gordon, M.J.; Mewett, O.P.; Howell, K.A.; Whelan, J.; Kazan, K.; Pogson, B.J. Systemic and intracellular responses to photooxidative stress in Arabidopsis. Plant Cell 2007, 19, 4091–4110. [Google Scholar] [CrossRef] [Green Version]
- Iida, A.; Kazuoka, T.; Torikai, S.; Kikuchi, H.; Oeda, K. A zinc finger protein RHL41 mediates the light acclimatization response in Arabidopsis. Plant J. 2000, 24, 191–203. [Google Scholar] [CrossRef]
- Vermeirssen, V.; De Clercq, I.; Van Parys, T.; Van Breusegem, F.; Van de Peer, Y. Arabidopsis ensemble reverse-engineered gene regulatory network discloses interconnected transcription factors in oxidative stress. Plant Cell 2014, 26, 4656–4679. [Google Scholar] [CrossRef]
- Seo, P.J.; Park, C.M. Signaling linkage between environmental stress resistance and leaf senescence in Arabidopsis. Plant Signal. Behav. 2011, 6, 1564–1566. [Google Scholar] [CrossRef] [Green Version]
- Zhou, X.; Jiang, Y.; Yu, D. WRKY22 transcription factor mediates dark-induced leaf senescence in Arabidopsis. Mol. Cells 2011, 31, 303–313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zentgraf, U.; Laun, T.; Miao, Y. The complex regulation of WRKY53 during leaf senescence of Arabidopsis thaliana. Eur. J. Cell Biol. 2010, 89, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Miao, Y.; Jiang, J.; Ren, Y.; Zhao, Z. The single-stranded DNA-binding protein WHIRLY1 represses WRKY53 expression and delays leaf senescence in a developmental stage-dependent manner in Arabidopsis. Plant Physiol. 2013, 163, 746–756. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, Y.; Huhn, K.; Brandt, R.; Potschin, M.; Bieker, S.; Straub, D.; Doll, J.; Drechsler, T.; Zentgraf, U.; Wenkel, S. Revoluta and WRKY53 connect early and late leaf development in Arabidopsis. Development 2014, 141, 4772–4783. [Google Scholar] [CrossRef] [Green Version]
- Denekamp, M.; Smeekens, S.C. Integration of wounding and osmotic stress signals determines the expression of the AtMYB102 transcription factor gene. Plant Physiol. 2003, 132, 1415–1423. [Google Scholar] [CrossRef] [Green Version]
- Efroni, I.; Han, S.K.; Kim, H.J.; Wu, M.F.; Steiner, E.; Birnbaum, K.D.; Hong, J.C.; Eshed, Y.; Wagner, D. Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses. Dev. Cell 2013, 24, 438–445. [Google Scholar] [CrossRef] [Green Version]
- Di Giacomo, E.; Iannelli, M.A.; Frugis, G. TALE and Shape: How to Make a Leaf Different. Plants 2013, 2, 317–342. [Google Scholar] [CrossRef] [Green Version]
- Bar, M.; Ori, N. Compound leaf development in model plant species. Curr. Opin. Plant Biol. 2015, 23, 61–69. [Google Scholar] [CrossRef]
- Ung, N.; Lal, S.; Smith, H.M. The role of pennywise and pound-foolish in the maintenance of the shoot apical meristem in Arabidopsis. Plant Physiol. 2011, 156, 605–614. [Google Scholar] [CrossRef] [Green Version]
- Lin, W.C.; Shuai, B.; Springer, P.S. The Arabidopsis lateral organ boundaries-domain gene ASYMMETRIC LEAVES2 functions in the repression of KNOX gene expression and in adaxial-abaxial patterning. Plant Cell 2003, 15, 2241–2252. [Google Scholar] [CrossRef] [Green Version]
- Nadakuduti, S.S.; Holdsworth, W.L.; Klein, C.L.; Barry, C.S. KNOX genes influence a gradient of fruit chloroplast development through regulation of GOLDEN2-LIKE expression in tomato. Plant J. 2014, 78, 1022–1033. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Fan, Z.; Zhang, Q.; Wang, C.; Gao, Y.; Deng, Y.; Zhu, B.; Zhu, H.; Chen, J.; Shan, W.; et al. BEL1-LIKE HOMEODOMAIN 11 regulates chloroplast development and chlorophyll synthesis in tomato fruit. Plant J. 2018, 94, 1126–1140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, T.; Tan, W.; Yang, H.; Zhang, L.; Li, T.; Liu, B.; Zhang, D.; Lin, H. Regulation of anthocyanin accumulation via MYB75/HAT1/TPL-mediated transcriptional repression. PLoS Genet. 2019, 15, e1007993. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Jia, K.P.; Lian, H.L.; Yang, X.; Li, L.; Yang, H.Q. Jasmonic acid enhancement of anthocyanin accumulation is dependent on phytochrome: A signaling pathway under far-red light in Arabidopsis. Biochem. Biophys. Res. Commun. 2014, 454, 78–83. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, A.; Mansfield, S.D.; Hall, H.C.; Douglas, C.J.; Ellis, B.E. MYB75 functions in regulation of secondary cell wall formation in the Arabidopsis inflorescence stem. Plant Physiol. 2010, 154, 1428–1438. [Google Scholar] [CrossRef] [Green Version]
- Bui, L.T.; Giuntoli, B.; Kosmacz, M.; Parlanti, S.; Licausi, F. Constitutively expressed ERF-VII transcription factors redundantly activate the core anaerobic response in Arabidopsis thaliana. Plant Sci. 2015, 236, 37–43. [Google Scholar] [CrossRef]
- Kosmacz, M.; Parlanti, S.; Schwarzlander, M.; Kragler, F.; Licausi, F.; Van Dongen, J.T. The stability and nuclear localization of the transcription factor RAP2.12 are dynamically regulated by oxygen concentration. Plant Cell Env. 2015, 38, 1094–1103. [Google Scholar] [CrossRef]
- Briggs, G.C.; Mouchel, C.F.; Hardtke, C.S. Characterization of the plant-specific BREVIS RADIX gene family reveals limited genetic redundancy despite high sequence conservation. Plant Physiol. 2006, 140, 1306–1316. [Google Scholar] [CrossRef] [Green Version]
- Maugarny-Cales, A.; Cortizo, M.; Adroher, B.; Borrega, N.; Goncalves, B.; Brunoud, G.; Vernoux, T.; Arnaud, N.; Laufs, P. Dissecting the pathways coordinating patterning and growth by plant boundary domains. PLoS Genet. 2019, 15, e1007913. [Google Scholar] [CrossRef]
- Kierzkowski, D.; Runions, A.; Vuolo, F.; Strauss, S.; Lymbouridou, R.; Routier-Kierzkowska, A.L.; Wilson-Sanchez, D.; Jenke, H.; Galinha, C.; Mosca, G.; et al. A growth-based framework for leaf shape development and diversity. Cell 2019, 177, 1405–1418. [Google Scholar] [CrossRef]
- Bray, N.L.; Pimentel, H.; Melsted, P.; Pachter, L. Near-Optimal probabilistic RNA-seq quantification. Nat. Biotechnol 2016, 34, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3--new capabilities and interfaces. Nucleic Acids Res 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maroufi, A.; Van Bockstaele, E.; De Loose, M. Validation of reference genes for gene expression analysis in chicory (Cichorium intybus) using quantitative real-time PCR. BMC Mol. Biol. 2010, 11, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muller, P.Y.; Janovjak, H.; Miserez, A.R.; Dobbie, Z. Processing of gene expression data generated by quantitative real-time RT-PCR. Biotechniques 2002, 32, 1372–1379. [Google Scholar] [PubMed]
- Lohse, M.; Nagel, A.; Herter, T.; May, P.; Schroda, M.; Zrenner, R.; Tohge, T.; Fernie, A.R.; Stitt, M.; Usadel, B. Mercator: A fast and simple web server for genome scale functional annotation of plant sequence data. Plant Cell Env. 2014, 37, 1250–1258. [Google Scholar] [CrossRef] [Green Version]
- Langfelder, P.; Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinf. 2008, 9, 559. [Google Scholar] [CrossRef] [Green Version]
- Wu, T.D.; Watanabe, C.K. GMAP: A genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 2005, 21, 1859–1875. [Google Scholar] [CrossRef] [Green Version]
- Wei, T.Y.; Simko, V. Visualization of a Correlation Matrix: Package ‘corrplot’. 2017. Available online: https://rdrr.io/cran/corrplot/man/corrplot-package.html (accessed on 22 October 2019).
- Maxwell, K.; Johnson, G.N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 2000, 51, 659–668. [Google Scholar] [CrossRef]
- Cerovic, Z.G.; Masdoumier, G.; Ghozlen, N.B.; Latouche, G. A new optical leaf-clip meter for simultaneous non-destructive assessment of leaf chlorophyll and epidermal flavonoids. Physiol. Plant 2012, 146, 251–260. [Google Scholar] [CrossRef]
- Sumanta, N.; Haque, C.I.; Nishika, J.; Suprakash, R. Spectrophotometric analysis of chlorophylls and carotenoids from commonly grown fern species by using various extracting solvents. Res. J. Chem. Sci. 2014, 4, 63–69. [Google Scholar]
- Castro-Esau, K.L.; Sánchez-Azofeifa, G.A.; Rivard, B. Comparison of spectral indices obtained using multiple spectroradiometers. Remote Sens. Env. 2006, 103, 276288. [Google Scholar]
- Hodges, D.M.; DeLong, J.M.; Forney, C.F.; Prange, R.K. Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds. Planta 1999, 207, 604–611. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]









| Cultivar | Parameters1 | |||||
|---|---|---|---|---|---|---|
| Fv/Fm (Relative Units) | Φ PSII (Relative Units) | ETR (μmol elec. m−2 s−1) | Abs (Relative Units) | Chlorophyll Content (µg cm−2) | PRI (Relative Units) | |
| Domari | 0.793 ± 0.001 c | 0.405 ± 0.004 b | 55.73 ± 0.53 b | 0.746 ± 0.006 b | 19.87 ± 0.96 b | 0.043 ± 0.001 a |
| Myrna | 0.807 ± 0.001 b | 0.375 ± 0.003 c | 49.42 ± 0.47 c | 0.715 ± 0.002 c | 17.25 ± 1.98 bc | 0.038 ± 0.001 ab |
| Flester | 0.828 ± 0.001 a | 0.375 ± 0.003 c | 46.64 ± 0.47 d | 0.675 ± 0.009 d | 14.51 ± 1.01 c | 0.033 ± 0.001 b |
| Confiance | 0.781 ± 0.001 d | 0.419 ± 0.005 a | 60.16 ± 0.76 a | 0.778 ± 0.002 a | 23.57 ± 0.58 a | 0.042 ± 0.001 a |
| P | 0.001 | 0.001 | 0.001 | 0.003 | 0.002 | 0.005 |
| Ce ID | Category | Ath ID | Gene Symbol | Score to Core | K | Ce Gene Name |
|---|---|---|---|---|---|---|
| Ce_contig37589 | TF | AT4G30080 | AUXIN RESPONSE FACTOR 16 (ARF16) | 0.925 | 3 | ARF_11 |
| Ce_contig68433 | TF | AT2G25180 | RESPONSE REGULATOR 12 (RR12) | 0.964 | 3 | ARR-B_7 |
| Ce_contig18469 | TF | AT1G14920 | GIBBERELLIC ACID INSENSITIVE (GAI) | 0.936 | 3 | GRAS_16 |
| Ce_contig31926 | TF | AT5G17490 | RGA-LIKE PROTEIN 3 (RGL3) | 0.972 | 3 | GRAS_23 |
| Ce_contig82634 | TF | AT4G00150 | LOST MERISTEMS 3 (LOM3/SCL6-IV) | 0.918 | 3 | GRAS_43 |
| Ce_contig6575 | TF | AT4G00150 | LOST MERISTEMS 3 (LOM3/SCL6-IV) | 0.948 | 3 | GRAS_7 |
| Ce_contig71598 | TF | AT3G13960 | GROWTH-REGULATING FACTOR 5 (GRF5) | 0.941 | 3 | GRF_8 |
| Ce_contig83059 | TF | AT4G32880 | ATHB-8 | 0.933 | 3 | HD-ZIP_41 |
| Ce_contig21296 | TF | AT2G22540 | SHORT VEGETATIVE PHASE (SVP) | 0.938 | 3 | MIKC-MADS_10 |
| Ce_contig36777 | TF | AT5G65060 | MADS AFFECTING FLOWERING 3 (MAF3) | 0.973 | 3 | MIKC-MADS_13 |
| Ce_contig30145 | TF | AT5G60440 | AGAMOUS-LIKE 62 (AGL62) | 0.936 | 3 | Mtype_MADS_10 |
| Ce_contig1183 | TF | AT3G61250 | MYB DOMAIN PROTEIN 17 (MYB17) | 0.898 | 3 | MYB_2 |
| Ce_contig13510 | PHOTO | AT2G18790 | PHYTOCHROME B (PHYB) | 0.928 | 3 | PHYB |
| Ce_contig73618 | TF | AT2G33810 | SQUAMOSA PROMOTER BINDING PROTEIN-LIKE 3 (SPL3) | 0.899 | 3 | SBP_14 |
| Ce_contig85637 | TF | AT2G36990 | SIGMA FACTOR 6 (SIG6) | 0.955 | 3 | SIG_5 |
| Ce_contig49086 | TF | AT3G27010 | TEOSINTE BRANCHED 1, CYCLOIDEA, PCF (TCP) 20 (TCP20) | 0.908 | 3 | TCP_12 |
| Ce_contig33120 | TF | AT3G47620 | TEOSINTE BRANCHED, CYCLOIDEA, PCF (TCP) 14 (TCP14) | 0.986 | 3 | TCP_7 |
| Ce_contig38256 | TF | AT1G19850 | MONOPTEROS (MP) | 0.947 | 6 | ARF_12 |
| Ce_contig18795 | TF | AT1G27730 | SALT TOLERANCE ZINC FINGER (STZ) (ZAT10) | 0.985 | 6 | C2H2_22 |
| Ce_contig46000 | TF | AT1G27730 | SALT TOLERANCE ZINC FINGER (STZ) (ZAT10) | 0.955 | 6 | C2H2_40 |
| Ce_contig76425 | TF | AT1G27730 | SALT TOLERANCE ZINC FINGER (STZ) (ZAT10) | 0.967 | 6 | C2H2_62 |
| Ce_contig81969 | TF | AT1G27730 | SALT TOLERANCE ZINC FINGER (STZ) (ZAT10) | 0.988 | 6 | C2H2_71 |
| Ce_contig56990 | TF | AT4G34410 | REDOX RESPONSIVE TRANSCRIPTION FACTOR 1 (RRTF1) | 0.976 | 6 | ERF_75 |
| Ce_contig5939 | TF | AT5G48150 | PHYTOCHROME A SIGNAL TRANSDUCTION 1 (PAT1) | 0.946 | 6 | GRAS_6 |
| Ce_contig26571 | TF | AT4G37740 | GROWTH-REGULATING FACTOR 2 (GRF2) | 0.954 | 6 | GRF_1 |
| Ce_contig19510 | TF | AT5G66870 | ASYMMETRIC LEAVES 2-LIKE 1 (ASL1) | 0.934 | 6 | LBD_6 |
| Ce_contig49004 | TF | AT4G21440 | MYB-LIKE 102 (MYB102) | 0.974 | 6 | MYB_36 |
| Ce_contig8173 | TF | AT2G31180 | MYB DOMAIN PROTEIN 14 (MYB14) | 0.945 | 6 | MYB_9 |
| Ce_contig7 | TF | AT3G10500 | NAC DOMAIN CONTAINING PROTEIN 53 (NAC053) | 0.971 | 6 | NAC_1 |
| Ce_contig47321 | TF | AT5G13180 | VND-INTERACTING 2 (VNI2) | 0.990 | 6 | NAC_49 |
| Ce_contig85206 | TF | AT2G27990 | BEL1-LIKE HOMEODOMAIN PROTEIN 8 (BLH8) | 0.893 | 6 | TALE_22 |
| Ce_contig23430 | TF | AT4G01250 | WRKY22 | 0.957 | 6 | WRKY_25 |
| Ce_contig27558 | TF | AT4G23810 | WRKY53 | 0.974 | 6 | WRKY_29 |
| Ce_contig33086 | TF | AT3G56400 | WRKY70 | 0.963 | 6 | WRKY_34 |
| Ce_contig60019 | TF | AT4G01250 | WRKY22 | 0.909 | 6 | WRKY_50 |
| Ce_contig75830 | TF | AT5G13080 | WRKY75 | 0.913 | 6 | WRKY_59 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Testone, G.; Baldoni, E.; Iannelli, M.A.; Nicolodi, C.; Di Giacomo, E.; Pietrini, F.; Mele, G.; Giannino, D.; Frugis, G. Transcription Factor Networks in Leaves of Cichorium endivia: New Insights into the Relationship between Photosynthesis and Leaf Development. Plants 2019, 8, 531. https://doi.org/10.3390/plants8120531
Testone G, Baldoni E, Iannelli MA, Nicolodi C, Di Giacomo E, Pietrini F, Mele G, Giannino D, Frugis G. Transcription Factor Networks in Leaves of Cichorium endivia: New Insights into the Relationship between Photosynthesis and Leaf Development. Plants. 2019; 8(12):531. https://doi.org/10.3390/plants8120531
Chicago/Turabian StyleTestone, Giulio, Elena Baldoni, Maria Adelaide Iannelli, Chiara Nicolodi, Elisabetta Di Giacomo, Fabrizio Pietrini, Giovanni Mele, Donato Giannino, and Giovanna Frugis. 2019. "Transcription Factor Networks in Leaves of Cichorium endivia: New Insights into the Relationship between Photosynthesis and Leaf Development" Plants 8, no. 12: 531. https://doi.org/10.3390/plants8120531