Abstract
Whole-plant corn silages on family farms were sampled in Erdos (S1), Baotou (S2), Ulanqab (S3), and Hohhot (S4) in North China, after 300 d of ensiling. The microbial communities, metabolites, and aerobic stability were assessed. Lactobacillus buchneri, Acinetobacter johnsonii, and unclassified Novosphingobium were present at greater abundances than others in S2 with greater bacterial diversity and metabolites. Lactobacillus buchneri, Lactobacillus parafarraginis, Lactobacillus kefiri, and unclassified Lactobacillus accounted for 84.5%, and 88.2%, and 98.3% of bacteria in S1, S3, and S4, respectively. The aerobic stability and fungal diversity were greater in S1 and S4 with greater abundances of unclassified Kazachstania, Kazachstania bulderi, Candida xylopsoci, unclassified Cladosporium, Rhizopus microspores, and Candida glabrata than other fungi. The abundances of unclassified Kazachstania in S2 and K. bulderi in S3 were 96.2% and 93.6%, respectively. The main bacterial species in S2 were L. buchneri, A. johnsonii, and unclassified Novosphingobium; Lactobacillus sp. dominated bacterial communities in S1, S3, and S4. The main fungal species in S1 and S4 were unclassified Kazachstania, K. bulderi, C. xylopsoci, unclassified Cladosporium, R. microspores, and C. glabrata; Kazachstania sp. dominated fungal communities in S2 and S3. The high bacterial diversity aided the accumulation of metabolites, and the broad fungal diversity improved the aerobic stability.
1. Introduction
Family farms, as new agricultural business entities, partake in more commercially oriented agricultural production and management than their forerunners (“farming households”) in China, even though their main labour force still consists of family members, and the major source of their income remains agricultural work [1]. Family farms are the main mode of management in the Inner Mongolian pastoral area of North China [2]. Whole-plant corn silage, alfalfa hay, grass hay, crop straw, and natural grass are the major forages of family farms in the desert steppe of Inner Mongolia. In general, these farmers ensile whole-plant corn without any additives to save processing costs [3]. Moreover, the microbial communities present in this form of whole-plant corn silage are still unclear, and understanding them is critical to explaining the fermentation quality and aerobic stability of such silage.
In the past decade, the development of next-generation sequencing technologies has helped to improve our understanding of the microbial communities present in silage [4]. Lactobacillus dominates the bacterial succession and determines the fermentation quality of whole-plant corn silage during fermentation process [5,6]. The main bacterial species present in the silage at the end of the process are Lactobacillus acetotolerans, Lactobacillus silagei, Lactobacillus parafarraginis, Lactobacillus buchneri, and Lactobacillus odoratitofui [7]. The geographical location where at the corn was grown influences the bacterial succession process taking place in whole-plant corn silage and the bacterial community in the final silage [8,9]. Drouin et al. [10] reported that ensiling whole-plant corn with lactic acid bacteria (LAB) increases the Shannon indexes of bacterial and fungal communities and modifies the aerobic stability, and Saccharomyces, Issatchenkia, and Kazachstania cause aerobic deterioration during aerobic exposure. Nevertheless, the effect of the geographical location on the fungal community and the aerobic stability of whole-plant corn silage has not yet been reported on.
Recently, the silage metabolome has attracted new interest [11]. Some studies have reported the effect of inoculating on the metabolite contents of whole-plant corn silage, alfalfa silage, and sainfoin silage with LAB [6,9,12]; the dynamics of metabolites in whole-plant corn silages during fermentation process [6]; and the correlation of the main metabolites with the main bacterial species and fermentation quality in whole-plant corn silages [6]. Additionally, Wu et al. [13] analysed the metabolic profiles of high-moisture sweetcorn kernel silages. Biogenic amines—a class of metabolites in silage—affect the palatability of silages, and the feed intake and performance of ruminants [14]. However, the concentration and composition of the metabolites—especially the biogenic amines in silages on the desert steppe—remain unclear.
In the present study, whole-plant corn silages on family farms in the desert steppe of Inner Mongolia in North China were assessed after about 300 d of ensiling. The hypothesis of the present study was the differences in microbial communities, metabolites, and aerobic stability between whole-plant corn silages from different areas. The objective was to analyse the bacterial and fungal communities, the metabolites, the fermentation quality and the aerobic stability of whole-plant corn silages.
2. Materials and Methods
2.1. Sampling
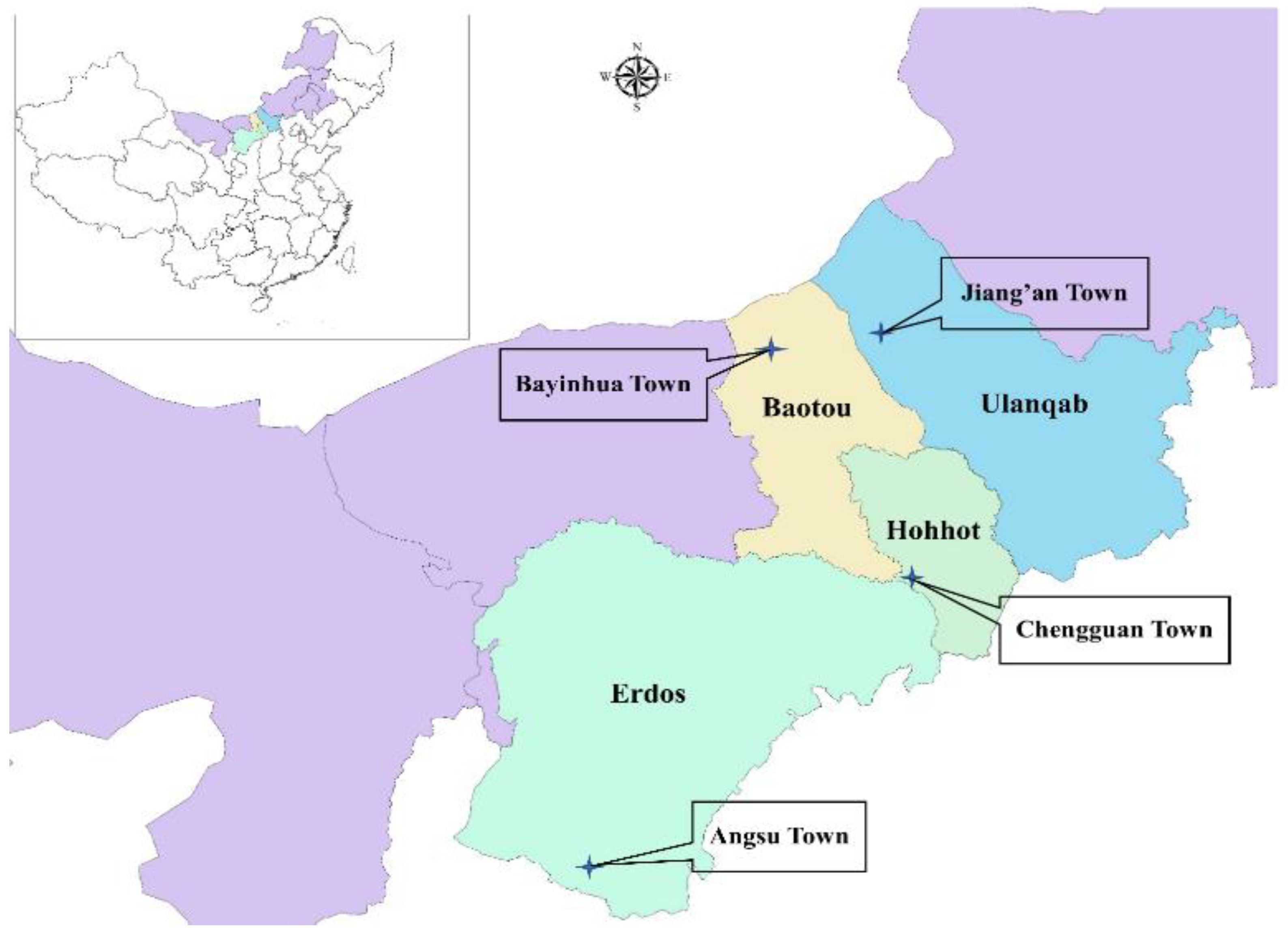
The corn (Zea mays L.) used for ensiling came from replicates grown on 3 family farms of each sampling area (silage 1 (S1), Angsu Town, Erdos City, 108°4′29.093″ E, 38°14′24.212″ N, 1378 m; S2, Bayinhua Town, Baotou City, 109°54′15.260″ E, 42°16′34.133″ N, 1251 m; S3, Jiang’an Town, Ulanqab City, 110°51′14.249″ E, 43°10′06.845″ N, 1077 m; S4, Chengguan Town, Hohhot City, 111°39′31.784″ E, 39°55′19.160″ N, 1137 m) in Inner Mongolia (desert steppe), North China (Figure 1). The variety of corn used was 23 Yu (No. 2008022, corn hybrid, Henan Dajingjiu Seed Industry Co., Ltd., Shangqiu, China). The corn plants were harvested at the 2/3, 1/3, 1/3, and 1/2 milk-line stages in Erdos (on 15 September 2018), Baotou (on 1 September 2018), Ulanqab (on 4 September 2018), and Hohhot (on 8 September 2018), respectively according to local tradition, and chopped into 1–2 cm pieces by combine-harvester (John Deere (China) Investment Co., Ltd., Beijing, China). The chopped forages on each family farm were ensiled in a silo (length, 100 m; width, 10 m; height, 2 m) without any additives at a density of more than 700 kg/m3. The samples of chopped forage were taken at filling to the 1/3, 1/2, and 2/3 of the length of the silo, respectively, and transported in ice boxes to the laboratory for analysis. After 30 d of ensiling, the silos were opened, and the silages were collected (around 30 cm thickness, from the front phase of the silo) for animal feeding on every morning. At about 300 d, the silages were sampled after the animals were fed. Subsamples were collected at 5 points along the face of a silo from each family farm: point 1, 1 m from the right wall, top film; point 2, 1 m from the left wall, top film; point 3, 1 m from the right wall, bottom; point 4, 1 m from the left wall, bottom; point 5, the centre of the silo’s face. The subsamples (5 kg) were collected with a needle forage sampler about 30 cm long and uniformly mixed as a composite sample. The samples were transported in ice boxes to the laboratory for analysis.
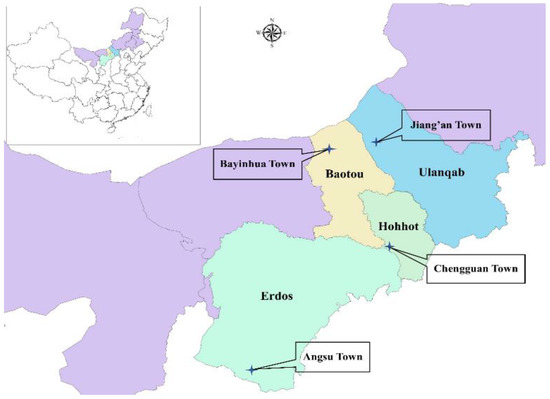
Figure 1.
Map of sampling locations.
2.2. Analyses
The silage from each silo was randomly divided into 4 batches (4 kg each). Every batch from each silo was placed into a separate plastic drum (diameter, 40 cm; height, 50 cm), and the tops of the drums were covered with 2 layers of cheesecloth. One batch from each silo was used for assessing the aerobic stability, which was measured with a temperature recorder (SMOWO Multi-Channel Data Logger, MDL-1048A; Shanghai Tianhe Automation Instrument Co., Ltd., Shanghai, China) via the method of Wang et al. [15]. The recorder’s temperature sensor probe was placed into the centre of the silage in the drum. The temperatures of room and silage were recorded every hour. One of the three batches from each silo was randomly selected at 0, 2, and 5 d for assessment of microbial counts and fermentation quality; the silages were assessed at 0 d for microbial communities and metabolites.
Extracts of forage or silage were prepared by homogenizing 20 g of chopped forage or silage with 180 mL of sterile water for 100 s using a flap-sterile homogenizer (LW-09, Shanghai Jingxin Industrial Development Co., Ltd., Shanghai, China) and then filtering through 2 layers of cheesecloth according to the methods of Owens et al. [16]. The pH of the extracts was detected with a pH meter (PB-10, Sartorius, Gottingen, Germany). Counts of LAB, Escherichia coli, bacteria, and yeast/moulds were taken by culturing on De Man Rogosa Sharpe agar, violet-red bile agar, nutrient agar, and potato dextrose agar, respectively, at 37 °C for 48 h in an incubator (LRH-70, Shanghai Yiheng Science Instruments Co., Ltd., Shanghai, China) [17]. Water-soluble carbohydrates (WSC) were measured via anthrone–sulfuric acid colorimetry using a spectrophotometer (Genesys 10; Thermo Fisher Scientific, Waltham, MA, USA) at 620 nm.
The organic acids (lactic acid, acetic acid, propionic acid, and butyric acid) in the silages were assessed using high-performance liquid chromatography (DAD, 210 nm, SPD-20A, Shimadzu Co., Ltd., Kyoto, Japan) (detector, SPD-20A diode array detector (210 nm); column, Shodex RS Pak KC-811 (50 °C, Showa Denko K.K., Kawasaki, Japan); mobile phase, 3 mM HClO4 (1.0 mL/min)) [18]. The ammonia nitrogen content was determined via the Kjeldahl method using an autoanalyzer (Kjeltec 8400; FOSS Co., Ltd., Hillerød, Denmark).
The chopped forages or silages were dried using a forced-air oven (BPG-9240A, Shanghai Yiheng Scientific Instrument Co., Ltd., Shanghai, China) operated at 65 °C for 48 h, ground through a 1 mm screen with a mill (FS-6D; Fichi Machinery Equipment Co., Ltd., Jinan, Shandong, China), and then dried at 105 °C, until reaching a constant mass suitable for measuring dry matter (DM) content. The total nitrogen (TN) was assessed via the Kjeldahl method using an autoanalyzer with copper as the catalyst, and the crude protein (CP) concentration was calculated by multiplying the TN concentration by 6.25. The neutral detergent fibre (NDF) and acid detergent fibre (ADF) contents were determined via the method of Van Soest et al. [19] using an Ankom fiber analyzer (2000, Ankom, Macedon, NY, USA) without heat-stable amylase; acid detergent lignin (ADL) content was determined using a 72% H2SO4 solution [19].
The total bacterial DNA and fungal DNA were extracted using a DNA isolation kit (OMEGA-D4015-04) following the manufacturer’s specifications. The DNA quality was evaluated via 1% (w/v) agarose gel electrophoresis. The full-length 16S rRNA gene was amplified via PCR using forward primer 5′-TAGRGTTYGATYMTGGCTCAG-3′ and reverse primer 5′-RGYTACCTTGTTACGACTT-3′; the full-length internal transcribed spacer (ITS) gene was amplified via PCR using forward primer 5′-TCCGTAGGTGAACCTGCGG-3′ and reverse primer 5′-TCCTCCGCTTATTGATATGC-3′. The amplification conditions were as follows: 95 °C for 3 min; 25 cycles of 98 °C for 20 s, 57 °C for 30 s and 72 °C for 90 s; a final extension of 72 °C for 2 min. The 16S rRNA and ITS libraries were built with a Pacific Biosciences Template Prep Kit (Pacific Biosciences, Menlo Park, CA, USA). Amplicon sequencing was performed with a PacBio Sequel instrument (Pacific Biosciences). Raw circular consensus sequencing (CCS) reads were obtain using PacBio SMRT Link CCS software. The sequence data reported in this study have been submitted to the NCBI Sequence Read Archive database under the accession number PRJNA650284.
The silage extracts were prepared as follows: 5 g silage and 10 mL extraction liquid (70% methanol) were vortexed for 30 s, oscillated for 1 h at 4 °C, filtered through a 0.22 µm membrane, and then dried to 1 mL with a vacuum concentrator in a glass vial [20]. The metabolites in the silage were analysed via liquid chromatography–electrospray ionization–tandem mass spectrometry (HPLC, Shim-pack UFLC Shimadzu CBM30A system; MS, Applied Biosystems 4500 QTRAP system) using measured silage extracts. The conditions of analysis were as follows: HPLC column, Waters ACQUITY UPLC HSS T3 C18 (1.8 µm, 2.1 mm × 100 mm); mobile phase, water (0.04% acetic acid)/acetonitrile (0.04% acetic acid); gradient program, 100:0 (v:v) at 0 min, 5:95 at 11.0 min, 5:95 at 12.0 min, 95:5 at 12.1 min, and 95:5 at 15.0 min; flow rate, 0.40 mL/min; temperature, 40 °C; injection volume, 5 µL [7]. The effluent was then connected to an ESI triple-quadrupole linear ion trap (QqQ-LIT) mass spectrometer [7]. The qualitative and quantitative analyses, the pre-processing and the determination of the relative concentrations of metabolites were all undertaken according to Yan et al. [20] and Xu et al. [6]. Biogenic amines (tyramine, putrescine, phenylethylamine, spermidine, and noradrenaline) were selected from the metabolites, and their relative concentrations were recorded.
2.3. Statistical Analyses
The data of fermentation quality and microbial count were analysed via a 4 × 3 factorial design. The model included 4 sampling areas and 3 aerobic exposure times, and their interaction. Using one-factor analysis of variance via the general linear model (GLM) of SAS (SAS System for Windows, version 9.1.3; SAS Institute Inc., Cary, NC, USA), we analysed the differences in the pH, microbial counts, and nutritional composition of fresh forage from 4 sampling areas, in the fermentation quality and microbial counts of silage among 4 sampling areas and 3 aerobic exposure times, and in the sequencing data, alpha diversity, nutritional composition, total metabolites, and biogenic amines of silage from 4 sampling areas. The interaction between the sampling area and aerobic exposure time was analysed using the PDIFF procedure of SAS. The differences were compared via least significant differences, and significance was declared at p ≤ 0.05. The principal coordinates analysis (PCoA) of microbial beta diversity was performed using R 1.7.13 for Windows; the principal component analysis (PCA) of the metabolic profiles was performed using R 3.5.1.
3. Results
3.1. Characteristics of Whole-Plant Corns before Ensiling
The S1 and S4 had higher LAB counts than S2 and S3 (p < 0.05). The S2 and S3 displayed lower DM contents and greater WSC concentrations than S1 and S4 (p < 0.05), and S1 showed greater DM than S4 (p < 0.05). The S2 had the greatest level of CP, S4 contained the most NDF and ADF, and S3 had the lowest contents of ADF and ADL (p < 0.05) (Table 1).

Table 1.
pH, microbial counts (log10 CFU/g fresh weight), dry matter content (DM, g/kg), and chemical component concentrations (g/kg DM) of whole-plant corn before ensiling (n = 3).
3.2. Microbial Counts and Diversity
At 0 d of aerobic exposure, S1 and S4 showed greater LAB and yeast counts than S2 and S3 (p < 0.05), and S1 had a greater bacterial count than other silages (p < 0.05); additionally, S3 contained more LAB and yeast than S2, and fewer bacteria than S2 and S4 (p < 0.05) (Table 2). During aerobic exposure, the LAB counts in S2 and S3 increased (p < 0.05), and the bacteria and yeast counts in S1 dropped (p < 0.05). The bacteria count decreased at 2 d and increased at 5 d in S2 (p < 0.05), while it increased at 2 d and decreased at 5 d in S3 and S4 (p < 0.05). The aerobic exposure time affected the LAB and bacteria counts (p < 0.05). The sampling area influenced the counts of LAB, bacteria, and yeast (p < 0.05), on which the aerobic exposure time and the sampling area had a compound effect (p < 0.05). Escherichia coli and moulds were not detected in all the silages.

Table 2.
Microbial counts (log10 CFU/g fresh weight) of whole-plant corn silages during aerobic exposure (n = 3).
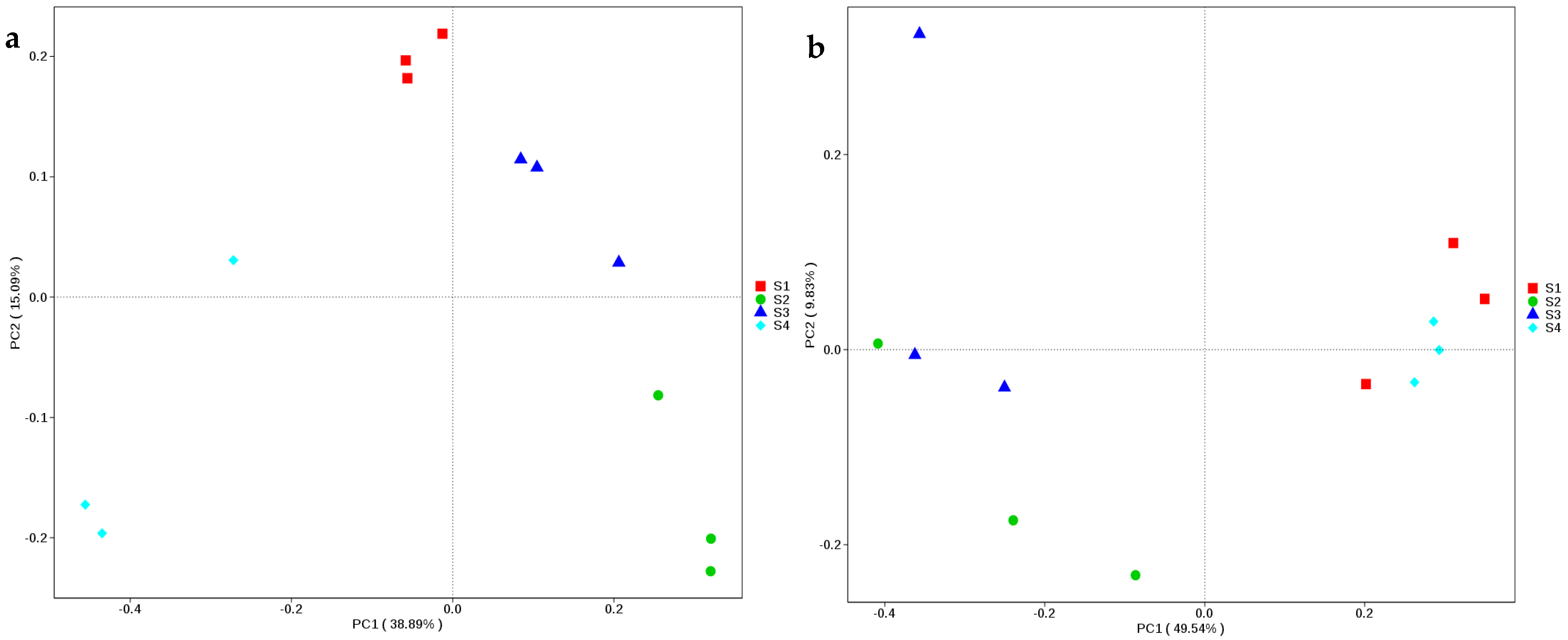
Totals of 166,552 and 141,221 clean reads of the full-length 16S rRNA and ITS genes, respectively, were obtained from 12 samples of whole-plant corn silage according to SMRT sequencing. The S4 had a greater number of 16S rRNA gene reads than S1 (p < 0.05), and these silages contained more than 10,000 clean reads. There were no differences in the numbers of clean reads of the ITS gene among the silages (p > 0.05), more than 10,000 clean reads were derived for each, except for S1 (9250). In terms of bacteria, S2 showed the highest Shannon and Simpson indexes among the silages, and it also had a higher Chao1 index and number of observed species than S4 (p < 0.05). As regards fungi, the numbers of observed species and the indexes of Shannon, Simpson and Chao1 indexes in S1 and S4 were higher than those in S2 and S3 (p < 0.05) (Table 3). According to PCoA, the bacterial communities in all the silages were clearly distinct from one another; however, the fungal communities in S1 and S4 were distinct from those in S2 and S3 (Figure 2).

Table 3.
Sequencing data and alpha diversity of bacteria and fungi in whole-plant corn silage (n = 3).
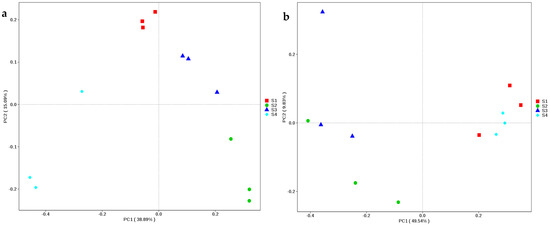
Figure 2.
Non-metric multi-dimensional scaling based on Bray-Curtis assessment of bacterial (a) and fungal (b) diversities in whole-plant corn silages (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot.
3.3. Bacterial Community
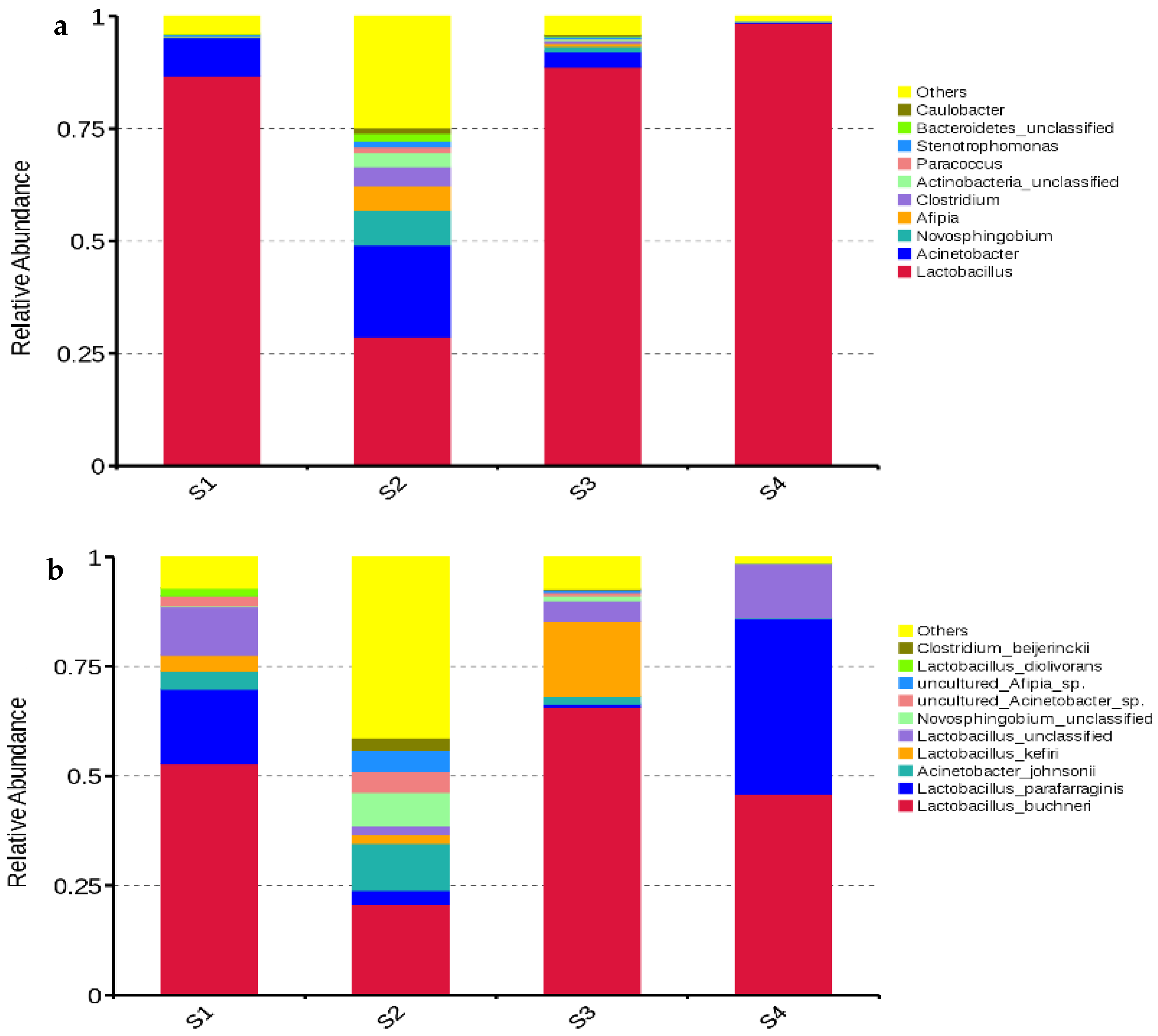
Lactobacillus dominated the bacterial communities in S1, S3, and S4 with abundances of 86.74%, 88.69%, and 98.49%, respectively. The main bacterial genera in S2 were Lactobacillus (28.70%), Acinetobacter (20.40%), Novosphingobium (7.80%), and Afipia (5.37%) (Figure 3a). The main bacterial species in S1, S3, and S4 were L. buchneri (52.74%, 65.79%, and 45.92%, respectively), L. parafarraginis (17.09%, 0.64%, and 40.06%, respectively), Lactobacillus kefiri (3.58%, 17.14%, and 0%, respectively) and unclassified Lactobacillus (11.05%, 4.68%, and 12.36%, respectively). The dominant bacterial species in S2 were L. buchneri (20.71%), Acinetobacter johnsonii (10.73%), unclassified Novosphingobium (7.64%), and L. parafarraginis (3.19%) (Figure 3b).
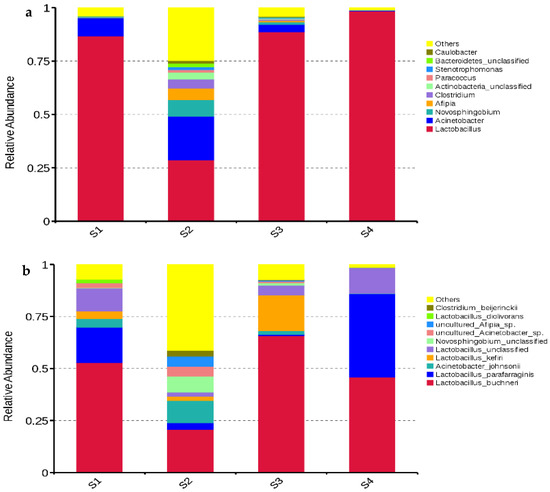
Figure 3.
Relative abundances of bacterial genera (a) and species (b) in whole-plant corn silage (n = 3). S1, silage collected frm Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot.
3.4. Fungal Community
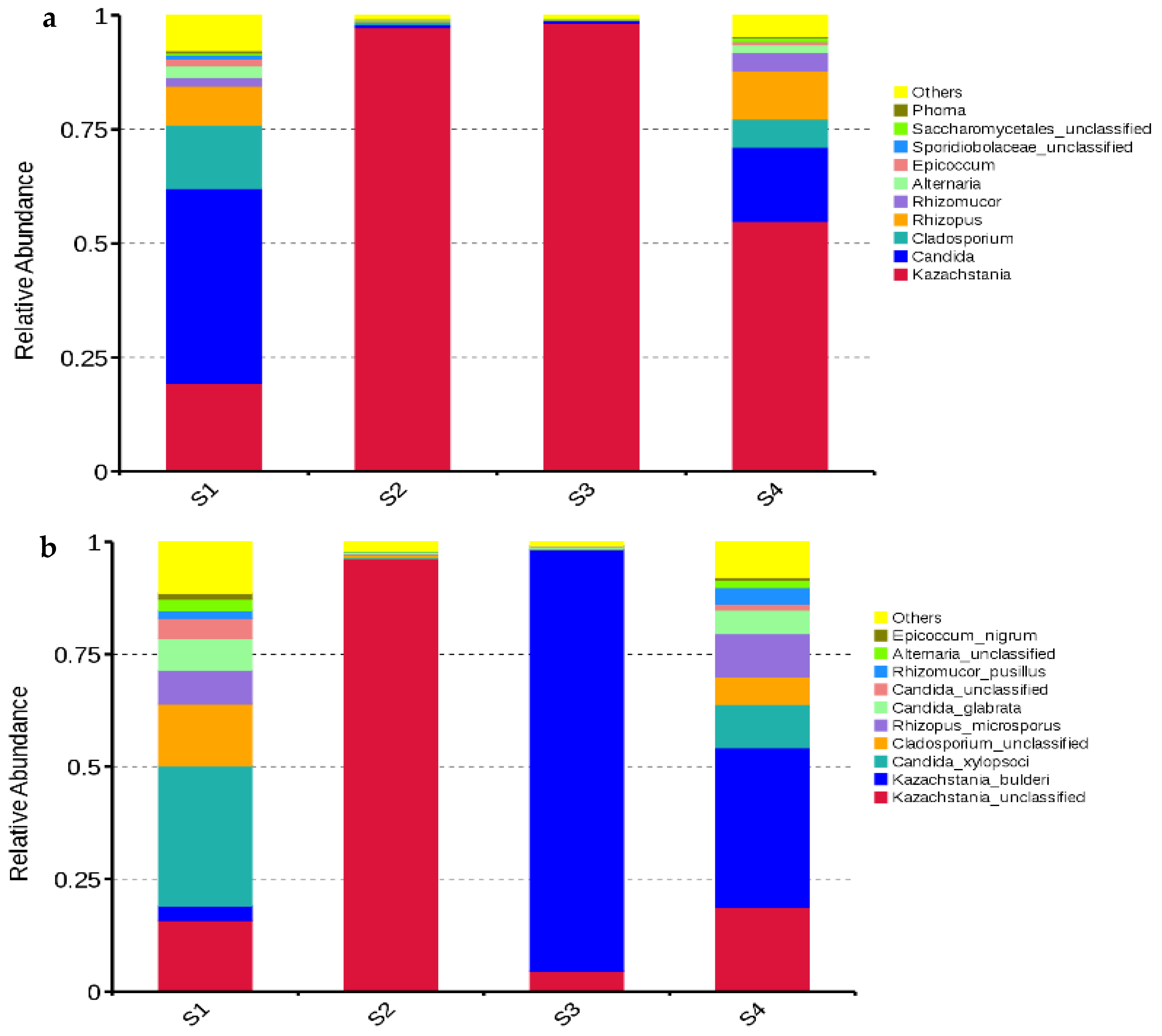
The dominant fungal genus in S2 and S3 was Kazachstania with abundances of 97.32% and 98.24%, respectively; Kazachstania (54.86%) was also dominant in S4, followed by Candida (16.28%), Rhizomucor (10.56%), and Cladosporium (6.15%). The main fungal genera in S1 were Candida (42.73%), Kazachstania (19.34%), Cladosporium (13.90%), and Rhizopus (8.52%) (Figure 4a). The main fungal species in S1 and S4 were unclassified Kazachstania (15.84% and 18.72%, respectively), Kazachstania bulderi (3.13% and 35.53%, respectively), Candida xylopsoci (31.19% and 9.64%, respectively), unclassified Cladosporium (13.79% and 6.14%, respectively), Rhizopus microsporus (7.54% and 9.60%, respectively), and Candida glabrata (7.03% and 5.25%, respectively). The dominant fungal species in S2 was unclassified Kazachstania (96.21%), while Kazachstania bulderi (93.61%) dominated in S3 (Figure 4b).
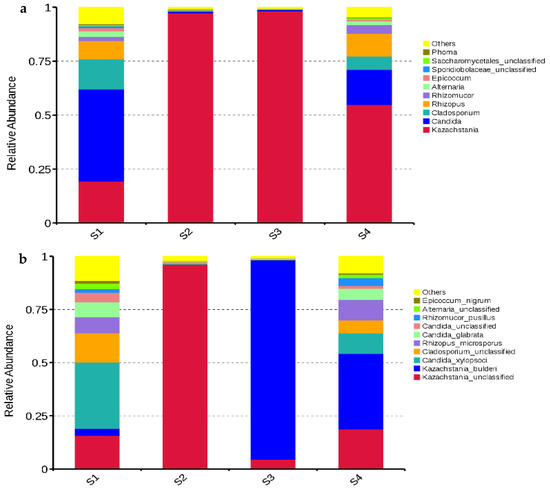
Figure 4.
Relative abundances of fungal genera (a) and species (b) in whole-plant corn silage (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot.
3.5. Metabolites
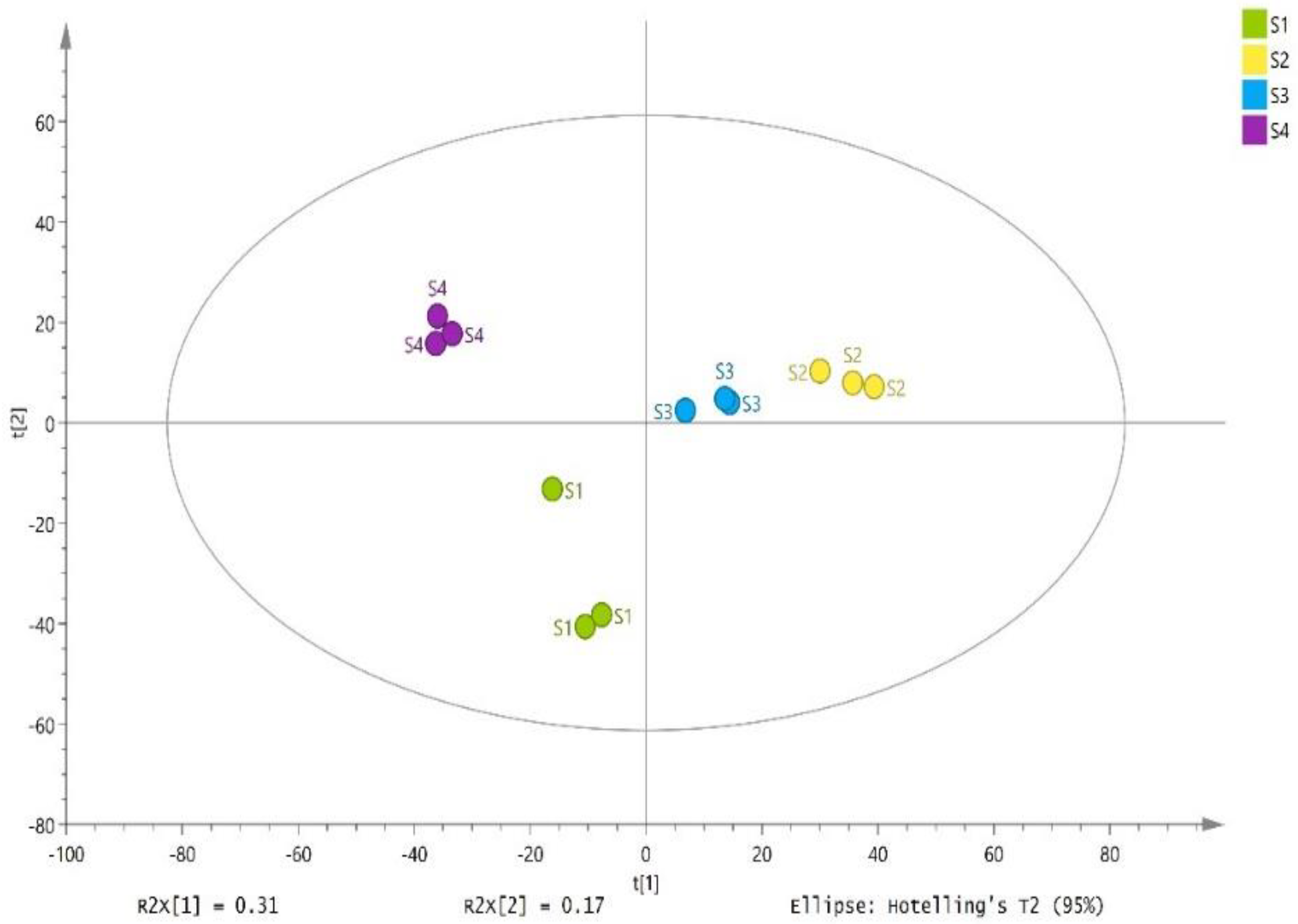
The silages collected from different sites (Erdos, Baotou, Ulanqab, and Hohhot) were clearly separated by PC1 and PC2 with 31% and 17% variation, respectively according to principal component analysis (Figure 5). A total of 668 substances were detected in 12 samples of whole-plant corn silages, which 292 substances were identified (Supplementary Table S1). The S2 had greater relative concentrations of total metabolites than other silages (p < 0.05) (Table 4). Five biogenic amines were detected in the whole-plant corn silages. The S2 showed the greatest abundances of phenylethylamine and spermidine among the silages (p < 0.05) (Table 4).
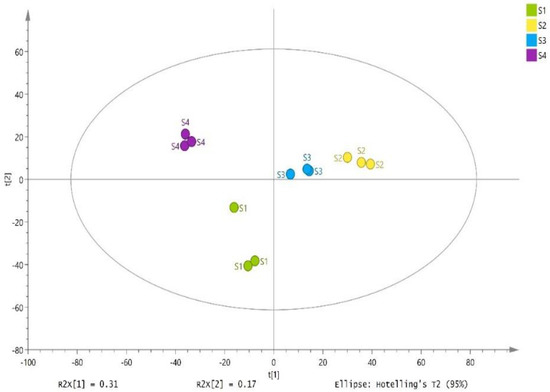
Figure 5.
Principal component analysis of the metabolites in whole-plant corn silages (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot.

Table 4.
Relative concentrations of total metabolites and biogenic amines in whole-plant corn silage (n = 3).
3.6. Fermentation Quality and Chemistry Composition
At 0 d of aerobic exposure, S1 and S4 displayed higher pH and acetic acid, but lower lactic acid, than S2 and S3 (p < 0.05); S1 contained more lactic acid and less acetic acid than S4 (p < 0.05). The S3 contained less ammonia nitrogen than other silages (p < 0.05), and S1 and S2 contained less than S4 (p < 0.05). Propionic and butyric acids were not detected in the silages at 0 d of aerobic exposure (Table 5). The contents of lactic acid and ammonia nitrogen (except for in S4) decreased (p < 0.05) by aerobic exposure, but pH increased (p < 0.05) except for in S1. The acetic acid in S2 and S4 decreased (p < 0.05), while it decreased in S3 at 2 d and then increased at 5 d (p < 0.05). The aerobic exposure time and sampling area had single and interactive effects on the pH and the contents of lactic acid, acetic acid, propionic acid, and ammonia nitrogen in the silages (p < 0.05) (Table 5).

Table 5.
pH, organic acid contents (g/kg dry matter) and ammonia nitrogen contents (g/kg total nitrogen) of whole-plant corn silages during aerobic exposure (n = 3).
The DM content in S1 was higher than that in other silages (p < 0.05). S2 and S3 had greater CP than S1 and S4, with S1 showing higher than S4 (p < 0.05). The NDF concentration in S4 was greater than that in other silages (p < 0.05) (Table 6).

Table 6.
Dry matter content (DM, g/kg) and chemical component concentrations (g/kg DM) of whole-plant corn silages (n = 3).
3.7. Aerobic Stability
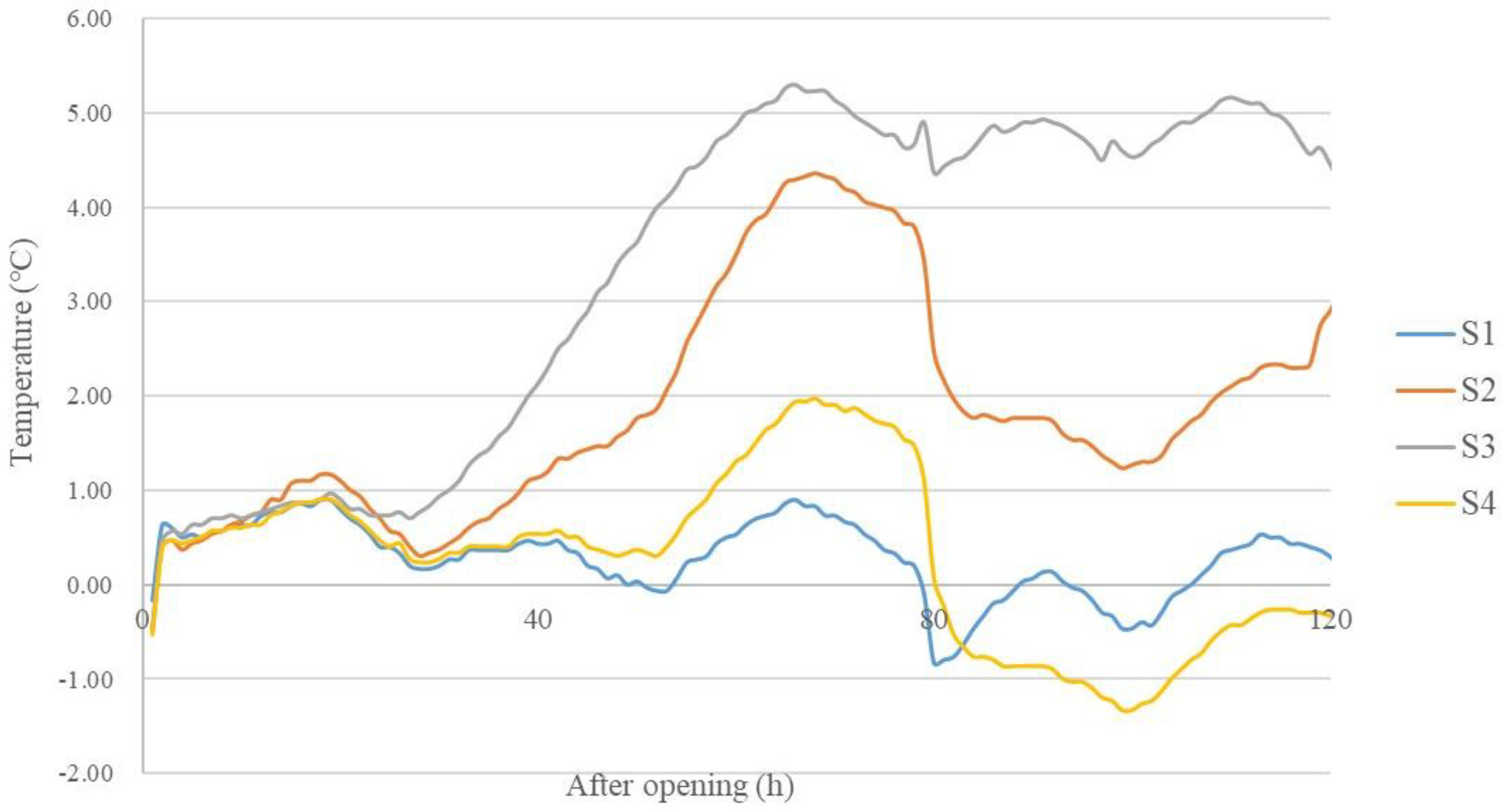
The temperatures of S1 and S4 did not exceed 2 °C above ambient temperature for 5 days after opening. The temperatures of S2 and S3 exceeded 2 °C above ambient temperature at 53 h and 39 h, respectively (Figure 6).
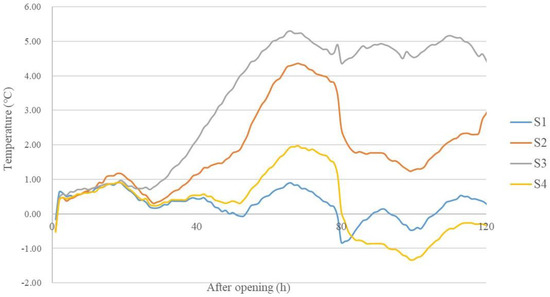
Figure 6.
Temperature dynamics of silage reaching above-ambient temperature during aerobic exposure (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot.
4. Discussion
In the present study, the materials displayed sufficient LAB counts (>5 log10 CFU/g FW) and WSC concentrations (>97 g/kg DM) (Table 1), resulting in a satisfactory fermentation quality for the whole-plant corn silages (Table 5). Before ensiling, S2 and S3 contained greater LAB and WSC contents and a lower DM content than S1 and S4 pre-ensiling (Table 1), indicating the conditions of the former two were more conducive to LAB fermentation during the ensiling process. This contributed to the lower pH and higher lactic acid contents in S2 and S3 (Table 5).
As the microbial community composition plays a crucial role in the fermentation quality and aerobic stability of silage, it is necessary to assess the bacterial and fungal communities in order to explain the different fermentation quality and aerobic stability among silages. In the present study, Lactobacillus was the most dominant bacterial genus in all silages, followed by Acinetobacter in S1 and S3, and by Acinetobacter, Novosphingobium, Afipia, Clostridium, and unclassified Actinobacteria in S2 (Figure 3a). This indicated that Lactobacillus is generally the dominant bacterial genus in whole-plant corn silages after ensiling for about 300 d. Moreover, previous studies also reported Lactobacillus at the greatest abundance in whole-plant corn silages collected from five regions in Southwest China [9] and three sites in Iran [8]. In silage with a good fermentation quality, the bacterial community is dominated by Lactobacillus [21], explaining the satisfactory fermentation quality of whole-plant corn silages in our study. Acinetobacter was found in sugarcane top silage [22], and Novosphingobium was detected in whole-plant corn silage [23] and red clover silage [24]. However, the effect of these genera to fermentation quality and aerobic stability are not clear and require further research.
In the present study, the main bacterial species in S2 were L. buchneri, Acinetobacter johnsonii, and unclassified Novosphingobium with abundances of 20.7%, 10.7% and 7.64%, respectively (Figure 3b). Lactobacillus buchneri, L. parafarraginis, L. kefiri, and unclassified Lactobacillus dominated the bacterial communities in S1, S3, and S4 with total abundances of 84.5%, 88.2%, and 98.3%, respectively (Figure 3b). The main LAB species in the silages after ensiling for about 300 d were L. buchneri, L. parafarraginis, L. kefiri, and L. diolivorans (Figure 3b), which belong to L. buchneri group as the sole heterofermentative LAB [25]. However, in previous studies, the main LAB species were L. acetotolerans, L. silagei, L. buchneri, L. odoratitofui, L. farciminis, and L. parafarraginis in whole-plant corn silages (90 d) with low pH (from 3.68 to 3.74) [7], L. plantarum, L. buchneri, and L. brevisi in alfalfa silage (90 d) with high pH (from 4.9 to 5.2) [26], L. plantarum, L. hammesi, L. brevisi, L. coryniformis, and L. piscium in Italian ryegrass silages (42 d) with pH ranging from 4.40 to 5.52 [10]. These studies suggested that homofermentative LAB co-existed with heterofermentative LAB in short silage (42–90 d). The differences in LAB population between our study and previous studies might result from the differences in silage age (300 d vs. 42–90 d), in the epiphytic population of bacteria on materials, and in the quality of the silages. The effects of short- and long-term storage on the microbial community and fermentation quality of silage requires further study. In the present study, the activities of heterofermentative LAB were weak in S2 and S3 under more acidic conditions (pH = 3.61 and 3.63, respectively) for a long period (about 300 d after ensiling), while they were stronger in S1 and S4 (pH 3.97). This is reflected in the higher acetic acid contents and LAB counts, and the lower lactic acid concentrations, in the latter at 0 and 2 d, along with greater aerobic stability (Table 5). The bacteria count decreased in S2 in the first 2 d, while it increased in S3 and S4 (Table 2). This might be because S2 still had a lower pH (3.69) and higher lactic acid content (121 g/kg DM) at 2 d than S1, S3, and S4 (Table 5). More acidic conditions might have reduced the bacterial activity in the silage, although the silage had been exposed to air for 2 d. Acinetobacter johnsonii, a known spoilage organism [27], was one of the more dominant bacterial species in S1 and S2, and it was present in S3 and S4 at low levels (Figure 3b). The effect of A. johnsonii on the fermentation quality and safety of silage requires further study.
The main fungal genera in S1 and S4 were Kazachstania, Candida, Cladosporium, Rhizopus, Rhizomucor, Alternaria, and Epicoccum, whereas Kazachstania dominated the fungal communities in S2 and S3 (Figure 4a). Other studies have reported dominant fungal genera of Candida and Monascus in whole-plant corn silage (90 d) [28], Pichia, Sporisorium, Meyerozyma, and Hannaella in sugarcane top silage (60 d) [22], Issatchenki in barley silage (60 d) [29], and Candida, Kazachstania, and Pichia in sugarcane top silage (90 d) [15]. These results suggest that, in contrast to the bacterial community, the dominant fungal community in the silage was different between the silages at the genera level. In the present study, Kazachstania dominated the fungal communities in S2 and S3 with poor aerobic stability (Figure 5). S2 and S3 had lower pH and acetic acid, and higher lactic acid than the other silages (Table 5). During aerobic exposure, the pH increased and the lactic acid decreased in S3, with acetic acid reducing at 2 d and increasing rapidly at 5 d. Between 2 and 5 d in S2, the pH increased quickly and the lactic acid decreased rapidly, with acetic acid disappearing at 5 d (Table 5). These results agree with the findings of Wang et al. for sugarcane top silage [15]. The results above indicate that Kazachstania might have a strong tolerance to lactic acid and be a yeast species that is crucially involved in initiating the aerobic deterioration of silage with a relatively low pH and acetic acid content. Kazachstania was also detected in barley silage [29] and sugarcane top silage [22] as minor taxa. Candida was the predominant fungal genus in S1 and S4 (Figure 4a), which had greater aerobic stability and higher acetic acid content (Table 5), and lower temperatures, than other silages (Figure 6) during aerobic exposure. However, Liu et al. [29] and Romero et al. [30] reported that a high abundance of Candida sp. was associated with a lower aerobic stability in silage, owing to Candida assimilating lactic acid after aerobic exposure [28]. The difference between our results and those of previous studies might be due to the higher acetic acid concentration in our study (42.8 g/kg) than in the studies of Romero et al. [4] and Liu et al. [29] (7.4 g/kg and 18.3 g/kg, respectively)—acetic acid inhibits fungus growth in silage and can enhance aerobic stability [4,28,30,31].
In the present study, the dominant fungal species in S1 and S4 were unclassified Kazachstania, K. bulderi, C. xylopsoci, unclassified Cladosporium, Rhizopus microspores, and C. glabrata with total abundances of 78.5% and 84.9% in fungal communities, respectively (Figure 4b). Unclassified Kazachstania and K. bulderi dominated the fungal communities in S2 and S3, respectively with abundances of 96.2% in S2 and 93.6% in S3 (Figure 4b). According to PCoA, the fungal species in S1 and S4 were connected, and were clearly separated from those of S2 and S3 (Figure 2). S1 and S4 thus had similar fungal species compositions, which were differentiated from those of S2 and S3. The differences in fungal species might be due to the differences in the fermentation processes of silage and in the epiphytic populations of fungi in the plant prior to ensiling [28]. Moreover, unclassified Kazachstania, and K. bulderi belonging to Kazachstania, were the dominant fungi in S2 and S3, respectively (Figure 4b), and they had poor aerobic stability (Figure 6). This indicates that the aerobic deterioration taking place in the two silages might result from Kazachstania sp. growth after opening. Wang et al. [15] found K. humilis to be the key fungal yeast initiating aerobic deterioration in sugarcane top silage. Kazachstania bulderi was the predominant fungus in S3 and S4 in the present study. This strain was formerly named Saccharomyces bulderi [32] and was first isolated from maize silage as a novel species [33]. The present study first detected C. xylopsoci, R. microspores, and Rhizomucor pusillus in the silage in the form of fungal pathogens. Candida xylopsoci is usually isolated via fuel ethanol fermentation processes [34], R. microsporus has been detected in the traditional Chinese liquor Daqu using 18S rRNA gene sequencing [35], and R. pusillus has been identified in immunocompromised patients using real-time PCR [36]. The effects of these fungi on the fermentation quality and aerobic stability of silage should be investigated in future studies.
Moreover, Drouin et al. [10] revealed that ensiling whole-plant corn with LAB inoculant improved the aerobic stability of the silage, by maintaining a higher microbial diversity (Shannon index of bacteria and fungi), avoiding the dominance of a few bacteria, and preventing fungi from damaging silage quality. However, in the present study, the fungal communities in S1 and S4 had higher Shannon, Simpson, and Chao1 indexes, and greater aerobic stability, than S2 and S3, but the bacterial communities in S1, S3, and S4 had lower Shannon, Simpson, and Chao1 indexes than S2 (Table 3, Figure 6). This suggests that high fungal diversity, rather than bacterial diversity, contributes most significantly to aerobic stability in whole-plant corn silage.
S2 displayed the highest bacterial Shannon and Simpson indexes among the silages (Table 3), suggesting S2 had greater bacterial diversity in the present study. Additionally, the relative concentration of total metabolites in S2 was the highest among the silages (Table 4), and containing 93, 103, and 81 more metabolites identified than S1, S3, and S4, respectively (Supplementary Tables S2–S4). Previous studies showed that inoculating with LAB at ensiling increased the relative concentrations of some metabolites (e.g., organic acids, amino acids, and fatty acids) as well as the α-diversity (Shannon index) in whole-plant corn silages, sainfoin silages, and alfalfa silages [6,7,9,12]. This indicates that the high bacterial diversity might contribute to the accumulation of metabolites in whole-plant corn silages during fermentation process.
Biogenic amines in silages are produced by the decarboxylation of amino acids via the activities of plant enzymes and microbial enzymes, which accumulate during fermentation process and influence the palatability of silages, as well as the feed intake and performance of ruminants [14,37,38]. The intake of adequate amounts of biogenic amines can promote normal physiological activities in man and animals, but excessive amounts may result in food poisoning [38]. In the present study, tyramine, putrescine, phenylethylamine, spermidine, and noradrenaline were detected in whole-plant corn silages; the relative concentrations of tyramine and putrescine were considerable in S1, S2 and S3. Steidlová and Kalac also found that the concentrations of tyramine and putrescine were greater than those of histamine, cadaverine, tryptamine, spermidine and spermine in maize silages in 1999 (62 silages) and 2000 (51 silages) [14]. Nishino et al. [37] reported that inoculating with Lactobacillus casei reduced the concentrations of biogenic amines (histamine, tyramine, putrescine and cadaverine) in grass silage, maize silage, and total mixed ration silage, and that adding L. buchneri lowered the contents of biogenic amines in grass silages. Additionally, the activities of some undesirable bacterial genera (Clostridia, Bacillus, Klebsiella, Escherichia, Pseudomonas, Citrobacter, Proteus, Salmonella, Shigella, and Photobacterium) might result in the accumulation of biogenic amines in silages during ensiling [38,39]. In the present study, S2 contained higher Bacillus, Pseudomonas, and Salmonella contents than other silages, and greater Escherichia and Klebsiella than S3 and S4 (Supplementary Table S5), which might explain the higher relative concentrations of biogenic amines in S2 (Table 4).
5. Conclusions
Whole-plant corn silages showed a satisfactory fermentation quality after 300 d of ensiling. The main bacterial species in S2 were L. buchneri, Acinetobacter johnsonii, and unclassified Novosphingobium, while L. buchneri, L. parafarraginis, L. kefiri, and unclassified Lactobacillus dominated the bacterial communities in S1, S3, and S4. The main fungal species in S1 and S4 were unclassified Kazachstania, K. bulderi, C. xylopsoci, unclassified Cladosporium, R. microspores, and C. glabrata, while unclassified Kazachstania and K. bulderi dominated the fungal communities in S2 and S3, respectively, and were associated with these silages’ poor aerobic stability. In whole-plant corn silage, high bacterial diversity helps the accumulation of metabolites, and high fungal diversity contributes to an improved aerobic stability.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/pr9050784/s1, Supplementary Table S1. Relative concentration of metabolites detected in whole-plant corn silages (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot. (page: 1–16). Supplementary Table S2. Difference in metabolites identified between S1 and S2 (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou. (page: 17–19). Supplementary Table S3. Difference in metabolites identified between S2 and S3 (n = 3). S2, silage collected from Baotou; S3, silage collected from Ulanqab. (page: 20–22). Supplementary Table S4. Difference in metabolites identified between S2 and S4 (n = 3). S2, silage collected from Baotou; S4, silage collected from Hohhot. (page: 23–25). Supplementary Table S5. Relative abundance of Bacillus, Pseudomonas, Escherichia, Klebsiella, and Salmonella in whole-plant corn silages (n = 3). S1, silage collected from Erdos; S2, silage collected from Baotou; S3, silage collected from Ulanqab; S4, silage collected from Hohhot. (page: 26).
Author Contributions
Conceptualization, C.W. and Y.X.; methodology, C.W.; software, L.S.; validation, C.W. and L.S.; formal analysis, L.S., H.X., and N.N.; investigation, C.W. and L.S.; resources, G.Y. and S.L.; data curation, S.L., H.X., and N.N.; writing—original draft preparation, C.W.; writing—review and editing, H.X., Y.J., and Y.X.; visualization, G.Y.; supervision, Y.X.; project administration, Y.X.; funding acquisition, Y.X. All authors have read and agreed to the published version of the manuscript.
Funding
This study was funded by the National Key R&D Program of China (grant number, 2017YFE0104300) and the Science and Technology Project of Inner Mongolia (grant number, 2020GG0049).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Gong, T.; Battese, G.E.; Villano, R.A. Family farms plus cooperatives in China: Technical efficiency in crop production. J. Asian Econ. 2019, 64, 101129. [Google Scholar] [CrossRef]
- Sachurina. The change of the pastoral management mode in Inner Mongolia: Joint households, cooperatives, household pastures and corporations. J. Arid Land Resour. Environ. 2017, 31, 56–63. [Google Scholar] [CrossRef]
- Wang, R.Z. Rational Forage and Livestock Allocation of Household Ranch in Inner Mongolia Pastoral Area. Master’s Thesis, Inner Mongolia Agricultural University, Hohhot, China, 2017. [Google Scholar]
- Romero, J.J.; Joo, Y.; Park, J.; Tiezzi, F.; Gutierrez-Rodriguez, E.; Castillo, M.S. Bacterial and fungal communities, fermentation, and aerobic stability of conventional hybrids and brown midrib hybrids ensiled at low moisture with or without a homo- and heterofermentative inoculant. J. Dairy Sci. 2018, 101, 3057–3076. [Google Scholar] [CrossRef]
- Sun, L.; Bai, C.; Xu, H.; Na, N.; Jiang, Y.; Yin, G.; Liu, S.; Xue, Y. Succession of bacterial community during the initial aerobic, intense fermentation, and stable phases of whole-plant corn silages treated with lactic acid bacteria suspensions prepared from other silages. Front. Microbiol. 2021, 12, 655095. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Wang, N.; Rinne, M.; Ke, W.; Weinberg, Z.G.; Da, M.; Bai, J.; Zhang, Y.; Li, F.; Guo, X. The bacterial community and metabolome dynamics and their interactions modulate fermentation process of whole crop corn silage prepared with or without inoculants. Microb. Biotechnol. 2020. [Google Scholar] [CrossRef]
- Xu, D.; Ding, W.; Ke, W.; Li, F.; Zhang, P.; Guo, X. Modulation of metabolome and bacterial community in whole crop corn silage by inoculating homofermentative Lactobacillus plantarum and heterofermentative Lactobacillus buchneri. Front. Microbiol. 2019. [Google Scholar] [CrossRef]
- Gharechahi, J.; Kharazian, Z.A.; Sarikhan, S.; Jouzani, G.S.; Aghdasi, M.; Salekdeh, G.H. The dynamics of the bacterial communities developed in maize silage. Microb. Biotechnol. 2017, 10, 1663–1676. [Google Scholar] [CrossRef]
- Guan, H.; Yan, Y.; Li, X.; Li, X.; Shuai, Y.; Feng, G.; Ran, Q.; Cai, Y.; Li, Y.; Zhang, X. Microbial communities and natural fermentation of corn silages prepared with farm bunker-silo in Southwest China. Bioresour. Technol. 2018, 265, 282–290. [Google Scholar] [CrossRef]
- Drouin, P.; Tremblay, J.; Renaud, J.; Apper, E. Microbiota succession during aerobic stability of maize silage inoculated with Lentilactobacillus buchneri NCIMB 40788 and Lentilactobacillus hilgardii CNCM-I-4785. MicrobiologyOpen 2020, 10, e1153. [Google Scholar] [CrossRef]
- Wilkinson, J.M.; Muck, R.E. Ensiling in 2050: Some challenges and opportunities. Grass Forage sci. 2019, 74, 178–187. [Google Scholar] [CrossRef]
- Xu, D.; Ding, Z.; Wang, M.; Bai, J.; Ke, W.; Zhang, Y.; Guo, X. Characterization of the microbial community, metabolome and biotransformation of phenolic compounds of sainfoin (Onobrychis viciifolia) silage ensiled with or without inoculation of Lactobacillus plantarum. Bioresour. Technol. 2020, 316, 123910. [Google Scholar] [CrossRef]
- Wu, Z.; Luo, Y.; Bao, J.; Luo, Y.; Yu, Z. Additives affect the distribution of metabolic profile, microbial communities and antibiotic resistance genes in high-moisture sweet corn kernel silage. Bioresour. Technol. 2020, 315, 123821. [Google Scholar] [CrossRef] [PubMed]
- Steidlová, Š.; Kalac, P. Levels of biogenic amines in maize silages. Anim. Feed Sci. Technol. 2002, 102, 197–205. [Google Scholar] [CrossRef]
- Wang, T.; Teng, K.; Cao, Y.; Shi, W.; Xuan, Z.; Zhou, J.; Zhang, J.; Zhong, J. Effects of Lactobacillus hilgardii 60TS-2, with or without homofermentative Lactobacillus plantarum B90, on the aerobic stability, fermentation quality and microbial community dynamics in sugarcane top silage. Bioresour. Technol. 2020, 312, 123600. [Google Scholar] [CrossRef]
- Owens, V.N.; Albrecht, K.A.; Muck, R. EProtein degradation and ensiling characteristics of red clover and alfalfa wilted under varying levels of shade. Can. J. Plant Sci. 1999, 79, 209–222. [Google Scholar] [CrossRef]
- Cai, Y. Identification and characterization of Enterococcus species isolated from forage crops and their influence on silage fermentation. J. Dairy Sci. 1999, 82, 2466–2471. [Google Scholar] [CrossRef]
- Zhang, Q.; Li, X.J.; Zhao, M.M.; Yu, Z. Isolating and evaluating lactic acid bacteria strains for effectiveness of Leymus chinensis silage fermentation. Lett. Appl. Microbiol. 2014, 59, 391–397. [Google Scholar] [CrossRef]
- Van Soest, P.J.; Robertson, J.B.; Lewis, B.A. Methods for dietary fiber neutral detergent fiber and nonstarch polysaccharides in relation to animal nutrition. J. Dairy Sci. 1991, 74, 3583–3594. [Google Scholar] [CrossRef]
- Yan, N.; Du, Y.; Liu, X.; Chu, M.; Shi, J.; Zhang, H.; Liu, Y. A comparative UHPLCQqQ-MS-based metabolomics approach for evaluating Chinese and North American wild rice. Food Chem. 2019, 275, 618–627. [Google Scholar] [CrossRef] [PubMed]
- McEniry, J.; O’Kiely, P.; Clipson, N.; Forristal, P.; Doyle, E. Bacterial community dynamics during the ensilage of wilted grass. J. Appl. Microbiol. 2008, 105, 359–371. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhou, X.; Gu, Q.; Liang, M.; Mu, S.; Zhou, B.; Huang, F.; Lin, B.; Zou, C. Analysis of the correlation between bacteria and fungi in sugarcane tops silage prior to and after aerobic exposure. Bioresour. Technol. 2019, 291, 121835. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, Y.; Meng, Q.; Zhou, Z.; Wu, H. A mixture of potassium sorbate and sodium benzoate improved fermentation quality of whole-plant corn silage by shifting bacterial communities. J. Appl. Microbiol. 2020, 128, 1312–1323. [Google Scholar] [CrossRef] [PubMed]
- Ali, N.; Wang, S.; Zhao, J.; Dong, Z.; Li, J.; Nazar, M.; Shao, T. Microbial diversity and fermentation profile of red clover silage inoculated with reconstituted indigenous and exogenous epiphytic microbiota. Bioresour. Technol. 2020, 123606. [Google Scholar] [CrossRef] [PubMed]
- Muck, R.E.; Nadeau, E.M.G.; McAllister, T.A.; Contreras-Govea, F.E.; Santos, M.C.; Kung, L. Silage review: Recent advances and future uses of silage additives. J. Dairy Sci. 2018, 101, 3980–4000. [Google Scholar] [CrossRef]
- Guo, X.S.; Ke, W.C.; Ding, W.R.; Ding, L.M.; Xu, D.M.; Wang, W.W.; Zhang, P.; Yang, F.Y. Profiling of metabolome and bacterial community dynamics in ensiled Medicago sativa inoculated without or with Lactobacillus plantarum or Lactobacillus buchneri. Sci. Rep. 2018, 8, 357. [Google Scholar] [CrossRef]
- Wang, X.Y.; Xie, J. Assessment of metabolic changes in Acinetobacter johnsonii and Pseudomonas fluorescens co-culture from bigeye tuna (Thunnus obesus) spoilage by ultrahigh-performance liquid chromatography-tandem mass spectrometry. LWT Food Sci. Technol. 2020, 123, 109073. [Google Scholar] [CrossRef]
- Keshri, J.; Chen, Y.; Pinto, R.; Kroupitski, Y.; Weinberg, Z.G.; Sela, S. Microbiome dynamics during ensiling of corn with and without Lactobacillus plantarum inoculant. Appl. Microbiol. Biot. 2018, 102, 4025–4037. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Huan, H.; Gu, H.; Xu, N.; Shen, Q.; Ding, C. Dynamics of a microbial community during ensiling and upon aerobic exposure in lactic acid bacteria inoculation-treated and untreated barley silages. Bioresour. Technol. 2019, 273, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Romero, J.J.; Zhao, Y.; Balseca-Paredes, M.A.; Tiezzi, F.; Gutierrez-Rodriguez, E.; Castillo, M.S. Laboratory silo type and inoculation effects on nutritional composition, fermentation, and bacterial and fungal communities of oat silage. J. Dairy Sci. 2017, 100, 1812–1828. [Google Scholar] [CrossRef]
- Li, Y.; Nishino, N. Effects of inoculation of Lactobacillus rhamnosus and Lactobacillus buchneri on fermentation, aerobic stability and microbial communities in whole crop corn silage. Grassl. Sci. 2011, 57, 184–191. [Google Scholar] [CrossRef]
- Boyaci-Gunduz, C.P.; Erten, H. Predominant yeasts in the sourdoughs collected from some parts of Turkey. Yeast 2020, 37. [Google Scholar] [CrossRef] [PubMed]
- Middelhoven, W.J.; Kurtzman, C.P.; Vaughan-Martini, A. Saccharomyces bulderi sp. nov., a yeast that ferments gluconolactone. Antonie Van Leeuwenhoek 2000, 77, 223–228. [Google Scholar] [CrossRef] [PubMed]
- Basilio, A.C.M.; de Araujo, P.R.L.; de Morais, J.O.F.; da Silva Filho, E.A.; de Morais, M.A.; Simoes, D.A. Detection and identification of wild yeast contaminants of the industrial fuel ethanol fermentation process. Curr. Microbiol. 2008, 56, 322–326. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Chen, X.; Guang, J. Bacterial and fungal diversity in the traditional Chinese strong flavour liquor Daqu. J. Inst. Brew. 2019, 125, 443–452. [Google Scholar] [CrossRef]
- Hadaschik, E.; Koschny, R.; Willinger, B.; Hallscheidt, P.; Enk, A.; Hartschuh, W. Pulmonary, rhino-orbital and cutaneous mucormycosis caused by Rhizomucor pusillus in an immunocompromised patient. Clin. Exp. Dermatol. 2012, 37, 355–357. [Google Scholar] [CrossRef]
- Nishino, N.; Hattori, H.; Wada, H.; Touno, E. Biogenic amine production in grass, maize and total mixed ration silages inoculated with Lactobacillus casei or Lactobacillus buchneri. J. Appl. Microbiol. 2007, 103, 325–332. [Google Scholar] [CrossRef] [PubMed]
- Scherer, R.; Gerlach, K.; Südekum, K.H. Biogenic amines and gamma-amino butyric acid in silages: Formation, occurrence and influence on dry matter intake and ruminant production. Anim. Feed Sci. Technol. 2015, 210, 1–16. [Google Scholar] [CrossRef]
- Santos, M.H.S. Biogenic amines: Their importance in foods. Int. J. Food Microbiol. 1996, 29, 213–231. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).