Genome-Wide Identification and Characterization of Alternative Oxidase (AOX) Genes in Foxtail Millet (Setaria italica): Insights into Their Abiotic Stress Response
Abstract
1. Introduction
2. Results
2.1. Identification and Characterization of AOX Gene Family in Foxtail Millet
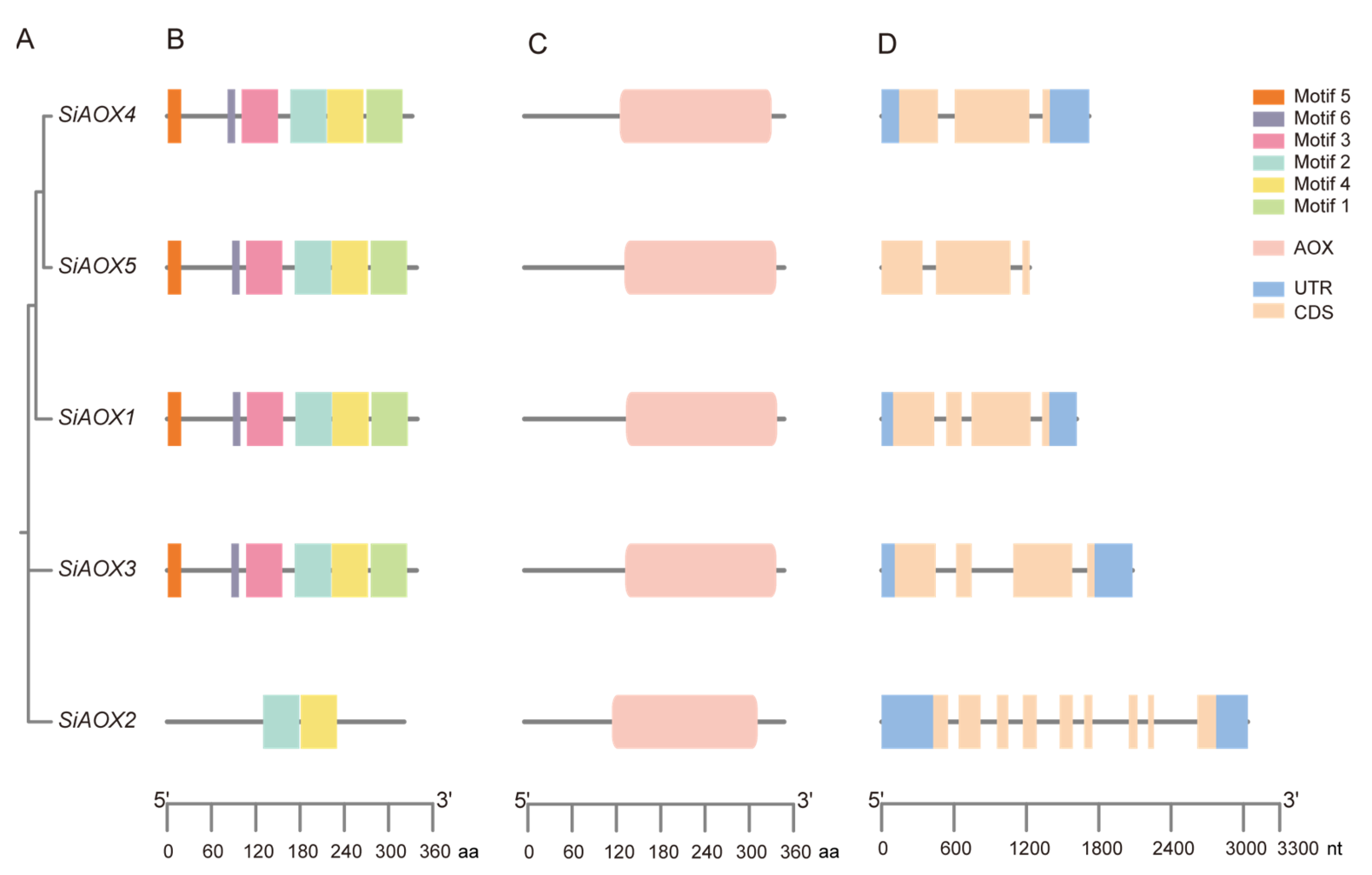
2.2. Structure Characterization and Three-Dimensional Modeling Analysis of AOX Genes in Foxtail Millet
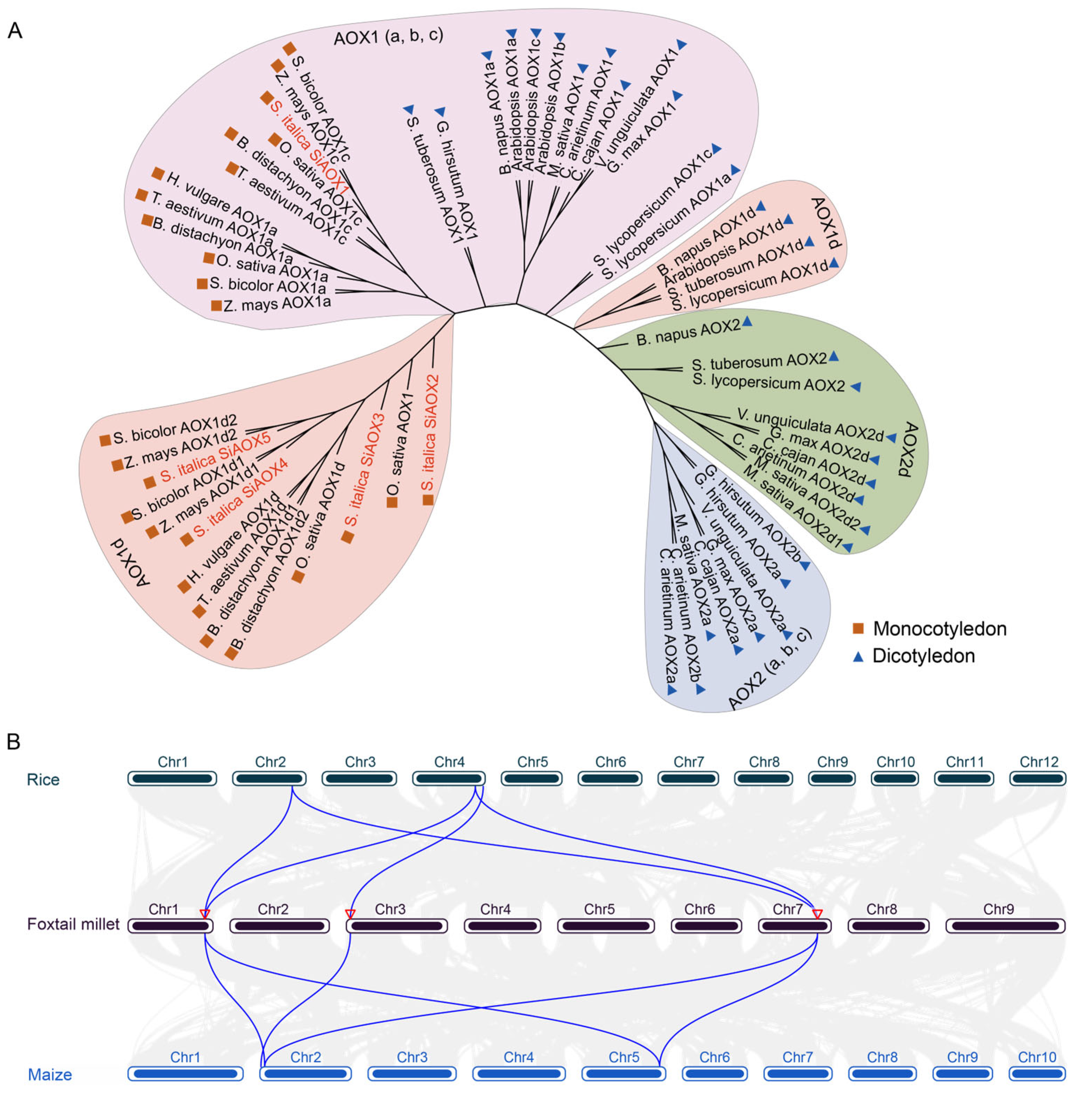
2.3. Phylogenetic Classification of AOX Proteins in Foxtail Millet
2.4. Promoter Analysis and Prediction of miRNA Target Sites of AOX Genes in Foxtail Millet
2.5. Colinearity and Selective Pressure Analysis of AOXs in Foxtail Millet
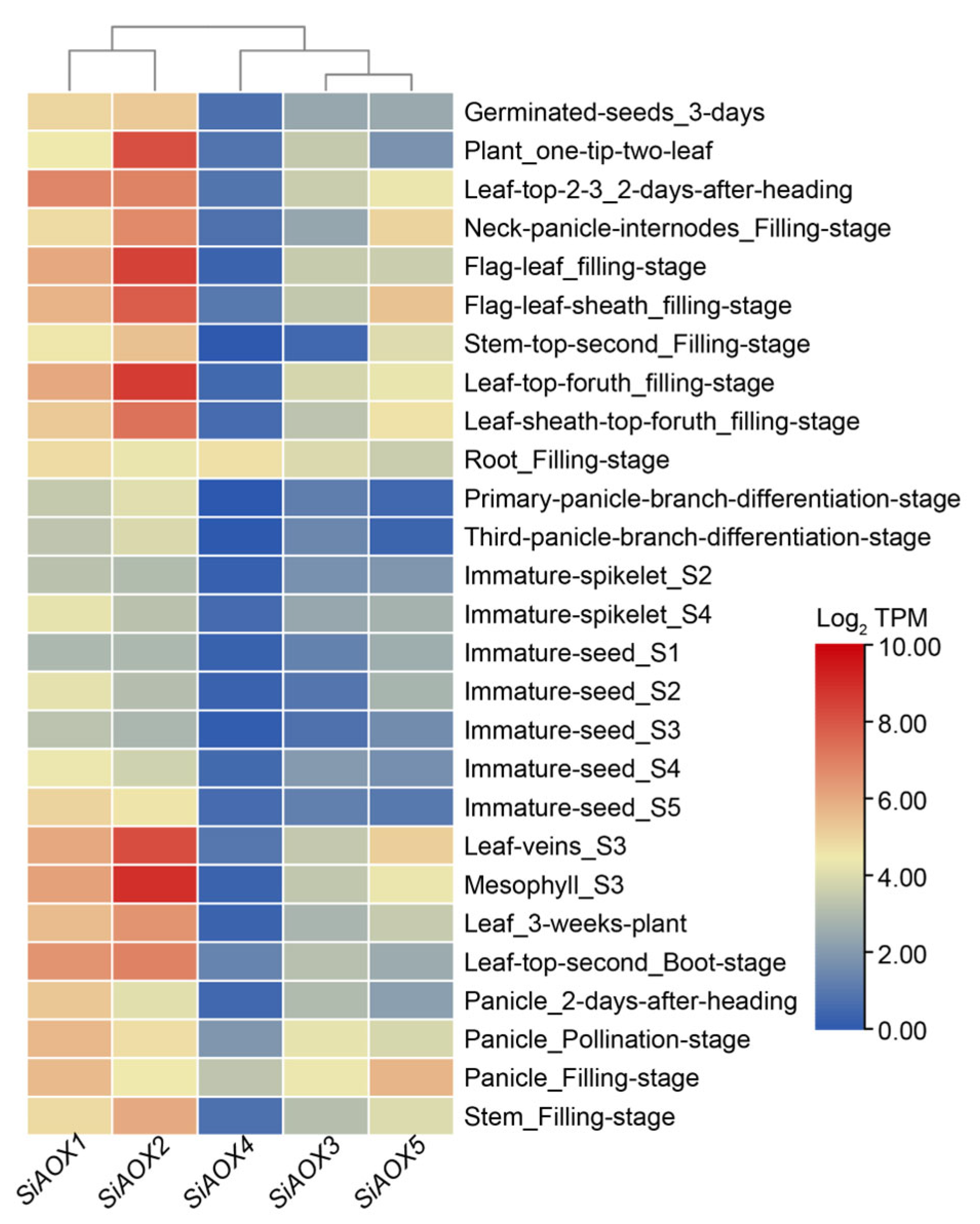
2.6. Expression Patterns of AOX Genes in Different Tissues in Foxtail Millet by RNA-seq
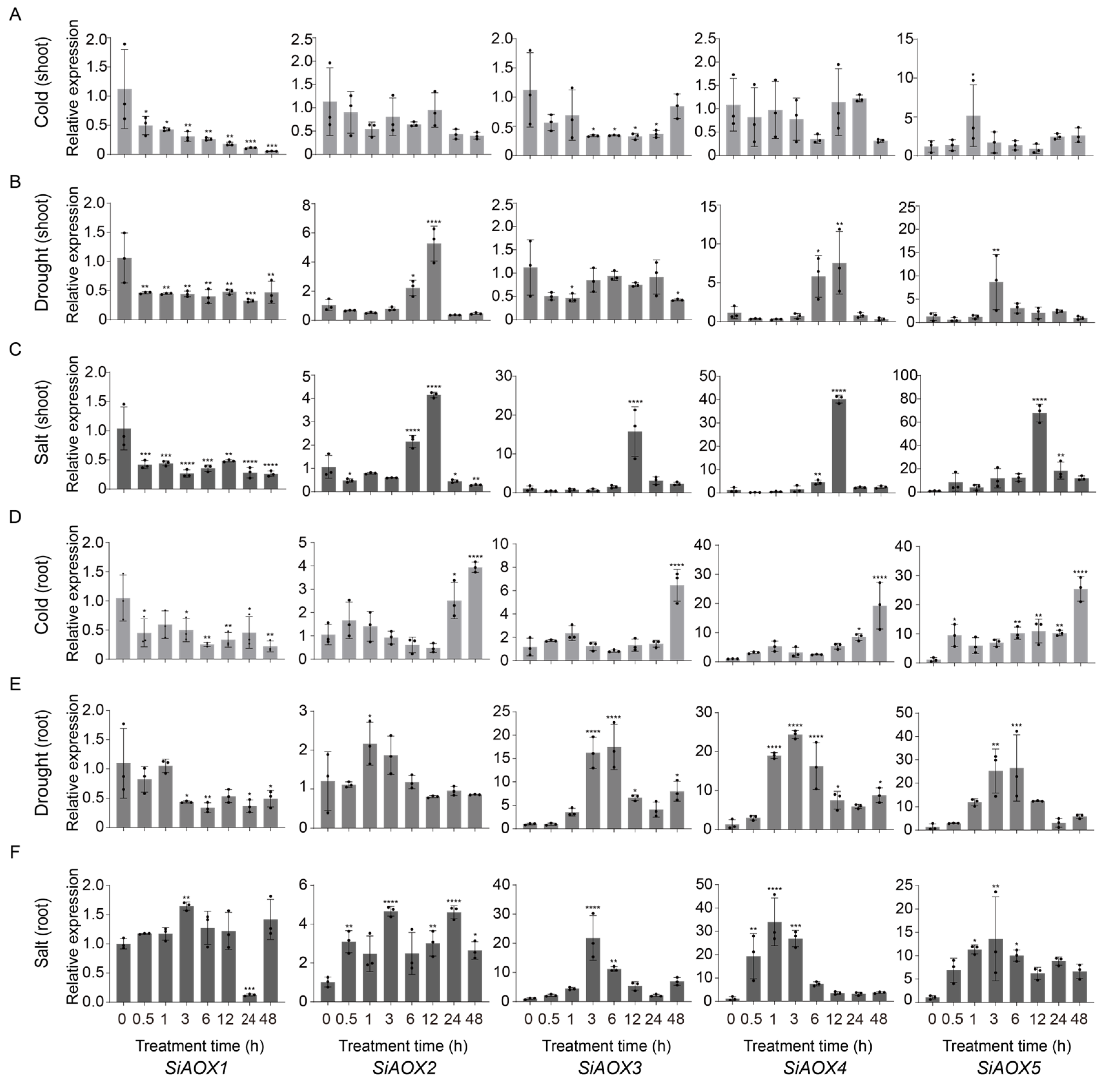
2.7. Response of SiAOX Genes under Various Abiotic Stress by Quantitative Real-Time PCR (RT-qPCR)
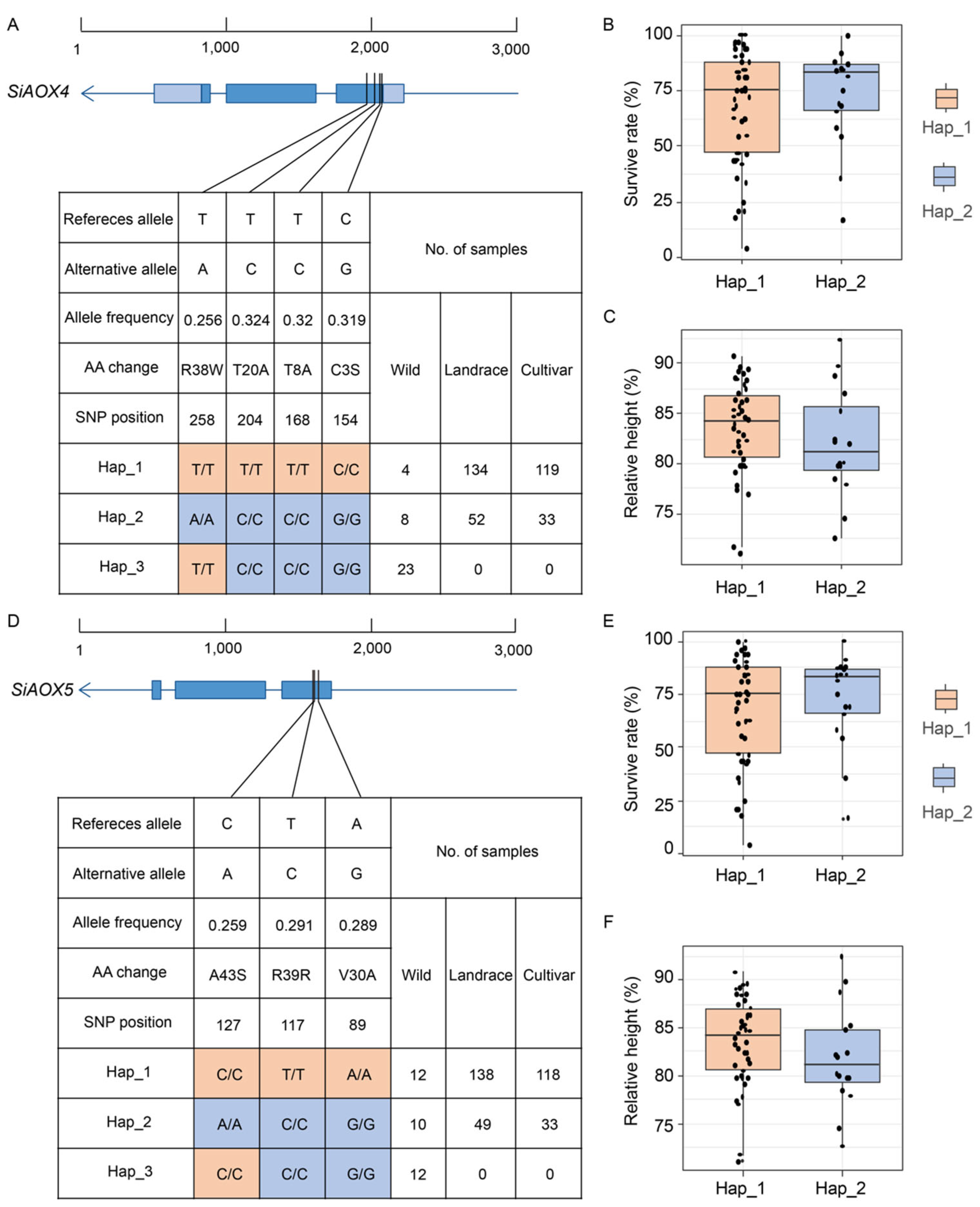
2.8. Co-Expression Network and Haplotype Variations Analysis of AOX Genes in Foxtail Millet
3. Discussion
3.1. SiAOX Genes and Their Classify in Foxtail Millet Genome
3.2. SiAOX Genes May Play a Crucial Role in the Response to Abiotic Stresses in Foxtail Millet
3.3. The Cluster Distribution of SiAOX4 and SiAOX5 May Enhance the Foxtail Millet’s Tolerance to Abiotic Stresses
4. Materials and Methods
4.1. Plant Growth Condition and Treatment
4.2. Genome-Wide Identification and Characterization of AOX Gene Family in Foxtail Millet
4.3. Chromosomal Localization, Gene Structure, Conserved Domain, Conserved Motif, and Three-Dimensional Modeling Analysis of AOX Genes in Foxtail Millet
4.4. Phylogenetic Tree, Selection Pressures and Collinearity Analysis of AOX Proteins in Foxtail Millet
4.5. cis-Regulatory Element Analysis, Prediction of miRNA Target Sites and Tissue Expression Analysis
4.6. RT-qPCR of SiAOX under Cold, Drought and Salt Stresses
4.7. Co-Expression Network Analysis and Haplotype Analysis of the SiAOX Genes and Association with Traits in Foxtail Millet
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Pardo, J.; VanBuren, R. Evolutionary innovations driving abiotic stress tolerance in C4 grasses and cereals. Plant Cell 2021, 33, 3391–3401. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Han, Y. Pan-genome brings opportunities to revitalize the ancient crop foxtail millet. Plant Commun. 2024, 5, 100735. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Zhang, H.; Li, X.; Shen, H.; Gao, J.; Hou, S.; Zhang, B.; Mayes, S.; Bennett, M.; Ma, J.; et al. A mini foxtail millet with an Arabidopsis-like life cycle as a C4 model system. Nat. Plants 2020, 6, 1167–1178. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Liang, H. Progress of the study on alternative oxidase. Chinese Bull. Bot. 1997, 14, 9–16. [Google Scholar]
- Florez-Sarasa, I.; Ribas-Carbo, M.; Del-Saz, N.F.; Schwahn, K.; Nikoloski, Z.; Fernie, A.R.; Flexas, J. Unravelling the in vivo regulation and metabolic role of the alternative oxidase pathway in C3 species under photoinhibitory conditions. New Phytol. 2016, 212, 66–79. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, J.; Li, R.; Ge, Y.; Li, Y.; Li, R. Plants’ response to abiotic stress: Mechanisms and strategies. Int. J. Mol. Sci. 2023, 24, 10915. [Google Scholar] [CrossRef]
- Kim, T.H.; Bohmer, M.; Hu, H.; Nishimura, N.; Schroeder, J.I. Guard cell signal transduction network: Advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu. Rev. Plant Biol. 2010, 61, 561–591. [Google Scholar] [CrossRef]
- Cutler, S.R.; Rodriguez, P.L.; Finkelstein, R.R.; Abrams, S.R. Abscisic acid: Emergence of a core signaling network. Annu. Rev. Plant Biol. 2010, 61, 651–679. [Google Scholar] [CrossRef]
- Manan, S.; Zhao, J. Role of Glycine max ABSCISIC ACID INSENSITIVE 3 (GmABI3) in lipid biosynthesis and stress tolerance in soybean. Funct. Plant Biol. 2021, 48, 171–179. [Google Scholar] [CrossRef]
- Manan, S.; Li, P.; Alfarraj, S.; Ansari, M.J.; Bilal, M.; Ullah, M.W.; Zhao, J. FUS3: Orchestrating soybean plant development and boosting stress tolerance through metabolic pathway regulation. Plant Physiol. Biochem. 2024, 213, 108803. [Google Scholar] [CrossRef]
- Selinski, J.; Scheibe, R.; Day, D.A.; Whelan, J. Alternative oxidase is positive for plant performance. Trends Plant Sci. 2018, 23, 588–597. [Google Scholar] [CrossRef] [PubMed]
- Giraud, E.; Van Aken, O.; Ho, L.H.; Whelan, J. The transcription factor ABI4 is a regulator of mitochondrial retrograde expression of ALTERNATIVE OXIDASE1a. Plant physiol. 2009, 150, 1286–1296. [Google Scholar] [CrossRef] [PubMed]
- Moore, A.L.; Siedow, J.N. The regulation and nature of the cyanide-resistant alternative oxidase of plant mitochondria. Biochim. Biophys. Acta BBA—Bioenerg. 1991, 1059, 121–140. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, D.M.; McIntosh, L. Isolation and characterization of a cDNA clone encoding an alternative oxidase protein of Sauromatum guttatum (Schott). Proc. Natl. Acad. Sci. USA 1991, 88, 2122–2126. [Google Scholar] [CrossRef]
- Martin, W.; Rotte, C.; Hoffmeister, M.; Theissen, U.; Gelius-Dietrich, G.; Ahr, S.; Henze, K. Early cell evolution, eukaryotes, anoxia, sulfide, oxygen, fungi first ?, and a tree of genomes revisited. IUBMB Life 2003, 55, 193–204. [Google Scholar] [CrossRef]
- Costa, J.H.; McDonald, A.E.; Arnholdt-Schmitt, B.; Fernandes de Melo, D. A classification scheme for alternative oxidases reveals the taxonomic distribution and evolutionary history of the enzyme in angiosperms. Mitochondrion 2014, 19, 172–183. [Google Scholar] [CrossRef] [PubMed]
- Vanlerberghe, G.C. Alternative oxidase: A mitochondrial respiratory pathway to maintain metabolic and signaling homeostasis during abiotic and biotic stress in plants. Int. J. Mol. Sci. 2013, 14, 6805–6847. [Google Scholar] [CrossRef]
- Zhang, D.W.; Yuan, S.; Xu, F.; Zhu, F.; Yuan, M.; Ye, H.X.; Guo, H.Q.; Lv, X.; Yin, Y.; Lin, H.H. Light intensity affects chlorophyll synthesis during greening process by metabolite signal from mitochondrial alternative oxidase in Arabidopsis. Plant Cell Environ. 2016, 39, 12–25. [Google Scholar] [CrossRef]
- Demircan, N.; Cucun, G.; Uzilday, B. Mitochondrial alternative oxidase (AOX1a) is required for the mitigation of arsenic-induced oxidative stress in Arabidopsis thaliana. Plant Biotechnol. Rep. 2020, 14, 235–245. [Google Scholar] [CrossRef]
- Song, C.; Zhao, Y.; Li, A.; Qi, S.; Lin, Q.; Duan, Y. Postharvest nitric oxide treatment induced the alternative oxidase pathway to enhance antioxidant capacity and chilling tolerance in peach fruit. Plant Physiol. Biochem. 2021, 167, 113–122. [Google Scholar] [CrossRef]
- Hou, L.; Zhao, M.; Huang, C.; He, Q.; Zhang, L.; Zhang, J. Alternative oxidase gene induced by nitric oxide is involved in the regulation of ROS and enhances the resistance of Pleurotus ostreatus to heat stress. Microb. Cell Fact. 2021, 20, 137. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Zou, L.; Li, Y.; Yao, X.; Xu, F.; Deng, X.; Zhang, D.; Lin, H. Mitochondrial alternative oxidase-dependent autophagy involved in ethylene-mediated drought tolerance in Solanum lycopersicum. Plant Biotechnol. J. 2018, 16, 2063–2076. [Google Scholar] [CrossRef] [PubMed]
- Challabathula, D.; Analin, B.; Mohanan, A.; Bakka, K. Differential modulation of photosynthesis, ROS and antioxidant enzyme activities in stress-sensitive and -tolerant rice cultivars during salinity and drought upon restriction of COX and AOX pathways of mitochondrial oxidative electron transport. J. Plant Physiol. 2022, 268, 153583. [Google Scholar] [CrossRef] [PubMed]
- Panda, S.K.; Sahoo, L.; Katsuhara, M.; Matsumoto, H. Overexpression of alternative oxidase gene confers aluminum tolerance by altering the respiratory capacity and the response to oxidative stress in tobacco cells. Mol. Biotechnol. 2013, 54, 551–563. [Google Scholar] [CrossRef]
- Garmash, E.V.; Velegzhaninov, I.O.; Ermolina, K.V.; Rybak, A.V.; Malyshev, R.V. Altered levels of AOX1a expression result in changes in metabolic pathways in Arabidopsis thaliana plants acclimated to low dose rates of ultraviolet B radiation. Plant Sci. 2020, 291, 110332. [Google Scholar] [CrossRef]
- Grabelnych, O.I.; Borovik, O.A.; Tauson, E.L.; Pobezhimova, T.P.; Katyshev, A.I.; Pavlovskaya, N.S.; Koroleva, N.A.; Lyubushkina, I.V.; Bashmakov, V.Y.; Popov, V.N.; et al. Mitochondrial energy-dissipating systems (alternative oxidase, uncoupling proteins, and external NADH dehydrogenase) are involved in development of frost-resistance of winter wheat seedlings. Biochemistry 2014, 79, 506–519. [Google Scholar] [CrossRef]
- Yang, H.; Deng, L.; Liu, H.; Fan, S.; Hua, W.; Liu, J. Overexpression of BnaAOX1b confers tolerance to osmotic and salt stress in rapeseed. G3 Genes Genom. Genet. 2019, 9, 3501–3511. [Google Scholar] [CrossRef]
- Oh, G.G.K.; O’Leary, B.M.; Signorelli, S.; Millar, A.H. Alternative oxidase (AOX) 1a and 1d limit proline-induced oxidative stress and aid salinity recovery in Arabidopsis. Plant Physiol. 2022, 188, 1521–1536. [Google Scholar] [CrossRef]
- Costa, J.H.; dos Santos, C.P.; e Lima, B.d.S.; Netto, A.N.M.; de Cruz Saraiva, K.D.; Arnholdt-Schmitt, B. In silico identification of alternative oxidase 2 (AOX2) in monocots: A new evolutionary scenario. J. Plant physiol. 2017, 210, 58–63. [Google Scholar] [CrossRef]
- Vanlerberghe, G.C.; Dahal, K.; Alber, N.A.; Chadee, A. Photosynthesis, respiration and growth: A carbon and energy balancing act for alternative oxidase. Mitochondrion 2020, 52, 197–211. [Google Scholar] [CrossRef]
- Borecký, J.; Nogueira, F.T.S.; de Oliveira, K.A.P.; Maia, I.G.; Vercesi, A.E.; Arruda, P. The plant energy-dissipating mitochondrial systems: Depicting the genomic structure and the expression profiles of the gene families of uncoupling protein and alternative oxidase in monocots and dicots. J. Exp. Bot. 2006, 57, 849–864. [Google Scholar] [CrossRef] [PubMed]
- Araújo Castro, J.; Gomes Ferreira, M.D.; Santana Silva, R.J.; Andrade, B.S.; Micheli, F. Alternative oxidase (AOX) constitutes a small family of proteins in Citrus clementina and Citrus sinensis L. Osb. PLoS ONE 2017, 12, e0176878. [Google Scholar] [CrossRef] [PubMed]
- Wanniarachchi, V.R.; Dametto, L.; Sweetman, C.; Shavrukov, Y.; Day, D.A.; Jenkins, C.L.D.; Soole, K.L. Alternative respiratory pathway component genes (AOX and ND) in rice and barley and their response to stress. Int. J. Mol. Sci. 2018, 19, 915. [Google Scholar] [CrossRef] [PubMed]
- Ding, C.; Chen, C.; Su, N.; Lyu, W.; Yang, J.; Hu, Z.; Zhang, M. Identification and characterization of a natural SNP variant in ALTERNATIVE OXIDASE gene associated with cold stress tolerance in watermelon. Plant Sci. 2021, 304, 110735. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Geng, X.; Bi, X.; Li, R.; Chen, Y.; Lu, C. Genome-wide identification of AOX family genes in Moso bamboo and functional analysis of PeAOX1b_2 in drought and salinity stress tolerance. Plant Cell Rep. 2022, 41, 2321–2339. [Google Scholar] [CrossRef]
- Zhang, S.; Yan, C.; Lu, T.; Fan, Y.; Ren, Y.; Zhao, J.; Shan, X.; Guan, Y.; Song, P.; Li, D.; et al. New insights into molecular features of the genome-wide AOX family and their responses to various stresses in common wheat (Triticum aestivum L.). Gene 2023, 888, 147756. [Google Scholar] [CrossRef]
- He, Q.; Wang, C.; He, Q.; Zhang, J.; Liang, H.; Lu, Z.; Xie, K.; Tang, S.; Zhou, Y.; Liu, B.; et al. A complete reference genome assembly for foxtail millet and Setaria-db, a comprehensive database for Setaria. Mol. Plant 2024, 17, 219–222. [Google Scholar] [CrossRef]
- Qiao, K.; Yao, X.; Zhou, Z.; Xiong, J.; Fang, K.; Lan, J.; Xu, F.; Deng, X.; Zhang, D.; Lin, H. Mitochondrial alternative oxidase enhanced ABA-mediated drought tolerance in Solanum lycopersicum. J. Plant Physiol. 2023, 280, 153892. [Google Scholar] [CrossRef]
- Zhang, H.; Wang, Y.; Zhao, B.; Zhang, L.; Qie, Q.; Han, Y.; Xukai, L. Identification of co-expression genes related to cold stress in foxtail millet by WGCNA. J. Agric. Sci. Technol. 2023, 25, 22–34. [Google Scholar]
- Yokoyama, R. AOX in action—Making plant mitochondrial metabolism special. Plant Physiol. 2023, 192, 2585–2587. [Google Scholar] [CrossRef]
- Zhou, S.; Chen, L.; Chen, H.; Li, Y.; Chen, G.; Lu, G.; Yang, H. Bioinformatics and expression analysis of alternative oxidase genes in sweetpotato. J. Nucl. Agric. Sci. 2022, 36, 270–281. [Google Scholar]
- Wang, H.; Ma, Y.; Qiao, Z.; Chang, Y.; Shu, K.; Ding, H.; Nie, Y.; Pan, G. Structural and functional characterization of AOX gene family. Biotechnol. Bull. 2022, 38, 160–170. [Google Scholar]
- Millar, A.H.; Whelan, J.; Soole, K.L.; Day, D.A. Organization and regulation of mitochondrial respiration in plants. Annu. Rev. Plant Biol. 2011, 62, 79–104. [Google Scholar] [CrossRef] [PubMed]
- Mróz, T.L.; Havey, M.J.; Bartoszewski, G. Cucumber possesses a single terminal alternative oxidase gene that is upregulated by cold stress and in the mosaic (MSC) mitochondrial mutants. Plant Mol. Biol. Rep. 2015, 33, 1893–1906. [Google Scholar] [CrossRef]
- Costa, J.H.; Jolivet, Y.; Hasenfratz-Sauder, M.P.; Orellano, E.G.; da Guia Silva Lima, M.; Dizengremel, P.; Fernandes de Melo, D. Alternative oxidase regulation in roots of Vigna unguiculata cultivars differing in drought/salt tolerance. J. Plant Physiol. 2007, 164, 718–727. [Google Scholar] [CrossRef]
- Clifton, R.; Millar, A.H.; Whelan, J. Alternative oxidases in Arabidopsis: A comparative analysis of differential expression in the gene family provides new insights into function of non-phosphorylating bypasses. Biochim. Biophys. Acta (BBA)—Bioenerg. 2006, 1757, 730–741. [Google Scholar]
- Clifton, R.; Lister, R.; Parker, K.L.; Sappl, P.G.; Elhafez, D.; Millar, A.H.; Day, D.A.; Whelan, J. Stress-induced co-expression of alternative respiratory chain components in Arabidopsis thaliana. Plant Mol. Biol. 2005, 58, 193–212. [Google Scholar] [CrossRef]
- Watling, J.R.; Robinson, S.A.; Seymour, R.S. Contribution of the alternative pathway to respiration during thermogenesis in flowers of the sacred lotus. Plant Physiol. 2006, 140, 1367–1373. [Google Scholar] [CrossRef] [PubMed]
- Florez-Sarasa, I.; Lambers, H.; Wang, X.; Finnegan, P.M.; Ribas-Carbo, M. The alternative respiratory pathway mediates carboxylate synthesis in white lupin cluster roots under phosphorus deprivation. Plant Cell Environ. 2014, 37, 922–928. [Google Scholar] [CrossRef]
- Song, X.; Li, Y.; Cao, X.; Qi, Y. MicroRNAs and their regulatory roles in plant-environment interactions. Annu. Rev. Plant Biol. 2019, 70, 489–525. [Google Scholar] [CrossRef]
- Zhang, L.; Song, J.; Lin, R.; Tang, M.; Shao, S.; Yu, J.; Zhou, Y. Tomato SlMYB15 transcription factor targeted by sly-miR156e-3p positively regulates ABA-mediated cold tolerance. J. Exp. Bot. 2022, 73, 7538–7551. [Google Scholar] [CrossRef]
- Khandal, H.; Singh, A.P.; Chattopadhyay, D. The MicroRNA397b-LACCASE2 module regulates root lignification under water and phosphate deficiency. Plant Physiol. 2020, 182, 1387–1403. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.; Udvardi, M. Global regulation of reactive oxygen species scavenging genes in alfalfa root and shoot under gradual drought stress and recovery. Plant Signal Behav. 2012, 7, 539–543. [Google Scholar] [CrossRef] [PubMed]
- Mehta, P.A.; Rebala, K.C.; Venkataraman, G.; Parida, A. A diurnally regulated dehydrin from Avicennia marina that shows nucleo-cytoplasmic localization and is phosphorylated by Casein kinase II in vitro. Plant Physiol. Biochem. 2009, 47, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Gao, J.; Song, J.; Guo, K.; Hou, S.; Wang, X.; He, Q.; Zhang, Y.; Zhang, Y.; Yang, Y.; et al. Multi-omics analyses of 398 foxtail millet accessions reveal genomic regions associated with domestication, metabolite traits, and anti-inflammatory effects. Mol. Plant 2022, 15, 1367–1383. [Google Scholar] [CrossRef]
- Li, X.; Shi, Z.; Gao, J.; Wang, X.; Guo, K. CandiHap: A haplotype analysis toolkit for natural variation study. Mol. Breed. 2023, 43, 21. [Google Scholar] [CrossRef]





| Gene Name | Gene Structure | Protein | Subcelluar Localization | ||||
|---|---|---|---|---|---|---|---|
| CDS Length (bp) | Intron | AA Length | MW (kDa) | pI | GRAVY | ||
| SiAOX1 | 1017 | 3 | 339 | 37.81 | 8.7 | −0.138 | Mitochondrial |
| SiAOX2 | 963 | 8 | 321 | 37.09 | 5.67 | −0.239 | Plasma Membrane |
| SiAOX3 | 1014 | 3 | 338 | 37.66 | 8.55 | −0.151 | Mitochondrial |
| SiAOX4 | 996 | 2 | 332 | 37.33 | 8.91 | −0.283 | Mitochondrial |
| SiAOX5 | 1014 | 2 | 338 | 37.21 | 7.31 | −0.205 | Mitochondrial |
| Gene Name | Gene ID | Ka | Ks | Ka/Ks |
|---|---|---|---|---|
| SiAOX1 | Z._mays_AOX1c | 0.0684 | 0.3500 | 0.1954 |
| SiAOX1 | S._bicolor_AOX1c | 0.0529 | 0.2425 | 0.2181 |
| SiAOX2 | O._sativa_AOX1 | 0.8616 | 1.8545 | 0.4646 |
| SiAOX3 | O._sativa_AOX1d | 0.1832 | 0.4413 | 0.4153 |
| SiAOX4 | Z._mays_AOX1d1 | 0.0404 | 0.1853 | 0.2180 |
| SiAOX4 | S._bicolor_AOX1d1 | 0.0450 | 0.1687 | 0.2665 |
| SiAOX5 | Z._mays_AOX1d2 | 0.0433 | 0.2028 | 0.2134 |
| SiAOX5 | S._bicolor_AOX1d2 | 0.0344 | 0.2358 | 0.1460 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, H.; Luo, Y.; Wang, Y.; Zhao, J.; Wang, Y.; Li, Y.; Pu, Y.; Wang, X.; Ren, X.; Zhao, B. Genome-Wide Identification and Characterization of Alternative Oxidase (AOX) Genes in Foxtail Millet (Setaria italica): Insights into Their Abiotic Stress Response. Plants 2024, 13, 2565. https://doi.org/10.3390/plants13182565
Zhang H, Luo Y, Wang Y, Zhao J, Wang Y, Li Y, Pu Y, Wang X, Ren X, Zhao B. Genome-Wide Identification and Characterization of Alternative Oxidase (AOX) Genes in Foxtail Millet (Setaria italica): Insights into Their Abiotic Stress Response. Plants. 2024; 13(18):2565. https://doi.org/10.3390/plants13182565
Chicago/Turabian StyleZhang, Hui, Yidan Luo, Yujing Wang, Juan Zhao, Yueyue Wang, Yajun Li, Yihao Pu, Xingchun Wang, Xuemei Ren, and Bo Zhao. 2024. "Genome-Wide Identification and Characterization of Alternative Oxidase (AOX) Genes in Foxtail Millet (Setaria italica): Insights into Their Abiotic Stress Response" Plants 13, no. 18: 2565. https://doi.org/10.3390/plants13182565
APA StyleZhang, H., Luo, Y., Wang, Y., Zhao, J., Wang, Y., Li, Y., Pu, Y., Wang, X., Ren, X., & Zhao, B. (2024). Genome-Wide Identification and Characterization of Alternative Oxidase (AOX) Genes in Foxtail Millet (Setaria italica): Insights into Their Abiotic Stress Response. Plants, 13(18), 2565. https://doi.org/10.3390/plants13182565







