Complete Chloroplast Genome of Rhipsalis baccifera, the only Cactus with Natural Distribution in the Old World: Genome Rearrangement, Intron Gain and Loss, and Implications for Phylogenetic Studies
Abstract
:1. Introduction
2. Results
2.1. Chloroplast Genome Organization and Features of Rhipsalis baccifera
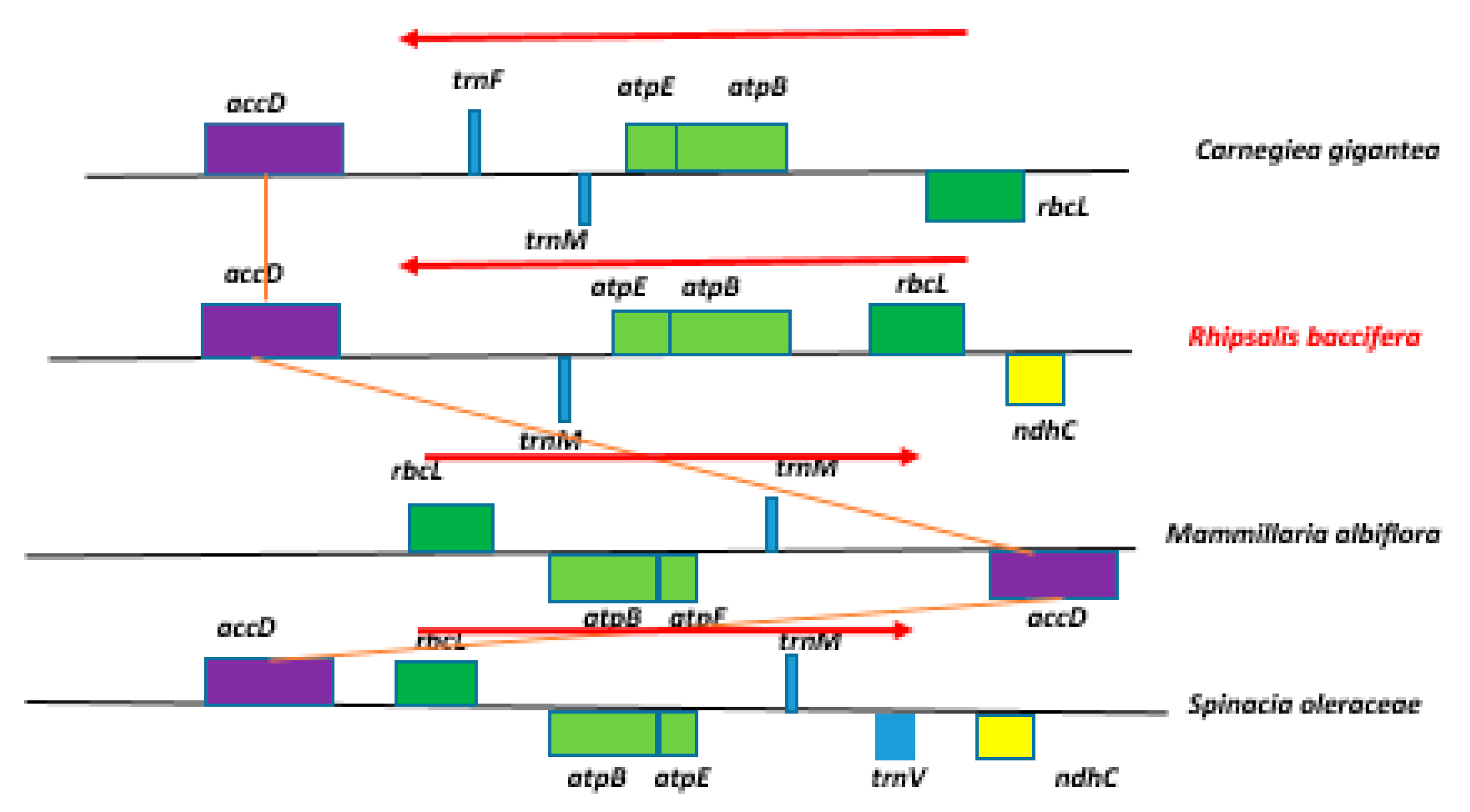
2.2. Inversions and Rearrangements in the cp Genome of Rhipsalis baccifera
2.2.1. rbcL - atpB - atpE-trnM Inversion
2.2.2. A 19-Gene Inversion in the LSC of Rhipsalis baccifera
2.2.3. Rearrangements in the SSC Region
2.3. Amino Acid Composition of Coded Proteins in R. baccifera cp Genome
2.4. Repeats
2.5. Boundaries between IRs and the Single Copy Regions
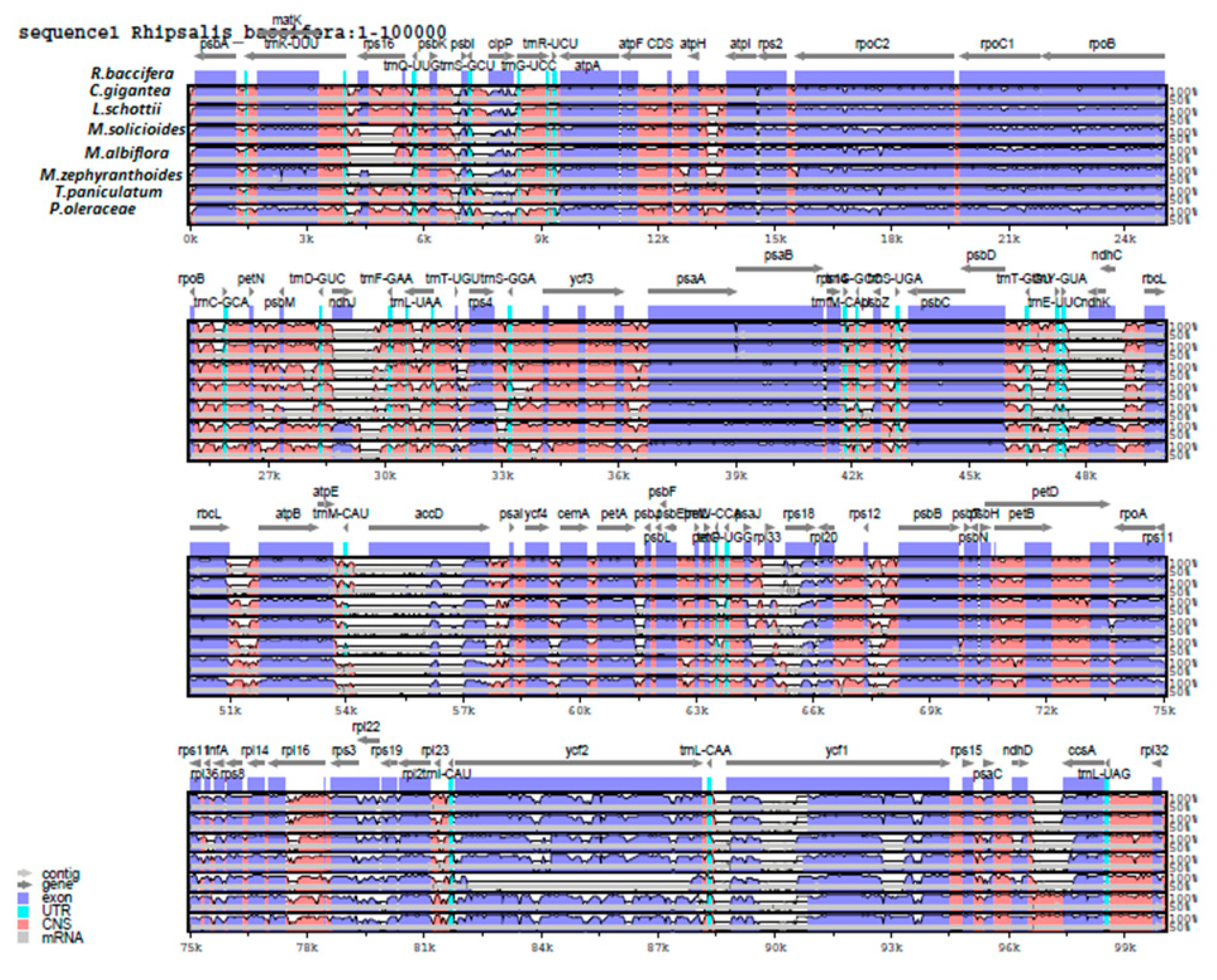
2.6. Genome Comparison and Sequence Divergence
2.7. Phylogenetic Analysis
3. Discussion
3.1. Chloroplast Genome Organization and Features of Rhipsalis baccifera
3.2. Inversions and Rearrangements in the cp Genome of Rhipsalis baccifera
3.3. Amino Acid Composition of Coded Proteins in R. baccifera
3.4. Repeats
3.5. Boundaries between IRs and the Single Copy Regions
3.6. Genome Comparison and Sequence Divergence
3.7. Phylogenetic Analysis
4. Materials and Methods
4.1. Chloroplast Genome Sequencing and Assembly
4.2. Genome Annotation and Codon Usage Analysis
4.3. Repeat Structure and Single Sequence Repeats (SSRs) Analysis
4.4. Genome Comparison and Sequence Divergence
4.5. Phylogenetic Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hershkovitz, M.A.; Zimmer, E.A. On the evolutionary origins of the cacti. Taxon 1997, 46, 217–232. [Google Scholar] [CrossRef]
- Nobel, P.S. CactiBiology and Uses; University of California Press: Berkeley, CA, USA; Los Angeles, CA, USA, 2002. [Google Scholar]
- Cronquist, A.; Thorne, R.F. Nomenclatural and Taxonomic History. In Caryophyllales; Behnke, H.D., Marby, T.J., Eds.; Springer: Berlin/Heidelberg, Germany, 1994; pp. 5–25. [Google Scholar]
- Nyffeler, R. The closest relatives of cacti: Insights from phylogenetic analyses of chloroplast and mitochondrial sequences with special emphasis on relationships in the tribe Anacampseroteae. Am. J. Bot. 2007, 94, 89–101. [Google Scholar] [CrossRef]
- Nyffeler, R.; Eggli, U. Disintegrating Portulacaceae: A new familial classification of the suborder Portulacineae (Caryophyllales) based on molecular and morphological data. TAXON 2010, 59, 227–240. [Google Scholar] [CrossRef] [Green Version]
- Nyffeler, R.; Eggli, U.; Ogburn, M.; Edwards, E. Variations On A Theme: Repeated Evolution Of Succulent Life Forms In the Portulacineae (Caryophyllales). Haseltonia 2008, 14, 26–36. [Google Scholar] [CrossRef]
- Ocampo, G.; Columbus, J.T. Molecular phylogenetics, historical biogeography, and chromosome number evolution of Portulaca (Portulacaceae). Mol. Phylogenetics Evol. 2012, 63, 97–112. [Google Scholar] [CrossRef] [PubMed]
- Barthlott, W.; Burstedde, K.; Geffert, J.; Ibisch, P.; Korotkova, N.; Miebach, A.; Rafiqpoor, M.D.; Stein, A.; Mutke, J. Biogeography and Biodiversity of Cacti. Schumannia 2015, 7, 208. [Google Scholar]
- Barthlott, W.; Taylor, N.P. Notes towards a Monograph of Rhipsalideae (Cactaceae). Bradleya 1995, 13, 43–79. [Google Scholar] [CrossRef]
- Hunt, D.; Taylor, N.; Charles, G. The new cactus lexicon; DH Books: Chapel Hill, NC, USA, 2006. [Google Scholar]
- Bomfim-Patrício, J.H.C.-S.M.C. Seed morphology, polyploidy and the evolutionary history of the epiphytic cactus Rhipsalis baccifera (Cactaceae). Polibotanica 2010, 29, 107–129. [Google Scholar]
- Barthlott, W. Biogeography and evolution in neo- and palaeotropical Rhipsalinae (Cactaceae). In Proceedings of the Dispersal and Distribution: An International Symposium, Hamburg, Germany, 10–12 June 1982; pp. 241–248. [Google Scholar]
- Korotkova, N.; Borsch, T.; Quandt, D.; Taylor, N.P.; Müller, K.F.; Barthlott, W.; Yuan, J.; Wang, B.-S.; Sun, J.; Pan, J.; et al. What does it take to resolve relationships and to identify species with molecular markers? An example from the epiphytic Rhipsalideae (Cactaceae). Am. J. Bot. 2011, 98, 1549–1572. [Google Scholar] [CrossRef]
- Facciola, S. Cornucopia II: A Source Book of Edible Plants; Kampong Publications: Vista, CA, USA, 1998. [Google Scholar]
- DeFilipps, R.A.; Maina, S.L.; Crepin, J. Medicinal Plants of the Guianas (Guyana, Surinam, French Guiana); National Museum of Natural History (US), Department of Botany: Washington, DC, USA, 2004.
- Christenhusz, M.J.M.; Chase, M.W. Biogeographical patterns of plants in the Neotropics—Dispersal rather than plate tectonics is most explanatory. Bot. J. Linn. Soc. 2012, 171, 277–286. [Google Scholar] [CrossRef] [Green Version]
- Calvente, A.; Andreata, R.H.P.; Vieira, R.C. Stem anatomy of Rhipsalis (Cactaceae) and its relevance for taxonomy. Plant Syst. Evol. 2008, 276, 1–7. [Google Scholar] [CrossRef]
- Tan, E.A. The phylogeography and cytogenetics of Rhipsalis baccifera: The cactus that made it to the Old World. Ph.D. Theses, Smith College, Northampton, MA, USA, 2016. [Google Scholar]
- Korotkova, N. Phylogeny and Evolution of the Epiphytic Rhipsalideae (Cactaceae). Ph.D. Thesis, University of Bonn, Bonn, Germany, 2011. [Google Scholar]
- Ravi, V.; Khurana, J.P.; Tyagi, A.K.; Khurana, P. An update on chloroplast genomes. Plant Syst. Evol. 2007, 271, 101–122. [Google Scholar] [CrossRef]
- Burke, S.V.; Grennan, C.P.; Duvall, M.R. Plastome sequences of two New World bamboos-Arundinaria gigantea and Cryptochloa strictiflora (Poaceae)-extend phylogenomic understanding of Bambusoideae. Am. J. Bot. 2012, 99, 1951–1961. [Google Scholar] [CrossRef]
- Dong, W.; Xu, C.; Cheng, T.; Zhou, S. Complete Chloroplast Genome of Sedum sarmentosum and Chloroplast Genome Evolution in Saxifragales. PLoS ONE 2013, 8, e77965. [Google Scholar] [CrossRef] [Green Version]
- Huang, H.; Shi, C.; Liu, Y.; Mao, S.-Y.; Gao, L.-Z. Thirteen Camellia chloroplast genome sequences determined by high-throughput sequencing: Genome structure and phylogenetic relationships. BMC Evol. Boil. 2014, 14, 151. [Google Scholar] [CrossRef] [Green Version]
- Yi, D.-K.; Lee, H.-L.; Sun, B.-Y.; Chung, M.Y.; Kim, K.-J. The Complete Chloroplast DNA Sequence of Eleutherococcus senticosus (Araliaceae); Comparative Evolutionary Analyses with Other Three Asterids. Mol. Cells 2012, 33, 497–508. [Google Scholar] [CrossRef] [Green Version]
- Bell, C.D.; Soltis, U.E.; Soltis, P.S. The age and diversification of the angiosperms re-revisited. Am. J. Bot. 2010, 97, 1296–1303. [Google Scholar] [CrossRef]
- Drouin, G.; Daoud, H.; Xia, J. Relative rates of synonymous substitutions in the mitochondrial, chloroplast and nuclear genomes of seed plants. Mol. Phylogenetics Evol. 2008, 49, 827–831. [Google Scholar] [CrossRef] [PubMed]
- Henry, R.J. Plant Diversity and Evolution: Genotypic and Phenotypic Variation in Higher Plants; CABI Publishing: Wallingford, UK, 2005; 332p. [Google Scholar] [CrossRef]
- Wolfe, K.H.; Li, W.-H.; Sharp, P.M. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. In Proceedings of the Proceedings of the National Academy of Sciences. Proc. Natl. Acad. Sci. USA 1987, 84, 9054–9058. [Google Scholar] [CrossRef] [Green Version]
- Cosner, E.M.; Raubeson, A.L.; Jansen, R.K. Chloroplast DNA rearrangements in Campanulaceae: Phylogenetic utility of highly rearranged genomes. BMC Evol. Boil. 2004, 4, 27. [Google Scholar] [CrossRef] [Green Version]
- Solórzano, S.; Chincoya, D.A.; Sanchez-Flores, A.; Estrada, K.; Díaz-Velásquez, C.E.; González-Rodríguez, A.; Vaca-Paniagua, F.; Dávila, P.; Arias, S.; Flores, S.-; et al. De Novo Assembly Discovered Novel Structures in Genome of Plastids and Revealed Divergent Inverted Repeats in Mammillaria (Cactaceae, Caryophyllales). Plants 2019, 8, 392. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Isaac, N.J.B. Taxonomic inflation: Its influence on macroecology and conservation. Trends Ecol. Evol. 2004, 19, 464–469. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haig, S.M.; Beever, E.A.; Chambers, S.M.; Draheim, H.M.; Dugger, B.D.; Dunham, S.; Elliott-Smith, E.; Fontaine, J.B.; Kesler, D.C.; Knaus, B.J.; et al. Taxonomic Considerations in Listing Subspecies Under the U.S. Endangered Species Act. Conserv. Boil. 2006, 20, 1584–1594. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, X.; Li, Y.; Yang, H.; Zhou, B. Chloroplast Genome of the Folk Medicine and Vegetable Plant Talinum paniculatum (Jacq.) Gaertn.: Gene Organization, Comparative and Phylogenetic Analysis. Molecules 2018, 23, 857. [Google Scholar] [CrossRef] [Green Version]
- Jansen, R.K.; Raubeson, L.A.; Boore, J.L.; dePamphilis, C.W.; Chumley, T.W.; Haberle, R.C.; Wyman, S.K.; Alverson, A.J.; Peery, R.; Herman, S.J.; et al. Methods for Obtaining and Analyzing Whole Chloroplast Genome Sequences. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2005; Volume 395, pp. 348–384. [Google Scholar]
- Li, B.; Zheng, Y. Dynamic evolution and phylogenomic analysis of the chloroplast genome in Schisandraceae. Sci. Rep. 2018, 8, 9285. [Google Scholar] [CrossRef]
- Plunkett, G.M.; Downie, S.R. Expansion and Contraction of the Chloroplast Inverted Repeat in Apiaceae Subfamily Apioideae. Syst. Bot. 2000, 25, 648. [Google Scholar] [CrossRef]
- McPherson, M.A.; Fay, M.F.; Chase, M.W.; Graham, S.W. Parallel Loss of a Slowly Evolving Intron from Two Closely Related Families in Asparagales. Syst. Bot. 2004, 29, 296–307. [Google Scholar] [CrossRef]
- Jansen, R.K.; Wojciechowski, M.F.; Sanniyasi, E.; Lee, S.-B.; Daniell, H.W. Complete plastid genome sequence of the chickpea (Cicer arietinum) and the phylogenetic distribution of rps12 and clpP intron losses among legumes (Leguminosae). Mol. Phylogenetics Evol. 2008, 48, 1204–1217. [Google Scholar] [CrossRef] [Green Version]
- McNeal, J.R.; Kuehl, J.V.; Boore, J.L.; Leebens-Mack, J.; Depamphilis, C.W. Parallel Loss of Plastid Introns and Their Maturase in the Genus Cuscuta. PLoS ONE 2009, 4, e5982. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Yang, H.; Zhao, J.; Zhou, B.; Li, T.; Xiang, B. The complete chloroplast genome sequence of the folk medicinal and vegetable plant purslane (Portulaca oleracea L.). J. Hortic. Sci. Biotechnol. 2017, 93, 356–365. [Google Scholar] [CrossRef]
- Sanderson, M.J.; Copetti, D.; Búrquez, A.; Bustamante, E.; Charboneau, J.L.M.; Eguiarte, L.E.; Kumar, S.; Lee, H.O.; Lee, J.; McMahon, M.; et al. Exceptional reduction of the plastid genome of saguaro cactus (Carnegiea gigantea): Loss of the ndh gene suite and inverted repeat. Am. J. Bot. 2015, 102, 1115–1127. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Wang, Z.; Wang, H.; Su, Y.; Wang, T. Patterns and Rates of Plastid rps12 Gene Evolution Inferred in a Phylogenetic Context using Plastomic Data of Ferns. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, K.; Subramanian, A.R. Proteomic identification of all plastid-specific ribosomal proteins in higher plant chloroplast 30S ribosomal subunit. PSRP-2 (U1A-type domains), PSRP-3alpha/beta (ycf65 homologue) and PSRP-4 (Thx homologue). JBIC J. Boil. Inorg. Chem. 2003, 270, 190–205. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.-S.; Lee, B.Y.; Kwak, M. The complete chloroplast genome sequences of Lychnis wilfordii and Silene capitata and comparative analyses with other Caryophyllaceae genomes. PLoS ONE 2017, 12, e0172924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sloan, D.B.; Triant, D.A.; Forrester, N.J.; Bergner, L.M.; Wu, M.; Taylor, U.R. A recurring syndrome of accelerated plastid genome evolution in the angiosperm tribe Sileneae (Caryophyllaceae). Mol. Phylogenetics Evol. 2014, 72, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Kohler, M.; Reginato, M.; Souza-Chies, T.T.; Majure, L.C. Insights into chloroplast genome variation across Opuntioideae (Cactaceae). bioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- He, Z.; Mi, H. Functional Characterization of the Subunits N, H, J, and O of the NAD(P)H Dehydrogenase Complexes in Synechocystis sp. Strain PCC 6803. Plant Physiol. 2016, 171, 1320–1332. [Google Scholar] [CrossRef] [Green Version]
- Downie, S.R.; Palmer, J.D. A Chloroplast DNA Phylogeny of the Caryophyllales Based on Structural and Inverted Repeat Restriction Site Variation. Syst. Bot. 1994, 19, 236. [Google Scholar] [CrossRef]
- Salim, H.M.W.; Cavalcanti, A.R.O. Factors influencing codon usage bias in genomes. J. Braz. Chem. Soc. 2008, 19. [Google Scholar] [CrossRef] [Green Version]
- Zotov, V.; Punina, N.V.; Dorokhov, D.B.; Schaad, N.W.; Ignatov, A. Phylogenetic Changes in Chloroplast Genomes. Comp. Evol. Genom. Proteom. 2006, 1, 249–252. [Google Scholar]
- Sharp, P.M.; Li, W.-H. The codon adaptation index-a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 1987, 15, 1281–1295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haberle, R.C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Extensive Rearrangements in the Chloroplast Genome of Trachelium caeruleum are Associated with Repeats and tRNA Genes. J. Mol. Evol. 2008, 66, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Vieira, M.L.C.; Santini, L.; Diniz, A.L.; Munhoz, C.D.F. Microsatellite markers: What they mean and why they are so useful. Genet. Mol. Boil. 2016, 39, 312–328. [Google Scholar] [CrossRef] [PubMed]
- Lavin, M.; Doyle, J.J.; Palmer, J.D. Evolutionary Significance of the Loss of the Chloroplast-DNA Inverted Repeat in the Leguminosae Subfamily Papilionoideae. Evolution 1990, 44, 390. [Google Scholar] [CrossRef]
- Palmer, J.D.; Osorio, B.; Thompson, W.F. Evolutionary significance of inversions in legume chloroplast DNAs. Curr. Genet. 1988, 14, 65–74. [Google Scholar] [CrossRef]
- Shinozaki, K.; Ohme, M.; Tanaka, M.; Wakasugi, T.; Hayashida, N.; Matsubayashi, T.; Zaita, N.; Chunwongse, J.; Obokata, J.; Yamaguchi-Shinozaki, K.; et al. The complete nucleotide sequence of the tobacco chloroplast genome: Its gene organization and expression. EMBO J. 1986, 5, 2043–2049. [Google Scholar] [CrossRef]
- Schmitz-Linneweber, C.; Williams-Carrier, R.E.; Williams-Voelker, P.M.; Kroeger, T.S.; Vichas, A.; Barkan, A. A Pentatricopeptide Repeat Protein Facilitates the trans-Splicing of the Maize Chloroplast rps12 Pre-mRNA. Plant Cell 2006, 18, 2650–2663. [Google Scholar] [CrossRef] [Green Version]
- Keller, J.; Rousseau-Gueutin, M.; Martin, G.; Morice, J.; Boutte, J.; Coissac, E.; Ourari, M.; Aïnouche, M.; Salmon, A.; Cabello-Hurtado, F.; et al. The evolutionary fate of the chloroplast and nuclear rps16 genes as revealed through the sequencing and comparative analyses of four novel legume chloroplast genomes from Lupinus. DNA Res. 2017, 24, 343–358. [Google Scholar] [CrossRef] [Green Version]
- Steyn, E.M.A.; Smith, G. Talinum paniculatum, a naturalized weed in South Africa. Bothalia Afr. Biodiv. Conserv. 2002, 31, 195–197. [Google Scholar]
- Veselova, T.; Dzhalilova, K.K.; Remizowa, M.; Timonin, A. Embryology of Talinum paniculatum (Jacq.) Gaertn. and T. triangulare (Jacq.) Willd. (Portulacaceae s.l., Caryophyllales). Wulfenia 2012, 19, 107–129. [Google Scholar]
- Doyle, J. DNA protocols for plants. In Molecular Techniques in Taxonomy; Hewitt, G.M., Johnston, A.W.B., Young, J.P.W., Eds.; Springer: Berlin/Heidelberg, Germany, 1991; pp. 283–293. [Google Scholar]
- Jin, J.-J.; Yu, W.-B.; Yang, J.-B.; Song, Y.; Depamphilis, C.W.; Yi, T.-S.; Li, D.-Z. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. bioRxiv 2018. [Google Scholar] [CrossRef] [Green Version]
- Tillich, M.; Lehwark, P.; Pellizzer, T.; Ulbricht-Jones, E.S.; Fischer, A.; Bock, R.; Greiner, S. GeSeq—Versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 2017, 45, W6–W11. [Google Scholar] [CrossRef] [PubMed]
- Lowe, T.M.; Chan, P.P. tRNAscan-SE On-line: Integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 2016, 44, W54–W57. [Google Scholar] [CrossRef] [PubMed]
- Qu, X.-J.; Moore, M.J.; Li, D.-Z.; Yi, T.-S. PGA: A software package for rapid, accurate, and flexible batch annotation of plastomes. Plant Methods 2019, 15, 50. [Google Scholar] [CrossRef] [Green Version]
- Lohse, M.; Drechsel, O.; Kahlau, S.; Bock, R. OrganellarGenomeDRAW—A suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucleic Acids Res. 2013, 41, W575–W581. [Google Scholar] [CrossRef]
- Sudhir Kumar, G.S.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar]
- Kurtz, S. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [Green Version]
- Thiel, T.; Michalek, W.; Varshney, R.K.; Graner, A. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor. Appl. Genet. 2003, 106, 411–422. [Google Scholar] [CrossRef]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef]
- Darling, E.A.; Mau, B.; Blattner, F.R.; Perna, N.T. Mauve: Multiple Alignment of Conserved Genomic Sequence with Rearrangements. Genome Res. 2004, 14, 1394–1403. [Google Scholar] [CrossRef] [Green Version]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.; Von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, D.; Gao, F.-L.; Jakovlić, I.; Zou, H.; Zhang, J.; Li, W.X.; Wang, G.T.; Zhou, H. PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol. Ecol. Resour. 2019, 20, 348–355. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van Der Mark, P.; Ayres, D.L.; E Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice Across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed] [Green Version]








| Region | Size (bp) | T (U) (%) | A (%) | G (%) | C (%) | Genes |
|---|---|---|---|---|---|---|
| LSC | 81,459 | 32.6 | 31.4 | 17.7 | 18.3 | 84 |
| SSC | 23,531 | 30.5 | 30.2 | 20.3 | 19.0 | 19 |
| IRA | 8530 | 31.4 | 31.7 | 19.0 | 17.9 | 4 |
| IRB | 8530 | 31.4 | 31.7 | 19.0 | 17.9 | 4 |
| Total | 122,333 | 32.1 | 31.2 | 18.3 | 18.4 | 110 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oulo, M.A.; Yang, J.-X.; Dong, X.; Wanga, V.O.; Mkala, E.M.; Munyao, J.N.; Onjolo, V.O.; Rono, P.C.; Hu, G.-W.; Wang, Q.-F. Complete Chloroplast Genome of Rhipsalis baccifera, the only Cactus with Natural Distribution in the Old World: Genome Rearrangement, Intron Gain and Loss, and Implications for Phylogenetic Studies. Plants 2020, 9, 979. https://doi.org/10.3390/plants9080979
Oulo MA, Yang J-X, Dong X, Wanga VO, Mkala EM, Munyao JN, Onjolo VO, Rono PC, Hu G-W, Wang Q-F. Complete Chloroplast Genome of Rhipsalis baccifera, the only Cactus with Natural Distribution in the Old World: Genome Rearrangement, Intron Gain and Loss, and Implications for Phylogenetic Studies. Plants. 2020; 9(8):979. https://doi.org/10.3390/plants9080979
Chicago/Turabian StyleOulo, Millicent Akinyi, Jia-Xin Yang, Xiang Dong, Vincent Okelo Wanga, Elijah Mbandi Mkala, Jacinta Ndunge Munyao, Victor Omondi Onjolo, Peninah Cheptoo Rono, Guang-Wan Hu, and Qing-Feng Wang. 2020. "Complete Chloroplast Genome of Rhipsalis baccifera, the only Cactus with Natural Distribution in the Old World: Genome Rearrangement, Intron Gain and Loss, and Implications for Phylogenetic Studies" Plants 9, no. 8: 979. https://doi.org/10.3390/plants9080979






