Differential Digestive Stability of Food-Derived microRNAs: The Case of miR-30c-5p and miR-92a-3p in Polyfloral Honey
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. In Vitro Digestion of Honey Samples
2.2.1. Pre-Digestion Procedures
2.2.2. INFOGEST Static In Vitro Gastrointestinal Digestion
2.3. Characterization of Exosome-like Nanoparticles in Honey
2.4. Total RNA Extraction from Honey and Its Digests
2.4.1. Column-Based Extraction (Manual)
2.4.2. Semi-Automated Extraction
2.5. Total RNA Quantification
2.6. qRT-PCR of Honey and Its Digests
2.7. Analysis and Visualization
3. Results
3.1. Honey Digestibility
3.2. Variation in Total RNA Yield According to Methods of Extraction
3.3. Vesicles Characterization of the Polyfloral Honey
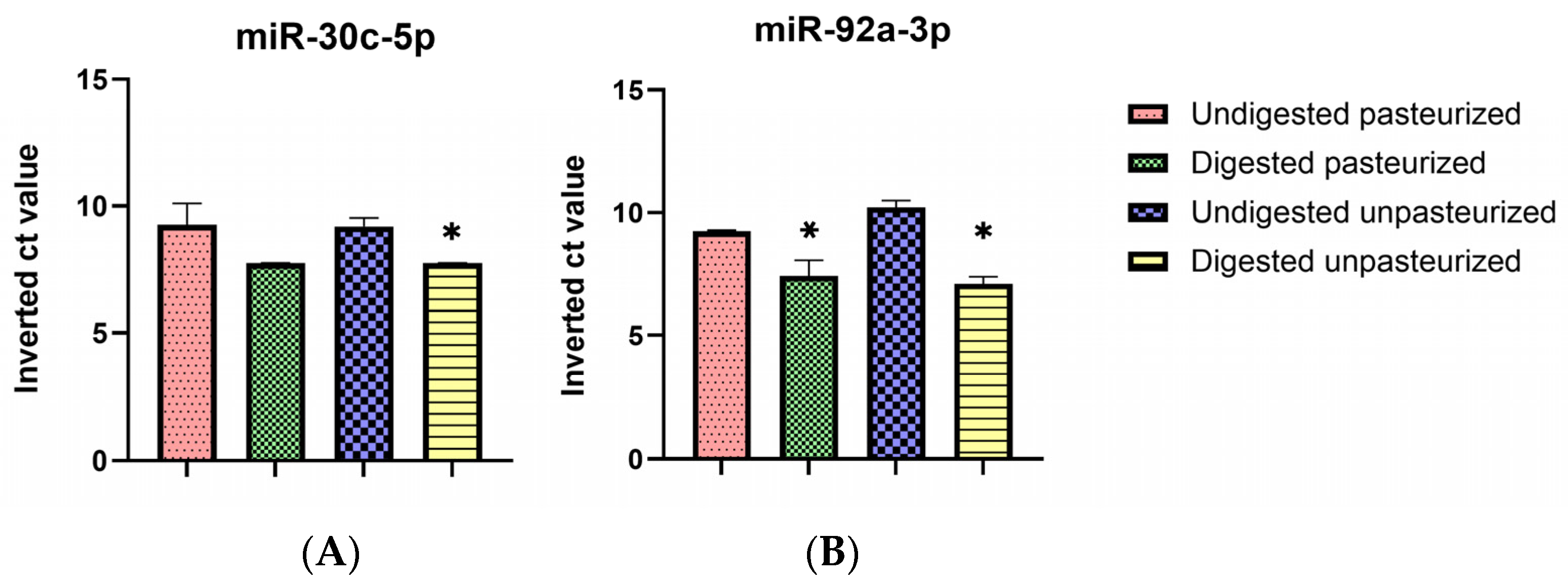
3.4. Selective Digestive Stability of miR-30c-5p and miR-92a-3p
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef] [PubMed]
- Tucci, P.; Agostini, M.; Grespi, F.; Markert, E.K.; Terrinoni, A.; Vousden, K.H.; Muller, P.A.J.; Dötsch, V.; Kehrloesser, S.; Sayan, B.S.; et al. Loss of p63 and its microRNA-205 target results in enhanced cell migration and metastasis in prostate cancer. Proc. Natl. Acad. Sci. USA 2012, 109, 15312–15317. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Sun, C.; Zhao, Y.; Wang, Q.; Guo, J.; Ye, B.; Yu, G. Overview of MicroRNAs as Diagnostic and Prognostic Biomarkers for High-Incidence Cancers in 2021. Int. J. Mol. Sci. 2022, 23, 1389. [Google Scholar] [CrossRef] [PubMed]
- Ho, P.T.B.; Clark, I.M.; Le, L.T.T. MicroRNA-Based Diagnosis and Therapy. Int. J. Mol. Sci. 2022, 23, 7167. [Google Scholar] [CrossRef] [PubMed]
- Baier, S.R.; Nguyen, C.; Xie, F.; Wood, J.R.; Zempleni, J. MicroRNAs Are Absorbed in Biologically Meaningful Amounts from Nutritionally Relevant Doses of Cow Milk and Affect Gene Expression in Peripheral Blood Mononuclear Cells, HEK-293 Kidney Cell Cultures, and Mouse Livers. J. Nutr. 2014, 144, 1495–1500. [Google Scholar] [CrossRef]
- Izumi, H.; Kosaka, N.; Shimizu, T.; Sekine, K.; Ochiya, T.; Takase, M. Bovine milk contains microRNA and messenger RNA that are stable under degradative conditions. J. Dairy Sci. 2012, 95, 4831–4841. [Google Scholar] [CrossRef]
- Sanchita; Trivedi, R.; Asif, M.H.; Trivedi, P.K. Dietary plant miRNAs as an augmented therapy: Cross-kingdom gene regulation. RNA Biol. 2018, 15, 1433–1439. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Li, L.; Jia, Y.; Xu, T.; Zhou, X. Advances in studies of circulating microRNAs: Origination, transportation, and distal target regulation. J. Cell Commun. Signal. 2023, 17, 445–455. [Google Scholar] [CrossRef]
- Zhang, L.; Hou, D.; Chen, X.; Li, D.; Zhu, L.; Zhang, Y.; Li, J.; Bian, Z.; Liang, X.; Cai, X.; et al. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res. 2012, 22, 107–126. [Google Scholar] [CrossRef]
- Kosaka, N.; Izumi, H.; Sekine, K.; Ochiya, T. microRNA as a new immune-regulatory agent in breast milk. Silence 2010, 1, 7. [Google Scholar] [CrossRef] [PubMed]
- Hatmal, M.M.; Al-Hatamleh, M.A.I.; Olaimat, A.N.; Alshaer, W.; Hasan, H.; Albakri, K.A.; Alkhafaji, E.; Issa, N.N.; Al-Holy, M.A.; Abderrahman, S.M.; et al. Immunomodulatory Properties of Human Breast Milk: MicroRNA Contents and Potential Epigenetic Effects. Biomedicines 2022, 10, 1219. [Google Scholar] [CrossRef] [PubMed]
- Martino, E.; D’Onofrio, N.; Balestrieri, A.; Colloca, A.; Anastasio, C.; Sardu, C.; Marfella, R.; Campanile, G.; Balestrieri, M.L. Dietary Epigenetic Modulators: Unravelling the Still-Controversial Benefits of miRNAs in Nutrition and Disease. Nutrients 2024, 16, 160. [Google Scholar] [CrossRef] [PubMed]
- Cavalieri, D.; Rizzetto, L.; Tocci, N.; Rivero, D.; Asquini, E.; Si-Ammour, A.; Bonechi, E.; Ballerini, C.; Viola, R. Plant microRNAs as novel immunomodulatory agents. Sci. Rep. 2016, 6, 25761. [Google Scholar] [CrossRef] [PubMed]
- Díez-Sainz, E.; Lorente-Cebrián, S.; Aranaz, P.; Riezu-Boj, J.I.; Martínez, J.A.; Milagro, F.I. Potential Mechanisms Linking Food-Derived MicroRNAs, Gut Microbiota and Intestinal Barrier Functions in the Context of Nutrition and Human Health. Front. Nutr. 2021, 8, 586564. [Google Scholar] [CrossRef]
- Gismondi, A.; Di Marco, G.; Canini, A. Detection of plant microRNAs in honey. PLoS ONE 2017, 12, e0172981. [Google Scholar] [CrossRef] [PubMed]
- Pulidindi, K.; Dhiman, M. Honey Market Size & Share|Growth Outlook 2024–2032; Global Market Insights Inc.: Selbyville, DE, USA, 2023. [Google Scholar]
- Masad, R.J.; Haneefa, S.M.; Mohamed, Y.A.; Al-Sbiei, A.; Bashir, G.; Fernandez-Cabezudo, M.J.; Al-Ramadi, B.K. The Immunomodulatory Effects of Honey and Associated Flavonoids in Cancer. Nutrients 2021, 13, 1269. [Google Scholar] [CrossRef] [PubMed]
- El-Seedi, H.R.; Eid, N.; Abd El-Wahed, A.A.; Rateb, M.E.; Afifi, H.S.; Algethami, A.F.; Zhao, C.; Al Naggar, Y.; Alsharif, S.M.; Tahir, H.E.; et al. Honey Bee Products: Preclinical and Clinical Studies of Their Anti-Inflammatory and Immunomodulatory Properties. Front. Nutr. 2022, 8, 761267. [Google Scholar] [CrossRef]
- Badolato, M.; Carullo, G.; Cione, E.; Aiello, F.; Caroleo, M.C. From the hive: Honey, a novel weapon against cancer. Eur. J. Med. Chem. 2017, 142, 290–299. [Google Scholar] [CrossRef]
- Behura, S.K.; Whitfield, C.W. Correlated expression patterns of microRNA genes with age-dependent behavioural changes in honeybee. Insect Mol. Biol. 2010, 19, 431–439. [Google Scholar] [CrossRef]
- Greenberg, J.K.; Xia, J.; Zhou, X.; Thatcher, S.R.; Gu, X.; Ament, S.A.; Newman, T.C.; Green, P.J.; Zhang, W.; Robinson, G.E.; et al. Behavioral plasticity in honey bees is associated with differences in brain microRNA transcriptome. Genes Brain Behav. 2012, 11, 660–670. [Google Scholar] [CrossRef] [PubMed]
- Al-Nakhle, H.; Burns, P.A.; Cummings, M.; Hanby, A.M.; Hughes, T.A.; Satheesha, S.; Shaaban, A.M.; Smith, L.; Speirs, V. Estrogen Receptor β1 Expression Is Regulated by miR-92 in Breast Cancer. Cancer Res. 2010, 70, 4778–4784. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, A.; Ohno, S.; Wu, W.; Borjigin, N.; Fujita, K.; Aoki, T.; Ueda, S.; Takanashi, M.; Kuroda, M. miR-92 is a key oncogenic component of the miR-17–92 cluster in colon cancer. Cancer Sci. 2011, 102, 2264–2271. [Google Scholar] [CrossRef] [PubMed]
- Valera, V.A.; Walter, B.A.; Linehan, W.M.; Merino, M.J. Regulatory Effects of microRNA-92 (miR-92) on VHL Gene Expression and the Hypoxic Activation of miR-210 in Clear Cell Renal Cell Carcinoma. J. Cancer 2011, 2, 515–526. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Yu, S.; Ren, P.; Sun, X.; Jin, M. Human microRNA-30 inhibits influenza virus infection by suppressing the expression of SOCS1, SOCS3, and NEDD4. Cell. Microbiol. 2020, 22, e13150. [Google Scholar] [CrossRef] [PubMed]
- Zheng, T.; Zhou, Y.; Xu, X.; Qi, X.; Liu, J.; Pu, Y.; Zhang, S.; Gao, X.; Luo, X.; Li, M.; et al. MiR-30c-5p loss-induced PELI1 accumulation regulates cell proliferation and migration via activating PI3K/AKT pathway in papillary thyroid carcinoma. J. Transl. Med. 2022, 20, 20. [Google Scholar] [CrossRef] [PubMed]
- Tingö, L.; Ahlberg, E.; Johansson, L.; Pedersen, S.A.; Chawla, K.; Sætrom, P.; Cione, E.; Simpson, M.R. Non-Coding RNAs in Human Breast Milk: A Systematic Review. Front. Immunol. 2021, 12, 725323. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Zhu, Y.; Sun, B.; Shao, Y.; Jing, A.; Wang, J.; Xiao, Z. Assessing the survival of exogenous plant microRNA in mice. Food Sci. Nutr. 2014, 2, 380–388. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Smith, C.; Cokcetin, N.; Truong, T.; Harry, E.; Hutvagner, G.; Bajan, S. Cataloguing the small RNA content of honey using next generation sequencing. Food Chem. Mol. Sci. 2021, 2, 100014. [Google Scholar] [CrossRef]
- Ceolotto, G.; Giannella, A.; Albiero, M.; Kuppusamy, M.; Radu, C.; Simioni, P.; Garlaschelli, K.; Baragetti, A.; Catapano, A.L.; Iori, E.; et al. miR-30c-5p regulates macrophage-mediated inflammation and pro-atherosclerosis pathways. Cardiovasc. Res. 2017, 113, 1627–1638. [Google Scholar] [CrossRef] [PubMed]
- Hirschberger, S.; Hinske, L.C.; Kreth, S. MiRNAs: Dynamic regulators of immune cell functions in inflammation and cancer. Cancer Lett. 2018, 431, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Cavallini, A.; Minervini, F.; Garbetta, A.; Lippolis, C.; Scamarcio, G.; Di Franco, C.; D’Alessandro, R. High degradation and no bioavailability of artichoke miRNAs assessed using an in vitro digestion/Caco-2 cell model. Nutr. Res. 2018, 60, 68–76. [Google Scholar] [CrossRef] [PubMed]
- Pieri, M.; Theori, E.; Dweep, H.; Flourentzou, M.; Kalampalika, F.; Maniori, M.-A.; Papagregoriou, G.; Papaneophytou, C.; Felekkis, K. A bovine miRNA, bta-miR-154c, withstands in vitro human digestion but does not affect cell viability of colorectal human cell lines after transfection. FEBS Open Bio 2022, 12, 925–936. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Ren, X.; Ning, L.; Wang, P.; Xu, K. Stability and absorption mechanism of typical plant miRNAs in an in vitro gastrointestinal environment: Basis for their cross-kingdom nutritional effects. J. Nutr. Biochem. 2020, 81, 108376. [Google Scholar] [CrossRef] [PubMed]
- Minekus, M.; Alminger, M.; Alvito, P.; Ballance, S.; Bohn, T.; Bourlieu, C.; Carrière, F.; Boutrou, R.; Corredig, M.; Dupont, D.; et al. A standardised static in vitro digestion method suitable for food-an international consensus. Food Funct. 2014, 5, 1113–1124. [Google Scholar] [CrossRef] [PubMed]
- Brodkorb, A.; Egger, L.; Alminger, M.; Alvito, P.; Assunção, R.; Ballance, S.; Bohn, T.; Bourlieu-Lacanal, C.; Boutrou, R.; Carrière, F.; et al. INFOGEST static in vitro simulation of gastrointestinal food digestion. Nat. Protoc. 2019, 14, 991–1014. [Google Scholar] [CrossRef]
- Sousa, R.; Recio, I.; Heimo, D.; Dubois, S.; Moughan, P.J.; Hodgkinson, S.M.; Portmann, R.; Egger, L. In vitro digestibility of dietary proteins and in vitro DIAAS analytical workflow based on the INFOGEST static protocol and its validation with in vivo data. Food Chem. 2023, 404, 134720. [Google Scholar] [CrossRef] [PubMed]
- Grundy, M.M.L.; Abrahamse, E.; Almgren, A.; Alminger, M.; Andres, A.; Ariëns, R.M.C.; Bastiaan-Net, S.; Bourlieu-Lacanal, C.; Brodkorb, A.; Bronze, M.R.; et al. INFOGEST inter-laboratory recommendations for assaying gastric and pancreatic lipases activities prior to in vitro digestion studies. J. Funct. Foods 2021, 82, 104497. [Google Scholar] [CrossRef]
- Hiolle, M.; Lechevalier, V.; Floury, J.; Boulier-Monthéan, N.; Prioul, C.; Dupont, D.; Nau, F. In vitro digestion of complex foods: How microstructure influences food disintegration and micronutrient bioaccessibility. Food Res. Int. 2020, 128, 108817. [Google Scholar] [CrossRef]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; How Huang, K.; Jen Lee, M.; Galas, D.J.; Wang, K. The MicroRNA Spectrum in 12 Body Fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- FAO. Honey [Internet]. Available online: https://www.fao.org/3/ca4657en/CA4657EN.pdf (accessed on 17 January 2019).
- Bogdanov, S.; Jurendic, T.; Sieber, R.; Gallmann, P. Honey for Nutrition and Health: A Review. J. Am. Coll. Nutr. 2008, 27, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Doner, L.W. The sugars of honey—A review. J. Sci. Food Agric. 1977, 28, 443–456. [Google Scholar] [CrossRef] [PubMed]
- Salles, C.; Chagnon, M.-C.; Feron, G.; Guichard, E.; Laboure, H.; Morzel, M.; Semon, E.; Tarrega, A.; Yven, C. In-Mouth Mechanisms Leading to Flavor Release and Perception. Crit. Rev. Food Sci. Nutr. 2010, 51, 67–90. [Google Scholar] [CrossRef] [PubMed]
- Braun, R.J.; Parrott, E.L. Influence of Viscosity and Solubilization on Dissolution Rate. J. Pharm. Sci. 1972, 61, 175–178. [Google Scholar] [CrossRef] [PubMed]
- Gittings, S.; Turnbull, N.; Henry, B.; Roberts, C.J.; Gershkovich, P. Characterisation of human saliva as a platform for oral dissolution medium development. Eur. J. Pharm. Biopharm. 2015, 91, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Wijenayake, S.; Eisha, S.; Tawhidi, Z.; Pitino, M.A.; Steele, M.A.; Fleming, A.S.; McGowan, P.O. Comparison of methods for pre-processing, exosome isolation, and RNA extraction in unpasteurized bovine and human milk. PLoS ONE 2021, 16, e0257633. [Google Scholar] [CrossRef] [PubMed]
- Schuh, C.M.A.P.; Aguayo, S.; Zavala, G.; Khoury, M. Exosome-like vesicles in Apis mellifera bee pollen, honey and royal jelly contribute to their antibacterial and pro-regenerative activity. J. Exp. Biol. 2019, 222, jeb208702. [Google Scholar] [CrossRef]
- Chen, X.; Liu, B.; Li, X.; An, T.T.; Zhou, Y.; Li, G.; Wu-Smart, J.; Alvarez, S.; Naldrett, M.J.; Eudy, J.; et al. Identification of anti-inflammatory vesicle-like nanoparticles in honey. J. Extracell. Vesicles 2021, 10, e12069. [Google Scholar] [CrossRef]
- Ståhl, A.; Johansson, K.; Mossberg, M.; Kahn, R.; Karpman, D. Exosomes and microvesicles in normal physiology, pathophysiology, and renal diseases. Pediatr. Nephrol. 2019, 34, 11–30. [Google Scholar] [CrossRef]
- Xu, R.; Greening, D.W.; Zhu, H.-J.; Takahashi, N.; Simpson, R.J. Extracellular vesicle isolation and characterization: Toward clinical application. J. Clin. Investig. 2016, 126, 1152–1162. [Google Scholar] [CrossRef] [PubMed]
- Wade, B.; Cummins, M.; Keyburn, A.; Crowley, T.M. Isolation and detection of microRNA from the egg of chickens. BMC Res. Notes 2016, 9, 283. [Google Scholar] [CrossRef] [PubMed]
- Andreassen, R.; Worren, M.M.; Høyheim, B. Discovery and characterization of miRNA genes in atlantic salmon (Salmo salar) by use of a deep sequencing approach. BMC Genom. 2013, 14, 482. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, Q.; Hou, J.; Huang, G.; Zhao, S.; Zheng, N.; Wang, J. Loss of bioactive microRNAs in cow’s milk by ultra-high-temperature treatment but not by pasteurization treatment. J. Sci. Food Agric. 2022, 102, 2676–2685. [Google Scholar] [CrossRef] [PubMed]
- Gareev, I.; Ahmad, A.; Wang, J.; Beilerli, A.; Ilyasova, T.; Sufianov, A.; Beylerli, O. Gastric juice non-coding RNAs as potential biomarkers for gastric cancer. Front. Physiol. 2023, 14, 1179582. [Google Scholar] [CrossRef]
- Mu, J.; Zhuang, X.; Wang, Q.; Jiang, H.; Deng, Z.-B.; Wang, B.; Zhang, L.; Kakar, S.; Jun, Y.; Miller, D.; et al. Interspecies communication between plant and mouse gut host cells through edible plant derived exosome-like nanoparticles. Mol. Nutr. Food Res. 2014, 58, 1561–1573. [Google Scholar] [CrossRef] [PubMed]
- Schulz, E.; Karagianni, A.; Koch, M.; Fuhrmann, G. Hot EVs—How temperature affects extracellular vesicles. Eur. J. Pharm. Biopharm. 2020, 146, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Zeng, Q.; Han, Q.; Xia, W. Effect of pH, temperature and freezing-thawing on quantity changes and cellular uptake of exosomes. Protein Cell 2019, 10, 295–299. [Google Scholar] [CrossRef]
- Sidhom, K.; Obi, P.O.; Saleem, A. A Review of Exosomal Isolation Methods: Is Size Exclusion Chromatography the Best Option? Int. J. Mol. Sci. 2020, 21, 6466. [Google Scholar] [CrossRef]
- Ahmed, F.E. miRNA as markers for the diagnostic screening of colon cancer. Expert Rev. Anticancer Ther. 2014, 14, 463–485. [Google Scholar] [CrossRef] [PubMed]
- Smyczynska, U.; Bartlomiejczyk, M.A.; Stanczak, M.M.; Sztromwasser, P.; Wesolowska, A.; Barbarska, O.; Pawlikowska, E.; Fendler, W. Impact of processing method on donated human breast milk microRNA content. PLoS ONE 2020, 15, e0236126. [Google Scholar] [CrossRef] [PubMed]


| Digestion Phase | Pasteurized | % Soluble Fraction | |
|---|---|---|---|
| With Human Saliva | With SSF | ||
| Oral | Yes | 96.32 | 92.05 |
| No | 96.57 | 94.29 | |
| Blank | 99.22 | 96.78 | |
| Gastric | Yes | 97.23 | 94.77 |
| No | 97.41 | 96.70 | |
| Blank | 97.91 | 97.21 | |
| Intestinal | Yes | 98.95 | 95.92 |
| No | 98.27 | 96.84 | |
| Blank | 97.41 | 96.94 | |
| Pasteurization 1 | µg/g Sample | A260/A280 | A260/A230 |
|---|---|---|---|
| Manual Extraction | |||
| Yes | 0.64 | 1.55 | 0.7 |
| No | 0.51 | 1.60 | 0.5 |
| Semi-Automatic Extraction 2 | |||
| Yes | 9.78 | 1.79 | 1.93 |
| No | 11.59 | 1.81 | 2.01 |
| State | Pasteurized | Phase | µg/g | A260/A280 | A260/A230 |
|---|---|---|---|---|---|
| Digested | Yes | Intestinal | 903 | 1.85 | 2.10 |
| No | Intestinal | 922 | 1.86 | 2.17 |
| miRNA | Pasteurized 1 | Manual 2 (n = 4) | Semi-Automated 3 (n = 4) | p-Value |
|---|---|---|---|---|
| miR-30c-5p | Yes | 32.96 ± 0.71 | 30.74 ± 1.20 | 0.154 |
| No | 32.11 ± 0.13 | 30.81 ± 0.57 | 0.067 | |
| miR-92a-3p | Yes | 33.60 ± 0.03 | 30.76 ± 0.06 | <0.001 |
| No | 31.94 ± 1.35 | 29.78 ± 0.39 | 0.162 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abrego-Guandique, D.M.; Ilori, O.A.; Caroleo, M.C.; Cannataro, R.; Cione, E.; Tucci, P. Differential Digestive Stability of Food-Derived microRNAs: The Case of miR-30c-5p and miR-92a-3p in Polyfloral Honey. Curr. Issues Mol. Biol. 2024, 46, 7473-7485. https://doi.org/10.3390/cimb46070443
Abrego-Guandique DM, Ilori OA, Caroleo MC, Cannataro R, Cione E, Tucci P. Differential Digestive Stability of Food-Derived microRNAs: The Case of miR-30c-5p and miR-92a-3p in Polyfloral Honey. Current Issues in Molecular Biology. 2024; 46(7):7473-7485. https://doi.org/10.3390/cimb46070443
Chicago/Turabian StyleAbrego-Guandique, Diana Marisol, Olubukunmi Amos Ilori, Maria Cristina Caroleo, Roberto Cannataro, Erika Cione, and Paola Tucci. 2024. "Differential Digestive Stability of Food-Derived microRNAs: The Case of miR-30c-5p and miR-92a-3p in Polyfloral Honey" Current Issues in Molecular Biology 46, no. 7: 7473-7485. https://doi.org/10.3390/cimb46070443










