Abstract
Persistent Vibrio-parahaemolyticus-associated vibriosis cases, attributed, in part, to the inefficient techniques for detecting viable-but-non-culturable (VBNC) Vibrio pathogens and the ingestion of undercooked seafood, is the leading cause of bacterial seafood-borne outbreaks, hospitalizations, and deaths in the United States. The effect of extreme heat processing on Vibrio biology and its potential food safety implication has been underexplored. In the present work, environmental samples from the wet market, lagoon, and estuarine environments were analyzed for V. parahaemolyticus recovery using a modified, temperature-dependent, two-step enrichment method followed by culture-based isolation, phenotype, and genotype characterizations. The work recovered novel strains (30% of 12 isolates) of V. parahaemolyticus from prolonged-heat-processing conditions (80 °C, 20 min), as confirmed by 16S rDNA bacterial identification. Select strains, VHT1 and VHT2, were determined to be hemolysis- and urease-positive pathogens. PCR analyses of chromosomal DNA implicated the tdh-independent, tlh-associated hemolysis in these strains. Both strains exhibited significant, diverse antibiotic profiles (p < 0.05). Turbidimetric and viable count assays revealed the pasteurization-resistant V. parahaemolyticus VHT1/VHT2 (62 °C, 8 h). These findings disclose the efficiency of Vibrio extremist recovery by the modified, two-step enrichment technique and improve knowledge of Vibrio biology essential to food safety reformation.
1. Introduction
Vibrio parahaemolyticus is a non-spore-forming [1], halophilic [2], and thermophilic [3] Gram-negative bacterium [4]. Attributed to this viable-but-nonculturable (VBNC) [5], infectious agent [3,4] and, hence, uneasy to detect, the number of V. parahaemolyticus-associated seafood-borne vibriosis [6,7] is increasing worldwide [3,8]. This gastroenteritis-causing pathogen [9] accounts for ca 45% [6,10] and 0.1% [11] of the total vibriosis and toll of foodborne illnesses (48 million) each year in the US, respectively, and is the leading cause of foodborne infections in China [12]. Infected patients, including all age groups, may carry various symptoms such as watery diarrhea, abdominal cramping, gastroenteritis, nausea, vomiting, fever, wound/soft tissue infections, bacteremia, low blood pressure, blistering skin lesions, and possibly death [6,13]. As such, the US Centers for Disease Control and Prevention (CDC) Cholera and Other Vibrio Illness Surveillance (COVIS) [10] and the Foodborne Disease Active Surveillance Network (FoodNet) [14] systems have been closely monitoring this pathogen since 1996. Additionally, the USDA Safe Minimum Internal Temperature chart [15] published cooking conditions for seafood, including shellfish, that requires a minimum internal cooking temperature of 63 °C to ensure safe consumption.
Persistent, increasing vibriosis incidents associated with V. parahaemolyticus are generally linked to the consumption of undercooked seafood contaminated with the pathogen [16,17] through their ecological niche [2] and improperly sanitized environment [18]. However, increasing V. parahaemolyticus prevalence has been detected in processed, ready-to-eat foods, suggesting that this pathogen can survive the harsh conditions of the food processes [9]. Hemolysin-related genes, such as thermostable-direct hemolysin (tdh), thermostable-direct-related hemolysin (trh), and thermolabile hemolysin (tlh) [19,20], are the primary virulence determinants of V. parahaemolyticus. In addition, most clinical strains possess the trh gene alongside the urease-producing gene (encoded by the uh gene) [21].
Unlike the thermal resuscitation technique, which uses extreme heat [22], VBNC V. parahaemolyticus resuscitation requires an enrichment-based cultivation technique, such as a one- or two-step enrichment technique, that involves alkaline peptone water (APW) selection alone or combined nutrient nourishment and APW (or drug) selection, respectively, followed by plating on selective agar containing thiosulfate-citrate-bile-salt-sucrose (TCBS) [23,24]. Of all the enrichment techniques documented, several groups noted that the heated saline enrichment (42 °C) technique was conducive to Vibrio growth [25,26,27,28,29,30,31]. However, both Delmore and Crisley [32] and Andrew et al. [33] noted that the heat process using temperatures between 49 and 55 °C was detrimental to this pathogen. Hence, these studies suggest that V. parahaemolyticus is a dormant-enabled, non-spore former, as spore former resuscitation requires a much higher temperature [22].
In light of the presence of VBNC Vibrio species, laboratories have employed antibody-labeled microscopy assays [13,34,35], gene-specific multiplex PCR [13,19,36], and one- and/or two-step enrichment-based cultivation techniques [37,38] for recovering this type of Vibrio. These techniques, however, compromise the feasibility of detecting all other 200 serogroups of cholera-causing Vibrio [39], as well as contribute to false-negative results as indigenous contaminants populate [40], which suggest that an efficient technique is needed for the accurate validation of food products for total pathogenic Vibrio contaminants, including the VBNC strains.
Collectively, these literature reports suggest the need of the present study to re-evaluate the conventional detection technique and cooking recommendation for V. parahaemolyticus containment in seafood products. To the best of our knowledge, the present work has not been noted in peer-reviewed literatures.
V. parahaemolycus-associated vibriosis that is deemed to be a persistent foodborne issue with the increasing number of cases worldwide [6,7] may be attributed, in part, to the presence of evasive V. parahaemolyticus from a conventional detection technique employed today, alterated cooking conditions, and lack of inspection benefits. The present study devised the conventional technique of APW enrichment [41], followed by heating selection [22], defined as a modified, two-step enrichment technique, to resuscitate/recover a group of heat-dependent, VBNC V. parahaemolyticus that could evade the conventional detection technique. The efficacy of the modified, two-step enrichment technique for detecting VBNC V. parahaemolyticus strains was simultaneously evaluated with the one-step enrichment technique (i.e., uses APW enrichment alone) [41]. The biology of this type of V. parahaemolyticus, which distinguishes it from the generally known, non-spore-like type of this pathogen, was characterized for virulence implications, which provides data for more effective validation for seafood safety. In this study, estuarine, lagoon, and wet-market seafoods were examined to isolate a group of VBNC V. parahaemolyticus requiring heat-dependent resuscitation and TCBS agar plating. Virulence determination tests in the literature, such as the Kanagawa phenomenon test, virulence genes PCR test, urease test, and antibiotic disk diffusion test, were subsequently applied to strains of V. parahaemolyticus and confirmed by 16S rDNA bacterial identification.
2. Materials and Methods
Sample collection. A total of eight shellfish samples (Table 1), including environmental samples (3, crab; 2, oyster) and wet market samples (3, shrimp), were collected from the Joe Patti seafood market (wet market, shrimp collection) around Santa Rosa Island (lagoon, oyster collection) and Gulf Island National beach (estuarine environment, crab collection) of Pensacola, Florida in early Fall (19 September 2017). All packaged samples were kept in an icebox (~1 °C) during transportation back to the laboratory, stored at refrigeration temperature (0–2 °C), and processed within a week. Environmental conditions at sample collection sites were determined to be pH 8.5 and temperature 30.3 °C.

Table 1.
List of environmental seafood samples used in this study.
Isolation, culture, and storage conditions. Strains of V. parahaemolyticus used in this study were isolated in the lab using a two-step enrichment technique described previously [37] with minor modifications (Figure 1). Briefly, microorganisms in a 25 g sample were homogenized in 225 g of enrichment broth containing APW (pH 8.6, 5% wt/v) using a Stomacher® 400 Circulator lab blender (Seward, Weber Scientific, Hamilton, NJ, USA), incubated at specific times, and bacterial dilutions were inoculated on thiosulfate-citrate-bile-salt-sucrose (TCBS, Difco, Detroit, MI, USA) agar plates or heat treated (i.e., 80 °C, 20 min followed by 4 °C, overnight) prior to the plating, followed by an incubation of 72 h at 35 °C before visible colonies were streaked on Brain Heart Infusion agar (BHI, Difco, Detroit, MI, USA) containing 3% NaCl for purification, with pure cultures being stored in a sterile solution containing BHI and glycerol (10% v/v) at −70 °C. Presumptive V. parahaemolyticus isolates were confirmed using 16S rDNA bacterial identification and species-gene-specific PCR (tlh). The bacterial cultures were prepared by inoculating thawed cultures (1/100) in sterile fresh culture solution containing BHI (pH 7.4) supplemented with 3% NaCl, followed by overnight incubation (~20 h) at 35 °C and a repeated subculture before experimental analyses, as described previously [18].
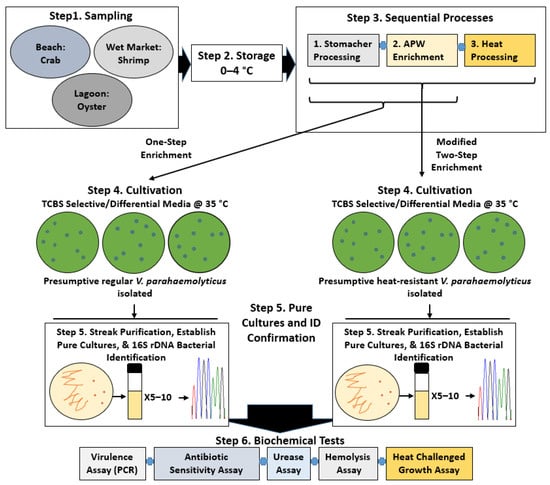
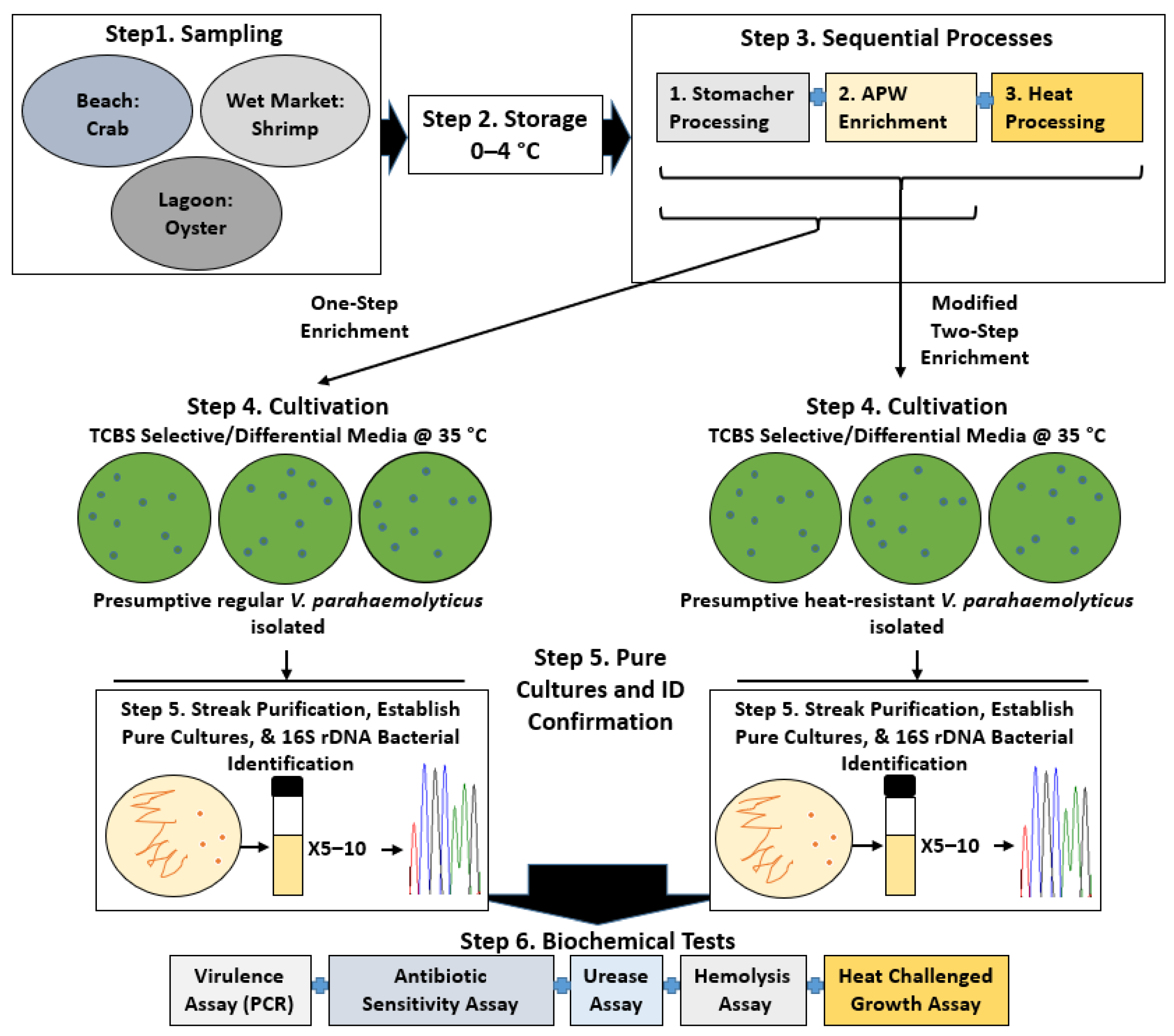
Figure 1.
Overview of the V. parahaemolyticus study. Step 1, sampling to collect environmental shellfish. Step 2, storage of samples before experimental analyses. Step 3, processes for the recovery of regular and heat-resistant V. parahaemolyticus. Step 4, cultivation of presumptive V. parahaemolyticus (from processed samples) on selective agar plates containing TCBS. Step 5, streak isolation, culture establishment, and bacterial identification of two forms of V. parahaemolyticus. Step 6, biochemical tests to characterize V. parahaemolyticus isolates.
Heat challenged viability assay. Sub-cultured, fresh, select strains of V. parahaemolyticus were serially diluted to a final cell concentration of 102–103 cfu/mL in LB broth. Dilution aliquots were incubated at 62 °C for 8 h in a water bath, followed by overnight incubation at 35 °C, and were then examined for growth by visible broth turbidity and viable plate count on agar containing BHI and 3% NaCl.
Hemolysis test. Hemolysis activity was examined using sheep erythrocytes [42] or human erythrocytes [16] as described previously with minor modifications. Select strains of V. parahaemolyticus sub-cultured twice in sterile BHI broth were spotted (5 µL) onto agar plates (pH 7.4) containing BHI, 1.5% human erythrocytes, and 1% NaCl. Inoculated plates were incubated at 35 °C for 24 h prior to hemolysis validation.
DNA extraction, PCR, and agarose gel electrophoresis conditions. Genomic DNA was prepared with a previously documented bead extraction method [43]. Briefly, pelleted fresh V. parahaemolyticus cultures (16 h) were resuspended in sterile DI water and spun down twice before being subjecting to bead collision with sterile micro-size beads (5 µm) in 0.1 mL sterile Tris buffer (10 mM, pH 7.4) to release the cytoplasmic components using a pulsing vortex mixer (SPW Industrial, Laguna Hills, CA, USA). Chromosomal DNA was phased out from cell debris using a high-speed microcentrifuge (VWR, Suwanee, GA, USA) and stored at −20 °C. The quality of DNA was analyzed using a UV spectrophotometer (Thermo Scientific, South San Francisco, CA, USA) and PCR using gene-specific primers.
PCR mixtures for the amplification of genes were formulated according to the GoTaq Flexi DNA Polymerase’s instructions (Promega, Madison, WI, USA). Briefly, each reaction mixture contained 5× PCR buffer (Promega), 0.4 μM of gene-specific primers (Table 2) (IDT, Coralville, IA, USA), 1.5 mM MgCl2 (Promega), 0.2 mM deoxynucleoside triphosphate mix (Fisher Scientific, Fair Lawn, NJ, USA), and 1.25 U of GoTaq polymerase (Promega). The PCR conditions were PCR step 1: 1 cycle of 5 min DNA denaturation at 95 °C; step 2: 40 cycles of 1 min DNA denaturation at 95 °C, 40 s primer-dependent annealing, 72 °C of DNA extension (gene-dependent incubation time); and followed by PCR step 3: 1 cycle of extended DNA extension for 10 min at 70 °C before infinite holding at 4 °C in a GeneAmp PCR System 9700 Thermal Cycler (Applied Biosystems, Thermo Scientific).

Table 2.
Primers used in this study.
PCR products (5 µL) were examined in a 1.6% agarose gel (formed in 1× Tris-borate-EDTA buffer) pre-stained with a GelStarTM Nucleic Acid Gel Stain (ratio 5 µL stain: 50 mL gel solution, Lonza Walkersville Inc., Walkersville, MD, USA) using a UV transilluminator.
Urease assay. V. parahaemolyticus urease activity was evaluated according to the protocol of a commercial urease testing kit. Briefly, a test reaction containing 1 mL of fresh cell solution of V. parahaemolyticus isolates (<20 h) and one tablet of Urease Test Tablets Key Scientific (Hardy diagnostics, Santa Maria, CA, USA) was mixed with a vortex machine and incubated at 37 °C for a maximum of 24 h, or until a pink color indicating a positive test developed.
Antibiotic disc diffusion test. A comparative antibiotic assay for distinguishing strains of V. parahaemolyticus was performed using the disc diffusion test with minor modifications [46]. Briefly, BHI agar plates supplemented with 3% NaCl were inoculated for bacterial lawns with 108 cfu/mL fresh V. parahaemolyticus cultures (~20 h) by using sterile glass rods. The antibiotic discs (BD; Sparks, MD, USA) chloramphenicol (CHL, 30 µg/disc), ciprofloxacin (CIP, 5 µg/disc), erythromycin (ERY, 15 µg/disc), gentamycin (GEN, 10 µg/disc), nalidixic acid (NAL, 30 µg/disc), neomycin (NEO, 30 µg/disc), streptomycin (STR, 10 µg/disc), and tetracycline (TET, 30 µg/disc) were distantly placed on the lawns. The inoculated plates were incubated for 24 h before antibiotic susceptibility diameters (mm) were determined and then scored per the Clinical and Laboratory Standards Institute (CLSI) M45 [47] and M100-S21(M2) (i.e., found in the antibiotic product user manual) guidelines for Vibrio species and enterococci/Escherichia coli, respectively.
3. Results
3.1. Colony Phenotype/Prevalence of V. parahaemolyticus from Thermally Treated Samples
In the present study, VBNC strains of V. parahaemolyticus were generated from wet-market and environmental shellfish seafoods. Using a conventional, one-step enrichment (i.e., APW enrichment) and a modified, two-step enrichment technique (i.e., APW enrichment + heat treatment), 12 of 50 isolates were confirmed as two biologically distinct groups, herein named regular and heat-resistant V. parahaemolyticus, respectively, and exclusively recovered from oyster samples at all incubation times tested (for the one-step enrichment technique) (Table 3) and 8/48 incubation hours tested (for the two-step enrichment technique) (Table 4 and Table 5). Regular V. parahaemolyticus exhibited a 1.4-fold higher number of isolates than the heat-resistant V. parahaemolyticus.

Table 3.
Colony-forming profiles of presumptive V. parahaemolyticus on TCBS agar media following the one-step enrichment method. Samples were plated after specific incubation times on TCBS. Bacterial colonies were identified with 16S rDNA sequencing.

Table 4.
Colony-forming profiles of presumptive V. parahaemolyticus on TCBS agar media following the two-step enrichment method. Enriched samples (0–72 h) were heated for 20 min at 80 °C and plated on TCBS agar. Bacterial colonies were identified with 16S rDNA sequencing.

Table 5.
List of V. parahaemolyticus isolates identified 1 in this study.
3.2. Heat Resistance in Heat-Resistant V. parahaemolyticus Vegetative Cells
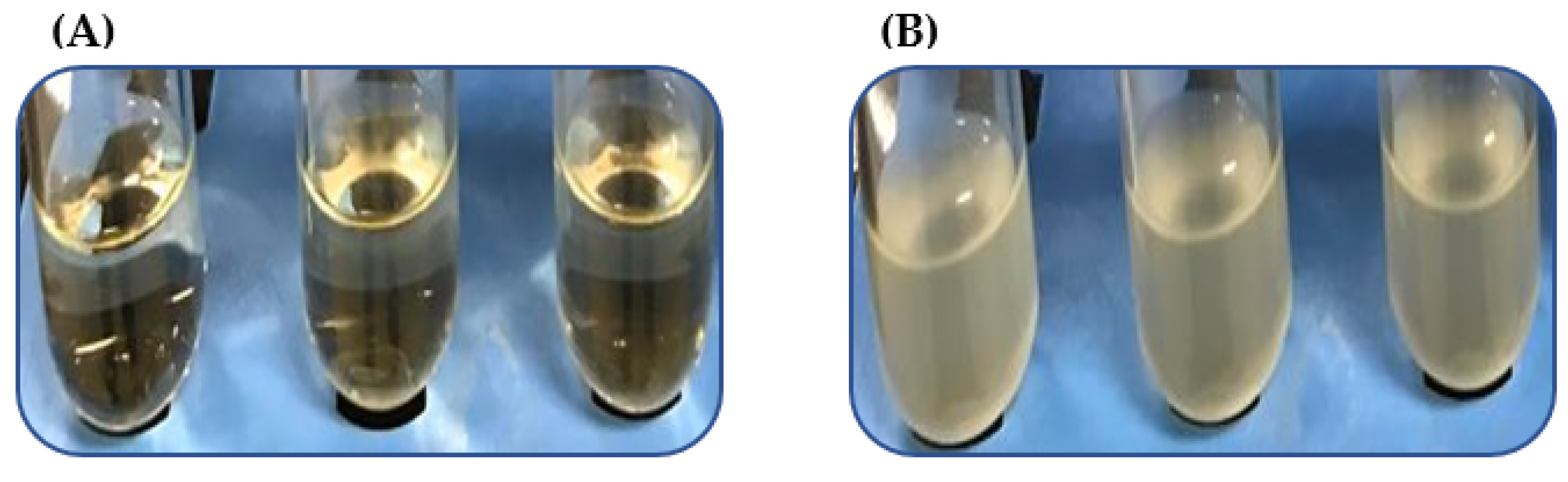
We analyzed a subset of heat-resistant strains, VHT1 and VHT2 (Table 5) of V. parahaemolyticus, for their susceptibility to the USDA-recommended pasteurization temperature of 62–65 °C for seafood [15]. VHT2 exhibited greater heat tolerance than VHT1, as demonstrated in growth turbidity replications (35 °C) (Figure 2 and Table 6) at post-heat treatment (8 h at 62 °C). It is worth noting that there was no noticeable growth turbidity for both VHT1 and VHT2 through the entire 8 h of incubation at 62 °C (data not shown). The viability of cells with turbidity was confirmed by plating (Figure 3).
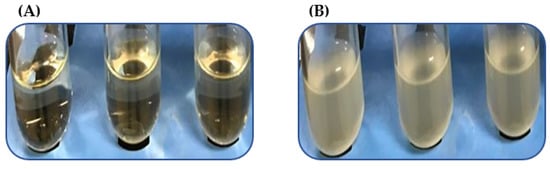

Figure 2.
Growth turbidity at post 35 °C incubation of V. parahaemolyticus pre-heated at 62 °C for 8 h. V. parahaemolyticus VHT1 (A) and VHT2 (B) were tested in triplicates.

Table 6.
Growth of V. parahaemolyticus strains VHT1 and VHT2 at post-heat treatment.
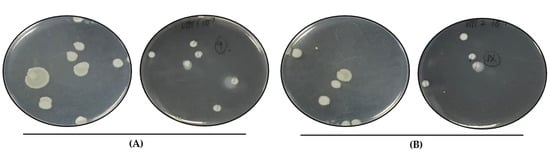
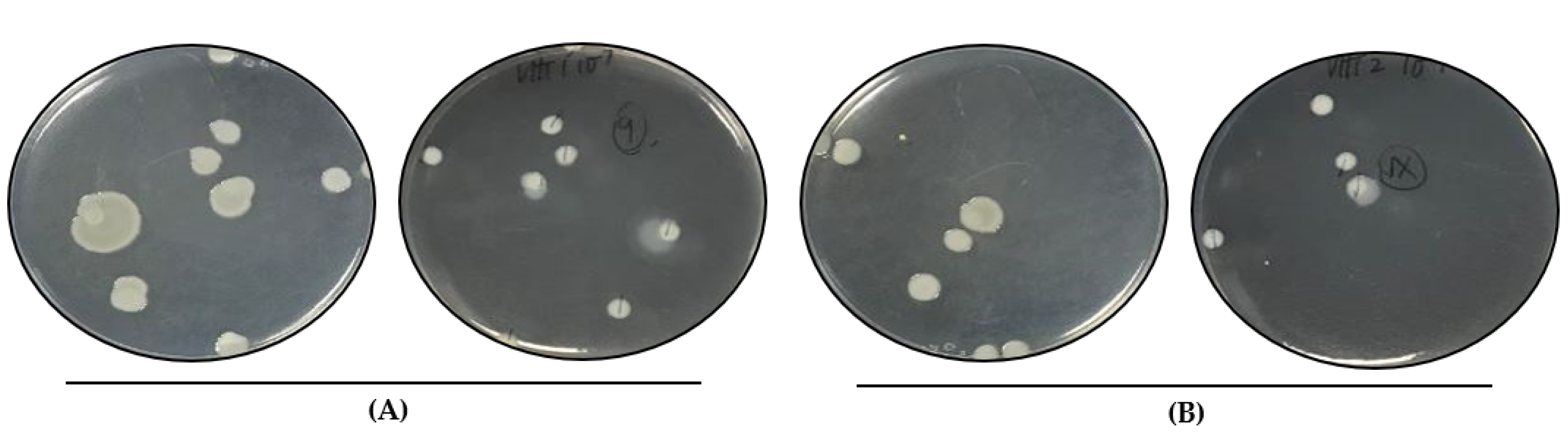
Figure 3.
Colony morphology of heated V. parahaemolyticus cells. Viable VHT1 (A) and VHT2 (B) cells heated at 62 °C for 8 h were recovered by plating and incubation at 35 °C. VHT79 (A), VHT80, and VHT81 (B) derived from the VHT1 and VHT2 colonies were established for identity confirmation analysis. (A) left, lid up; right, lid down. (B) left, lid up; right, lid down.
3.3. Virulence Determinants of the Heat-Resistant V. parahaemolyticus VHT1 and VHT2
- Kanagawa phenomenon
The heat resistant (VHT1, VHT2, VHT15, VHT16, VHT79, VHT80, or VHT81) and regular (VHT17 or VHT18) strains of V. parahaemolyticus (Figure 4) produced in this study were analyzed for virulence phenotype characterizations, such as the Kanagawa phenomenon (KP) and urease activity. Heat-resistant and regular strains (VHT1, VHT2, VHT15, VHT16, VHT17, and VHT18) examined with sheep erythrocytes did not exhibit positive KP activity as opposed to the use of human erythrocytes (tested in VHT1, VHT2, VHT79, VHT80, and VHT81) (Figure 4 and Table 7).
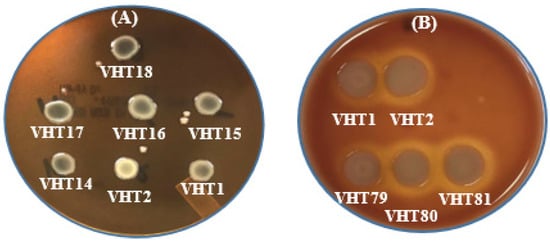
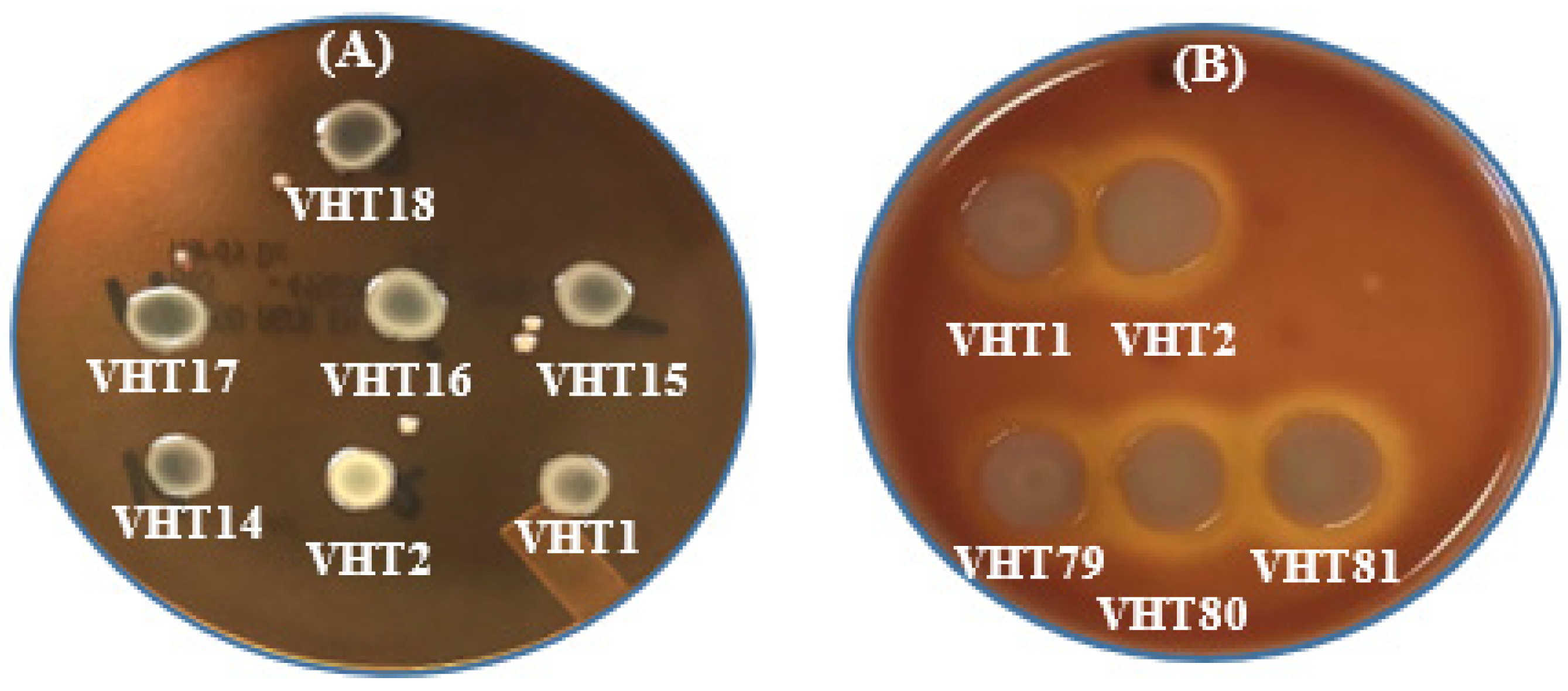
Figure 4.
In-vitro hemolysis assay for determining the KP of V. parahaemolyticus using sheep erythrocytes (A) or human erythrocytes (B). VHT1, VHT2, VHT14, VHT15, and VHT16 were heat-resistant strains of V. parahaemolyticus; VHT17 and VHT18 were regular strains of V. parahaemolyticus; VHT79, VHT80, and VHT81 were derived from VHT1/VHT2 at the post-pasteurization process. The same hemolytic results were observed in at least two separate replications.

Table 7.
Virulence phenotype (urease activity) and genotypes (tdh/tlh genes) of 7 strains of V. parahaemolyticus.
- PCR amplification of hemolysin genes
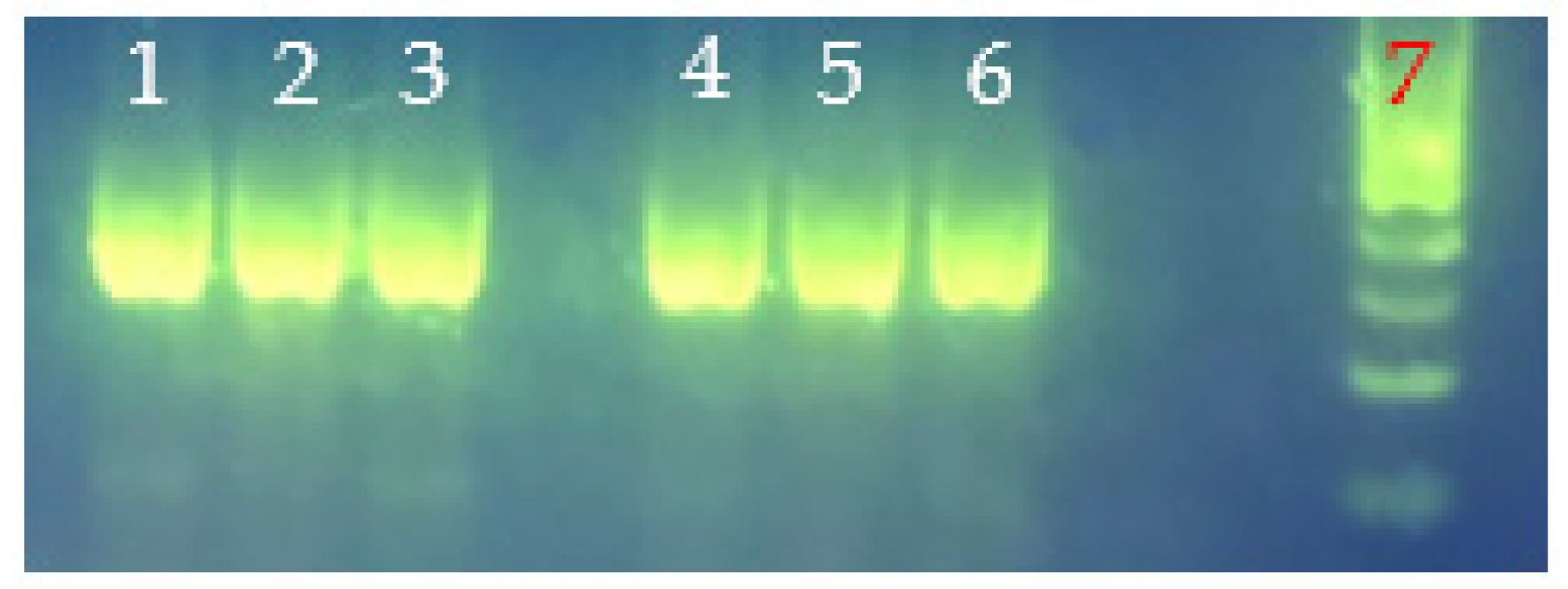
PCR analysis of hemolysin genes (i.e., tdh, tlh) in select heat-resistant and regular strains of V. parahaemolyticus with the gene-specific primers listed in Table 1 revealed V. parahaemolyticus with tdh−/tlh− (i.e., regular V. parahaemolyticus VHT17 and VHT18) or tdh−/tlh+ (i.e., heat-resistant V. parahaemolyticus VHT1, VHT2, VHT14, VHT15, and VHT16) genotypes (Figure 5 and Table 7) based on expected amplicon sizes (Table 1). Additionally, the derivatives of VHT1 (VHT79) and VHT2 (VHT80 and VHT81) retained the tlh gene of their parental strains as demonstrated in Figure 6.
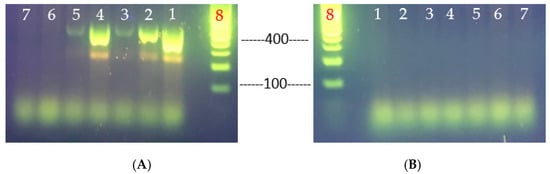
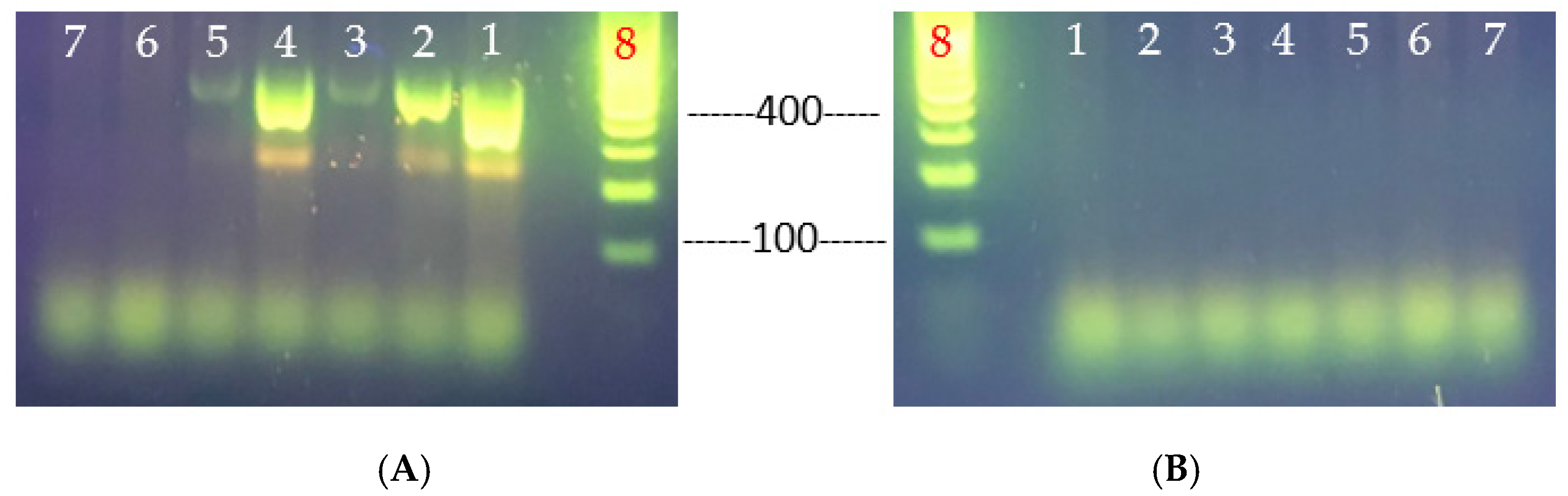
Figure 5.
Primer-specific PCR analyses for the determination of V. parahaemolyticus virulence genes, tlh (A) and tdh (B). V. parahaemolyticus strains tested: lane 1, VHT1; lane 2, VHT2; lane 3, VHT14; lane 4, VHT15; lane 5, VHT16; lane 6, VHT17; lane 7, VHT18; and lane 8, 100 bp DNA ladder.
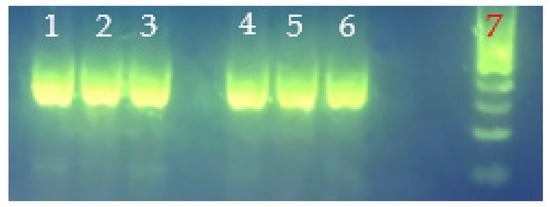
Figure 6.
tlh amplicons from genomic DNA of V. parahaemolyticus strains VHT79, VHT80, and VHT81 for strain purity validation. Lane 1, VHT79; lane 2, VHT80; lane 3, VHT81; lane 4, VHT79 (rep2); lane 5, VHT80 (rep2); lane 6 (rep2); VHT81; and lane 7, 100 bp DNA ladder.
- Urease activity
Urea hydrolysis conditions were adapted from the chemical supplier Hardy Diagnostics (product cat. # K650) since the urea tablets used for urease examination were less laborious to perform than a urea agar-based examination method. Urease activities were detected in all strains with no specific association between intense urease production and each of two forms of V. parahaemolyticus (Table 6) detected in this study.
3.4. Antibiotic Profile of the V. parahaemolyticus VHT1 and VHT2
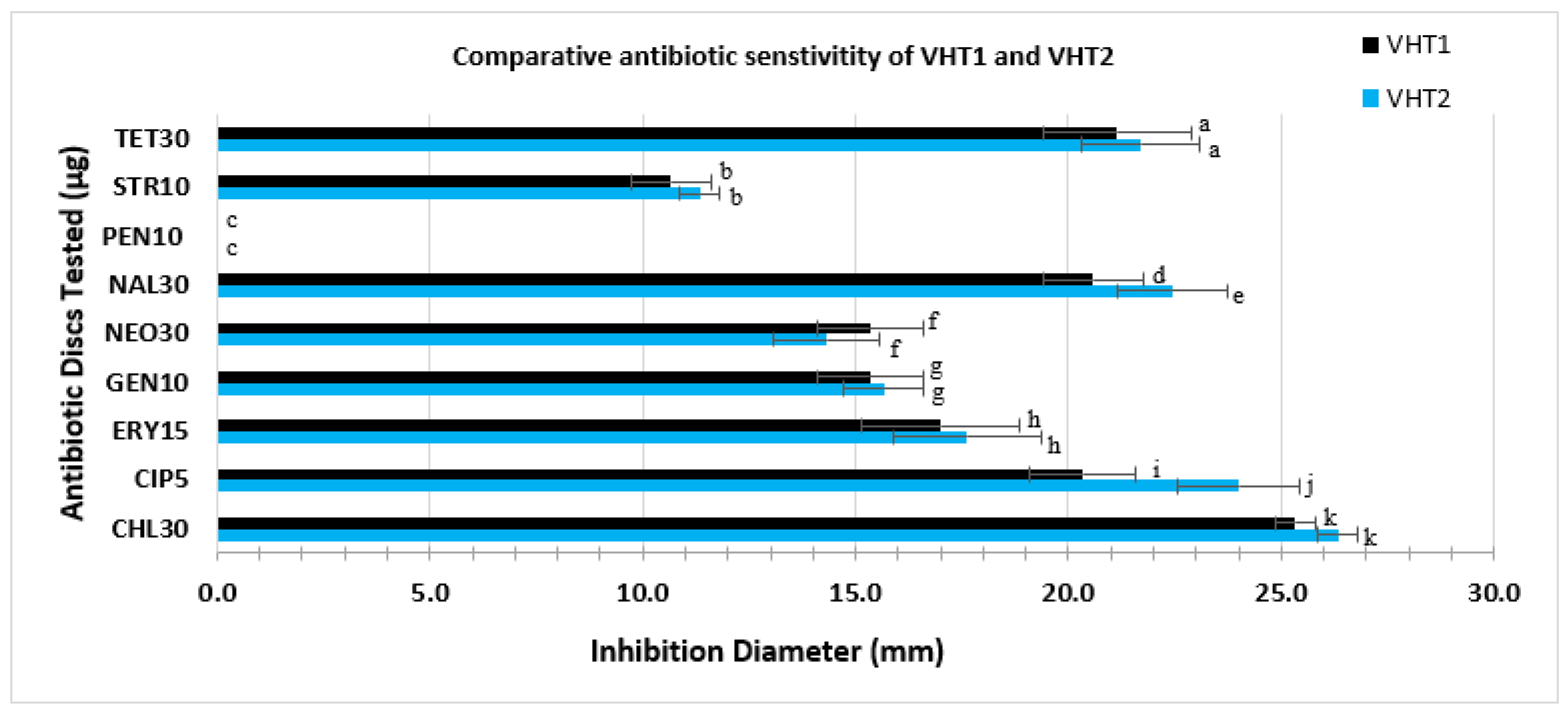
Antibiotic testing was carried out with a subset of two heat-resistant strains of V. parahaemolyticus (i.e., VHT1 and VHT2) obtained in this study. Different antibiotic susceptibility profiles were detected between VHT1 and VHT2. The latter strain was susceptible to more of the antibiotics tested (i.e., susceptible to chloramphenicol, ciprofloxacin, gentamycin, nalidixic acid, and tetracycline) as opposed to the VHT1 strain (i.e., susceptible to chloramphenicol, gentamycin, nalidixic acid, and tetracycline) (Table 8). A significantly (P < 0.05) high level of susceptibility was detected in VHT2 to ciprofloxacin and nalidixic acid as compared with the other strain tested (Figure 7). Both strains were highly resistant to penicillin (Figure 7) and scored an equal MAR index value of 0.22 (Table 8).

Table 8.
Antibiotic susceptibility interpretive data for the heat-resistant V. parahaemolyticus strains VHT1 and VHT2.
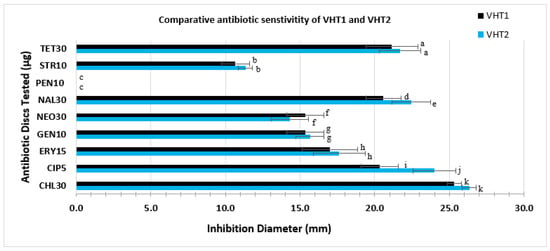
Figure 7.
Comparative antibiotic sensitivity profile of VHT1 and VHT2. Antibiotic discs with a specified concentration (µg/mL) used in this study include CHL30, CIP5, GEN10, NEO30, NAL30, PEN10, STR10, and TET30. Data and error bars are the means of at least triplicate replications and the standard deviations from the mean, respectively. Antibiotic activities with a significant difference, set at p < 0.05, are given as different letters, whereas antibiotic activities with the same letters are not significantly different.
4. Discussion
The ability of Vibrio species to form a viable-but-non-culturable (VBNC) state and become resistant to stresses, such as high salt content, pH, high temperatures [42,48], and antibiotics [49,50] has recently started to shed light on the persistent cases of vibriosis [35,51,52,53,54]. Previous studies separately reported by Mizunoe et al. [5], Okuda et al. [21], and Xu et al. [55] have suggested the presence of genotype and phenotype variabilities of V. parahaemolyticus, which could result in detection inconsistency/limit of one or many other detection techniques reported to date.
Conventionally, Vibrio spp. or spore-forming bacteria enrichment techniques use nutrient cultivation followed by selective enrichment [23,31,56] or heat treatment followed by cold incubation [22]. In this study, however, we combine SPW enrichment, heat-treatment, and cold incubation for resuscitating V. parahaemolyticus. This novel, combined enrichment and heat-selection technique (i.e., modified, two-step enrichment technique) resuscitated a group of environmental, heat-resistant V. parahaemolyticus strains (41.7%), in addition to regular V. parahaemolyticus strains (58.3%). This suggests the presence of a distinctive genotype and phenotype of V. parahaemolyticus that could remain unculturable by many conventional detection/cultivation techniques. Odeyemi noted that V. parahaemolyticus is more prevalent in oysters than other kinds of seafood investigated by various groups between 2003 and 2015, as demonstrated in a meta-analysis study [3]. This phenomenon was consistent with the prevalence findings (in this study), whereby V. parahaemolyticus could only be retrieved from oysters when analyzed along with other seafood samples. Hence, the observations suggest that oyster may be a V. parahaemolyticus’s preferred symbiont, including the heat-resistant strains reported for the first time in this study, compared with other kinds of shellfish seafood which are deemed frequent targets of contamination by V. parahaemolyticus. It is worth noting that this technique has not been documented in prior reports, and that the use of an extended SPW enrichment time (i.e., 72 h) in this study is novel (Table 3).
Delmore and Crisley [32] and Andrews et al. [33] noted that V. parahaemolyticus is susceptible to temperatures between 49 and 55 °C. Cebrián et al. [57] and Terano et al. [22] noted numerous non-Vibrio species of Gram-negative bacteria that are capable of withstanding pasteurization temperature (i.e., 65 °C). The heat-resistant strains of V. parahaemolyticus investigated in this study were able to survive extended pasteurization conditions (i.e., 62 °C, 8 h) as opposed to other vegetative cells of spore-forming bacteria [58] or non-spore forming bacteria [57]. This could suggest the differential expression of heat-shock proteins, such as DnaI [42], DnaK [48], GroEL, and GroeS [42,48] chaperones, whose role is implicated in bacterial maintenance of heat-deactivated proteins to revert their structure(s) and function(s) [59]. Both V. parahaemolyticus VHT1 and VHT2 strains exhibited different heat-tolerance phenotypes (VHT79, VHT80, VHT81) at post-pasteurization temperature (62 °C), as demonstrated by turbidimetric data (35 °C) (Figure 2) and followed by viable plate count (>8 log cfu/mL cells) data (Figure 3); nevertheless, these strains are susceptible to the resuscitation temperature (80 °C) used to recover them (Table 6). It is worth noting that the heat-resistant colonies of VHT80 and VHT81 derived from VHT2 (i.e., pasteurized VHT2) were different phenotypically and that our findings suggest, for the first time, that V. parahaemolyticus could tolerate pasteurization temperatures (62–65 °C). The latter strain (VHT81) appeared to have sticky colonies. This could be attributed to the heat-dependent alteration of Gram-negative bacteria’s outer membrane, which leads to non-lethal LPS damage [60] or homeoviscous adaptation [61]. In contrast to Delmore and Crisley [32], both strains survived a much higher incubation temperature than elevated temperatures between 49 and 55 °C.
KP was prepared using sheep erythrocytes as described by Ming et al. [62], as it is more economical than human erythrocytes. However, agar containing BHI was used for the reason that Wagatsuma agar [16] is costly, and that the V. parahaemolyticus strains did not grow well on agar plates containing Luria Bertani. In our study, human erythrocytes were able to demonstrate detectable hemolysis activity, disagreeing with Ming et al. [62]. The findings suggest that the V. parahaemolyticus heat-resistant strains VHT1 and VHT2 could possess one or more major hemolysin-producing gene(s) of the virulence strains, such as thermostable direct hemolysin (tdh) and/or tdh-related hemolysin (trh) [38], and that their derivative strains VHT79 (from VHT1), VHT80 (from VHT2), and VHT81 (VHT2) retained their hemolysis phenotypes post pasteurization (Figure 4 and Table 7). It is worth mentioning that all KP+ strains tested, including VHT1 and VHT2, did not exhibit hemolysis activity when cultured on agar media supplemented with sheep erythrocytes instead of human erythrocytes (Figure 4 and Table 7).
Wang et al. [63] noted that the Kanagawa hemolytic test could render false virulence results for V. parahaemolyticus and that tdh-specific PCR analysis could be used to confirm their virulence property. Letchumanan et al. [38] further noted that the non-virulence factor-associated hemolysin gene thermolabile hemolysin (tlh) could be employed for detecting total V. parahaemolyticus. Our PCR data revealed that regular V. parahaemolyticus strains and heat-resistant V. parahaemolyticus strains possess tdh−/tlh− and tdh−/tlh+ hemolysin genotypes, respectively (Table 7). It is worth noting that this is the first case of environmental V. parahaemolyticus with tlh−. Additionally, the data demonstrated that these KP+ V. parahaemolyticus strains (i.e., VHT79, VHT80, VHT81) possess tdh−/tlh+ virulence factors (Table 7) and that this hemolysin gene is phenotypically (Figure 4) and genotypically (Figure 6) stable to pasteurization conditions.
Xu et al. [55] and others [64,65,66,67,68] noted the presence of urease activity in V. parahaemolyticus and that it positively correlates/co-exists with the presence of the thermostable direct hemolysin-related hemolysin (trh) gene [21,64,65,68,69,70]. However, the other groups reported that urease activity is usually found in non-clinical strains of V. parahaemolyticus [65,66,67,68], which is in opposition to Xu et al. [55]. When urease activity was analyzed for regular (VHT17, 18) and heat-resistant (VHT1, 2, 14, 15, 16) strains of environmentally isolated V. parahaemolyticus in this study, both groups exhibited positive urease reaction (Table 7).
Park et al. [65] also noted that the urease and trh gene cluster does not positively correlate with the Kanagawa phenomenon (KP). However, the present study revealed that select strains, the heat-resistant, urease+ V. parahaemolyticus VHT1 and VHT2, were KP+ (Table 7). Letchumanan et al. [38] and Chung et al. [71] noted other V. parahaemolyticus hemolysins, such as the thermolabile hemolysin (tlh) and thermostable direct hemolysin (tdh), respectively, that could contribute to KP+. Hemolysin gene-specific PCR analyses revealed that the V. parahaemolyticus strains VHT1 and VHT2 were tdh− but tlh+ (Table 7). It is worth noting that this is the first report of the discovery of urease+, KP+ environmental V. parahaemolyticus [65].
The different antibiotics used in this study were adapted in part from previous documented work [18,46] and the availability of the antibiotics in the lab. Elexson et al. [72] and Letchumanan et al. [18] noted penicillin/ampicillin-resistant strains of V. parahaemolyticus from environmental and cultured seafood, respectively. When the antibiotic susceptibility test was compared between the heat-resistant V. parahaemolyticus VHT1 and VHT2, both demonstrated high levels of resistance to penicillin.
The availability of heat-resistant V. parahaemolyticus in seafoods suggest the potential cause of vibriosis food outbreak by a heat-resistant group of V. parahaemolyticus due to their ability to evade the conventional protocol of cooking and testing [15] and become a food safety threat and vibriosis cause. Our findings suggest, for the first time, that V. parahaemolyticus could survive food processing and cooking criteria [15], which could be an overlooked root cause of the persistent vibriosis in the US, and that they could be activated/recovered using the modified, two-step enrichment of combined nourishment (APW, 48 h), heating selection (80 °C, 20 min), and cooling steps. As previously noted, the prevalence of the V. parahaemolyticus variant [5,21,55], such as heat-resistant strains of environmental V. parahaemolyticus, could be attributed to specific gene transfers within the bacterial complex/mixture community, that could consist of Vibrio species and heat-resistant symbionts [73], and induced by global warming [74]. The results revealing positive KP activity and tlh amplicons suggest the presence of environmental V. parahaemolyticus pathogens [19,20], and that these strains (i.e., VHT1 and VHT2 derived strains, VHT79, VHT80, and VHT81) (Table 5 and Table 7) possess heat-stable hemolysin gene(s), such as tlh, and that it encodes a heat-stress protecting product, possibly suggesting a positive correlation between V. parahaemolyticus heat resistance (i.e., 62 °C, 8 h) and persistent vibriosis cases, as noted by Ueda et al. [75], whose work determined a positive correlation between thermolabile protein expression and thermotolerance in a member of Gram-negative bacteria. The PCR results confirm that this pathogen could use tdh-independent hemolysin(s), such as trh [38,71], tlh [2,38], VP3048 putative hemolysin [48], and/or others [71] for exhibiting KP activity (Table 7), and that urease-positive V. parahaemolyticus strains do not simultaneously possess a tdh-encoding gene, as noted by Wang et al. [63]. Additionally, the data suggest that tdh and tlh hemolysins are not essential in environmental V. parahaemolyticus metabolism, as these genes were absent in a subset (tlh gene) or all (tdh gene) of our culture isolates (Table 6). Further work (i.e., transcriptomic analyses) exploring the transcriptomic response of this group of V. parahaemolyticus strains to extreme temperature (i.e., 62 °C) is worthwhile for fostering a seafood processing/cooking and safety regimen. The results demonstrating positive urease activity suggest that both urease-positive heat-resistant and regular V. parahaemolyticus (Table 6) could possess a trh gene. Several groups reported that bacteria could produce urease when exposed to nitrogen limitation [76], acidic pH [77], or urea availability [78,79]. Berutti et al. [79] noted the increasing level of nitrogen content in environmental water since 2008, which suggests the adoption of urease in environmental V. parahaemolyticus. Thus, this indicates that environmental V. parahaemolyticus could readily produce urease in response to the urea availability supplemented in the urease assay. The results demonstrating high levels of resistance to penicillin could be attributed to the outer membrane of the Gram-negative bacteria, which interferes with antibiotic drugs’ entry and antimicrobial efficacy [46,80], as well as to the overuse of antibiotics in aquaculture that leads to bacterial resistance [81,82]. VHT2 was relatively more susceptible to ciprofloxacin and nalidixic acid (Figure 7) than VHT1, suggesting an antibiotic-resistance diversity of heat-resistant, environmental V. parahaemolyticus. Collectively, the results suggest that the V. parahaemolyticus strains of VHT1 and VHT2 (heat-resistant strains) possess tlh, trh (presumptive), urease, and KP activities.
5. Conclusions
A viable-but-nonculturable physiological state is a well-embraced contributing factor to bacterial resistance to, and evasive abilities from, hurdle technology that result in persistent foodborne bacterial infections, including V. parahaemolyticus-associated vibriosis. The present study reveals the first recovery of heat-resistant variants of environmental V. parahaemolyticus using the modified, two-step enrichment technique of SPW enrichment (step 1), heat enrichment, and refrigeration (step 2). These novel strains possessed tdh−, Trh+ (presumptive), urease+, Kanagawa phenomenon, and spore-like characteristics as opposed to the urease−/urease+, non-spore-like environmental strains determined in previous studies [32,33]. In addition, we disclose, for the first time, tlh− V. parahaemolyticus [38], urease+/KP+ environmental V. parahaemolyticus [65], and KP analysis inconsistency with sheep blood. Knowledge of different enrichment conditions that are capable of improving V. parahaemolyticus recovery, namely, the regular and heat-resistant forms, and understanding the biology of VBNC bacteria, may help detect total V. parahaemolyticus in seafoods and improve the accuracy of test results by manipulating the currently employed FDA-recommended conditions [24,30] for V. parahaemolyticus detection, both biologically and physically. These findings imply that the detection conditions that are recommended by the FDA may fail with the resuscitation of heat-dependent V. parahaemolyticus and that the modified, two-step enrichment technique that was examined should be adopted by Vibrio testing labs. In addition, this technique should be further investigated for the potential recovery of other species of VBNC Vibrio pathogens of public concern and additional samples from multiple locations are warranted due to these initial/novel results.
Author Contributions
The work has been contributed by the authors as follows: Conceptualization, H.K.T.; methodology, H.K.T.; validation, G.M. and H.M.; formal analysis, H.K.T., G.M. and H.M.; investigation, H.K.T., G.M. and H.M.; resources, H.K.T. and H.M.; writing—original draft preparation, H.M.; writing—review and editing, H.K.T. and G.M.; visualization, H.K.T. and G.M.; supervision, H.K.T.; project administration, H.K.T.; funding acquisition, H.K.T. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors would like to thank the dean of the University of West Alabama Biological and Environmental Sciences department, John McCall, for sample collection in Florida. Additionally, the authors would like to recognize students (Xiaoning Wei and Erica Johnson), faculty (Kevin Morse), and administrative groups (the University Office of Sponsored Programs and Research and Department of Biological and Environmental Sciences) for their bench work, reviewing, and financial contributions to this work, respectively.
Conflicts of Interest
The authors have no known conflict of interest to disclose.
References
- Rodrick, G.E. Indigenous pathogens: Vibrionaceae. In Microbiology of Marine Food Products, 1st ed.; Ward, D.R., Hackney, C., Eds.; Springer: Boston, MA, USA, 1991; pp. 285–300. [Google Scholar] [CrossRef]
- Broberg, C.A.; Calder, T.J.; Orth, K. Vibrio parahaemolyticus cell biology and pathogenicity determinants. Microbes Infect. 2011, 13, 992–1001. [Google Scholar] [CrossRef] [PubMed]
- Odeyemi, O.A. Incidence and prevalence of Vibrio parahaemolyticus in seafood: A systematic review and meta-analysis. SpringerPlus 2016, 5, 464–480. [Google Scholar] [CrossRef] [PubMed]
- Tarr, C.L.; Patel, J.S.; Puhr, N.D.; Sowers, E.G.; Bopp, C.A.; Strockbine, N.A. Identification of Vibrio Isolates by a multiplex PCR assay and rpoB sequence determination. J. Clin. Microbiol. 2007, 45, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Mizunoe, Y.; Wai, S.N.; Ishikawa, T.; Takade, A.; Yoshida, S. Resuscitation of viable but nonculturable cells of Vibrio parahaemolyticus induced at low temperature under starvation. FEMS Microbiol. Lett. 2000, 186, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Vibrio Species Causing Vibriosis. Available online: https://www.cdc.gov/vibrio/healthcare.html (accessed on 20 January 2022).
- Sims, J.N.; Isokpehi, R.D.; Cooper, G.A.; Bass, M.P.; Brown, S.D.; St John, A.L.; Gulig, P.A.; Cohly, H.H. Visual analytics of surveillance data on foodborne vibriosis, United States, 1973–2010. Environ. Health Insights 2011, 5, 71–85. [Google Scholar] [CrossRef]
- Martinez-Urtaza, J.; Bowers, J.C.; Trinanes, J.; DePaola, A. Climate anomalies and the increasing risk of Vibrio parahaemolyticus and Vibrio vulnificus illnesses. Food Res. Int. 2010, 43, 1780–1790. [Google Scholar] [CrossRef]
- Li, Y.; Xie, T.; Pang, R.; Wu, Q.; Zhang, J.; Lei, T.; Xue, L.; Wu, H.; Wang, J.; Ding, Y.; et al. Food-Borne Vibrio parahaemolyticus in China: Prevalence, antibiotic susceptibility, and genetic characterization. Front. Microbiol. 2020, 11, 1670. [Google Scholar] [CrossRef]
- Vibrio Species Causing Vibriosis. Available online: https://www.cdc.gov/vibrio/surveillance.html (accessed on 20 January 2022).
- Estimates of Foodborne Illness in the United States. Available online: https://www.cdc.gov/foodborneburden/index.html (accessed on 20 January 2022).
- Yang, Y.; Xie, J.F.; Li, H.; Tan, S.W.; Chen, Y.F.; Yu, H. Prevalence, antibiotic susceptibility and diversity of Vibrio parahaemolyticus isolates in seafood from south China. Front. Microbiol. 2017, 8, 2566. [Google Scholar] [CrossRef]
- McCarthy, S.A.; DePaola, A.; Cook, D.W.; Kaysner, C.A.; Hill, W.E. Evaluation of alkaline phosphatase- and digoxigenin-labelled probes for detection of the thermolabile hemolysin (tlh) gene of Vibrio parahaemolyticus. Lett. Appl. Microbiol. 1999, 28, 66–70. [Google Scholar] [CrossRef]
- Newton, A.; Kendall, M.; Vugia, D.J.; Henao, O.L.; Mahon, B.E. Increasing rates of vibriosis in the United States, 1996–2010: Review of surveillance data from 2 systems. Clin. Infect. Dis. 2012, 54, S391–S395. [Google Scholar] [CrossRef]
- Safe Minimum Internal Temperature Chart. Available online: https://www.fsis.usda.gov/food-safety/safe-food-handling-and-preparation/food-safety-basics/safe-temperature-chart (accessed on 20 January 2022).
- Gutierrez, C.K.; Klein, S.L.; Lovell, C.R. High frequency of virulence factor genes tdh, trh, and tlh in Vibrio parahaemolyticus strains isolated from a pristine estuary. Appl. Environ. Microbiol. 2013, 79, 2247–2252. [Google Scholar] [CrossRef] [PubMed]
- Trinanes, J.; Martinez-Urtaza, J. Future scenarios of risk of Vibrio infections in a warming planet: A global mapping study. Lancet Planet Health 2021, 5, e426–e435. [Google Scholar] [CrossRef]
- Letchumanan, V.; Yin, W.F.; Lee, L.H.; Chan, K.G. Prevalence and antimicrobial susceptibility of Vibrio parahaemolyticus isolated from retail shrimps in Malaysia. Front. Microbiol. 2015, 6, 33. [Google Scholar] [CrossRef]
- Raghunath, P. Roles of thermostable direct hemolysin (TDH) and TDH-related hemolysin (TRH) in Vibrio parahaemolyticus. Front. Microbiol. 2014, 5, 805. [Google Scholar] [CrossRef]
- Wang, R.; Zhong, Y.; Gu, X.; Yuan, J.; Saeed, A.F.; Wang, S. The pathogenesis, detection, and prevention of Vibrio parahaemolyticus. Front. Microbiol. 2015, 6, 144. [Google Scholar] [CrossRef] [PubMed]
- Okuda, J.; Ishibashi, M.; Hayakawa, E.; Nishino, T.; Takeda, Y.; Mukhopadhyay, A.K.; Garg, S.; Bhattacharya, S.K.; Nair, G.B.; Nishibuchi, M. Emergence of a unique O3:K6 clone of Vibrio parahaemolyticus in Calcutta, India, and isolation of strains from the same clonal group from Southeast Asian travelers arriving in Japan. J. Clin. Microbiol. 1997, 35, 3150–3155. [Google Scholar] [CrossRef] [PubMed]
- Terano, H.; Takahashi, K.; Sakakibara, Y. Characterization of spore germination of a thermoacidophilic spore-forming bacterium, Alicyclobacillus acidoterrestris. Biosci. Biotechnol. Biochem. 2005, 69, 1217–1220. [Google Scholar] [CrossRef]
- Hara-Kudo, Y.; Nishina, T.; Nakagawa, H.; Konuma, H.; Hasegawa, J.; Kumagai, S. Improved method for detection of Vibrio parahaemolyticus in seafood. Appl. Environ. Microbiol. 2001, 67, 5819–5823. [Google Scholar] [CrossRef]
- Food and Drug Agency (FDA). Bacteriological Analytical Manual (BAM) Chapter 9: Vibrio. Available online: https://www.fda.gov/food/laboratory-methods-food/bam-chapter-9-vibrio (accessed on 20 January 2022).
- DePaola, A.; Motes, M.L. Isolation of Vibrio parahaemolyticus from wild raccoons in Florida. In Vibrios in the Environment, 1st ed.; Colwell, R.R., Ed.; John Wiley & Sons, Inc.: New York, NY, USA, 1984; pp. 563–566. [Google Scholar]
- DePaola, A.; Kaysner, C.A.; McPhearson, R.M. Elevated temperature method for recovery of Vibrio cholerae from oysters (Crassostrea gigas). Appl. Environ. Microbiol. 1987, 53, 1181–1182. [Google Scholar] [CrossRef]
- Huq, A.; Grim, C.; Colwell, R.R.; Nair, G.B. Detection, isolation, and identification of Vibrio cholerae from the environment. Curr. Protoc. Microbiol. 2006, 2, 6A.5.1–6A.5.38. [Google Scholar] [CrossRef]
- Alam, M.; Chowdhury, W.B.; Bhuiyan, N.A.; Islam, A.; Hasan, N.A.; Nair, G.B.; Watanabe, H.; Siddique, A.K.; Huq, A.; Sack, R.B.; et al. Serogroup, virulence and genetic traits of Vibrio parahaemolyticus in the estuarine ecosystem of Bangladesh. Appl. Environ. Microbiol. 2009, 75, 6268–6274. [Google Scholar] [CrossRef] [PubMed]
- Oliver, J.D. Recent findings on the viable but nonculturable state in pathogenic bacteria. FEMS Microbiol. Rev. 2010, 34, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Huq, A.; Haley, B.J.; Taviani, E.; Chen, A.; Hasan, N.A.; Colwell, R.R. Detection, isolation, and identification of Vibrio cholerae from the environment. Curr. Protoc. Microbiol. 2012, 26, 6A.5.1–6A.5.51. [Google Scholar] [CrossRef] [PubMed]
- Otomo, Y.; Hossain, F.; Rabbi, F.; Yakuwa, Y.; Ahsan, C.R. Pre-enrichment of estuarine and fresh water environmental samples with sodium chloride yields in better recovery of Vibrio parahaemolyticus. Adv. Microbiol. 2015, 3, 21–25. [Google Scholar] [CrossRef]
- Delmore, R.P., Jr.; Crisley, F.D. Thermal Resistance of Vibrio parahaemolyticus in clam homogenate. J. Food Prot. 1979, 42, 131–134. [Google Scholar] [CrossRef]
- Andrews, L.S.; DeBlanc, S.; Veal, C.D.; Park, D.L. Response of Vibrio parahaemolyticus 03:K6 to a hot water/cold shock pasteurization process. Food Addit. Contam. 2003, 20, 331–334. [Google Scholar] [CrossRef]
- Chen, H.C.; Chang, T.C. Rapid detection of Vibrio parahaemolyticus in oysters by immunofluorescence microscopy. Int. J. Food Microbiol. 1996, 29, 311–319. [Google Scholar] [CrossRef]
- Lipp, E.K.; Rivera, I.N.G.; Gil, A.I.; Espeland, E.M.; Choopun, N.; Louis, V.R.; Russek-Cohen, E.; Huq, A.; Colwell, R.R. Direct Detection of Vibrio cholerae and ctxA in Peruvian coastal water and plankton by PCR. Appl. Environ. Microbiol. 2003, 69, 3676–3680. [Google Scholar] [CrossRef]
- Hossain, M.T.; Kim, Y.O.; Kong, I.S. Multiplex PCR for the detection and differentiation of Vibrio parahaemolyticus strains using the groEL, tdh and trh genes. Mol. Cell Probes 2013, 27, 171–175. [Google Scholar] [CrossRef]
- Blanco-Abad, V.; Ansede-Bermejo, J.; Rodriguez-Castro, A.; Martinez-Urtaza, J. Evaluation of different procedures for the optimized detection of Vibrio parahaemolyticus in mussels and environmental samples. Int. J. Food Microbiol. 2009, 129, 229–236. [Google Scholar] [CrossRef]
- Letchumanan, V.; Chan, K.G.; Lee, L.H. Vibrio parahaemolyticus: A review on the pathogenesis, prevalence, and advance molecular identification techniques. Front. Microbiol. 2014, 5, 705. [Google Scholar] [CrossRef] [PubMed]
- Yamai, S.; Okitsu, T.; Shimada, T.; Katsube, Y. Distribution of serogroups of Vibrio cholerae non-O1 non-O139 with specific reference to their ability to produce cholera toxin, and addition of novel serogroups. J. Jpn. Assoc. Infect. Dis. 1997, 71, 1037–1045. [Google Scholar] [CrossRef]
- Law, J.W.; Ab Mutalib, N.S.; Chan, K.G.; Lee, L.H. Rapid methods for the detection of foodborne bacterial pathogens: Principles, applications, advantages and limitations. Front. Microbiol. 2015, 5, 770. [Google Scholar] [CrossRef] [PubMed]
- Hara-Kudo, Y.; Sugiyama, K.; Nishibuchi, M.; Chowdhury, A.; Yatsuyanagi, J.; Ohtomo, Y.; Saito, A.; Nagano, H.; Nishina, T.; Nakagawa, H.; et al. Prevalence of pandemic thermostable direct hemolysin-producing Vibrio parahaemolyticus O3:K6 in seafood and the coastal environment in Japan. Appl. Environ. Microbiol. 2003, 69, 3883–3891. [Google Scholar] [CrossRef] [PubMed]
- Wong, H.C.; Peng, P.Y.; Lan, S.L.; Chen, Y.C.; Lu, K.H.; Shen, C.T.; Lan, S.F. Effects of heat shock on the thermotolerance, protein composition, and toxin production of Vibrio parahaemolyticus. J. Food Prot. 2002, 65, 499–507. [Google Scholar] [CrossRef] [PubMed]
- Tiong, H.K.; Muriana, P.M. RT-qPCR analysis of 15 genes encoding putative surface proteins involved in adherence of Listeria monocytogenes. Pathogens 2016, 5, 60. [Google Scholar] [CrossRef] [PubMed]
- Tanner, M.A.; Shoskes, D.; Shahed, A.; Pace, N.R. Prevalence of corynebacterial 16S rRNA sequences in patients with bacterial and “nonbacterial” prostatitis. J. Clin. Microbiol. 1999, 37, 1863–1870. [Google Scholar] [CrossRef]
- Ward, L.N.; Bej, A.K. Detection of Vibrio parahaemolyticus in shellfish by use of multiplexed real-time PCR with TaqMan fluorescent probes. Appl. Environ. Microbiol. 2006, 72, 2031–2042. [Google Scholar] [CrossRef]
- Tan, C.W.; Malcolm, T.; Kuan, C.H.; Thung, T.Y.; Chang, W.S.; Loo, Y.Y.; Premarathne, J.; Ramzi, O.B.; Norshafawatie, M.; Yusralimuna, N.; et al. Prevalence and antimicrobial susceptibility of Vibrio parahaemolyticus isolated from short mackerels (Rastrelliger brachysoma) in Malaysia. Front. Microbiol. 2017, 8, 1087. [Google Scholar] [CrossRef]
- CLSI. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria, 3rd ed.; CLSI guideline M45; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2015. [Google Scholar]
- Urmersbach, S.; Aho, T.; Alter, T.; Hassan, S.S.; Autio, R.; Huehn, S. Changes in global gene expression of Vibrio parahaemolyticus induced by cold- and heat-stress. BMC Microbiol. 2015, 15, 229. [Google Scholar] [CrossRef]
- Han, F.; Walker, R.D.; Janes, M.E.; Prinyawiwatkul, W.; Ge, B. Antimicrobial susceptibilities of Vibrio parahaemolyticus and Vibrio vulnificus isolates from Louisiana Gulf and retail raw oysters. Appl. Environ. Microbiol. 2007, 73, 7096–7098. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.C.; Brown, A.M.; Luscombe, G.M.; Wong, S.J.; Mendis, K. Antibiotic use for Vibrio infections: Important insights from surveillance data. BMC Infect. Dis. 2015, 15, 226. [Google Scholar] [CrossRef] [PubMed]
- Ayrapetyan, M.; Williams, T.; Oliver, J.D. Relationship between the Viable but Nonculturable State and Antibiotic Persister Cells. J. Bacteriol. 2018, 200, e00249-18. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Q.; Wang, B.; Wang, J.; Liu, Y.; Fang, X.; Liao, Z. Global proteomic analysis of the resuscitation state of Vibrio parahaemolyticus compared with the normal and viable but non-culturable state. Front. Microbiol. 2019, 10, 1045. [Google Scholar] [CrossRef] [PubMed]
- Ben Kahla-Nakbi, A.; Besbes, A.; Chaieb, K.; Rouabhia, M.; Bakhrouf, A. Survival of Vibrio alginolyticus in seawater and retention of virulence of its starved cells. Mar. Environ. Res. 2007, 64, 469–478. [Google Scholar] [CrossRef]
- Snoussi, M.; Noumi, E.; Cheriaa, J.; Usai, D.; Sechi, L.A.; Zanetti, S.; Bakhrouf, A. Adhesive properties of environmental Vibrio alginolyticus strains to biotic and abiotic surfaces. New Microbiol. 2008, 31, 489–500. [Google Scholar] [PubMed]
- Xu, F.; Ilyas, S.; Hall, J.A.; Jones, S.H.; Cooper, V.S.; Whistler, C.A. Genetic characterization of clinical and environmental Vibrio parahaemolyticus from the Northeast USA reveals emerging resident and non-indigenous pathogen lineages. Front. Microb. 2015, 6, 272. [Google Scholar] [CrossRef]
- Kumar, H.S.; Parvathi, A.; Karunasagar, I.; Karunasagar, I. A gyrB-based PCR for the detection of Vibrio vulnificus and its application for direct detection of this pathogen in oyster enrichment broths. Int. J. Food Microbiol. 2006, 111, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Cebrián, G.; Condón, S.; Mañas, P. Physiology of the inactivation of vegetative bacteria by thermal treatments: Mode of action, influence of environmental factors and inactivation kinetics. Foods 2017, 6, 107. [Google Scholar] [CrossRef]
- Desai, S.V.; Varadaraj, M.C. Behavioural pattern of vegetative cells and spores of Bacillus cereus as affected by time-temperature combinations used in processing of Indian traditional foods. J. Food Sci. Technol. 2010, 47, 549–556. [Google Scholar] [CrossRef][Green Version]
- Bukau, B. Regulation of the Escherichia coli heat-shock response. Mol. Microbiol. 1993, 9, 671–680. [Google Scholar] [CrossRef] [PubMed]
- Russell, A.D. Lethal effects of heat on bacterial physiology and structure. Sci. Prog. 2003, 86 Pt 1–2, 115–137. [Google Scholar] [CrossRef] [PubMed]
- Paracini, N.; Clifton, L.A.; Skoda, M.W.A.; Lakey, J.H. Liquid crystalline bacterial outer membranes are critical for antibiotic susceptibility. Proc. Natl. Acad. Sci. USA 2018, 115, E7587–E7594. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Yamamoto, K.; Honda, T.; Ming, X. Construction and characterization of an isogenic mutant of Vibrio parahaemolyticus having a deletion in the thermostable direct hemolysin-related hemolysin gene (trh). J. Bact. 1994, 176, 4757–4760. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.P.; Zhang, J.L.; Jiang, T.; Bao, Y.X.; Zhou, X.M. Insufficiency of the Kanagawa hemolytic test for detecting pathogenic Vibrio parahaemolyticus in Shanghai, China. Diagn. Microbiol. Infect. Dis. 2011, 69, 7–11. [Google Scholar] [CrossRef]
- Okuda, J.; Ishibashi, M.; Abbott, S.L.; Janda, J.M.; Nishibuchi, M. Analysis of the thermostable direct hemolysin (tdh) gene and the tdh-related hemolysin (trh) genes in urease-positive strains of Vibrio parahaemolyticus isolated on the West Coast of the United States. J. Clin. Microbiol. 1997, 35, 1965–1971. [Google Scholar] [CrossRef]
- Park, K.S.; Iida, T.; Yamaichi, Y.; Oyagi, T.; Yamamoto, K.; Honda, T. Genetic characterization of DNA region containing the trh and ure genes of Vibrio parahaemolyticus. Infect. Immun. 2000, 68, 5742–8748. [Google Scholar] [CrossRef]
- Honda, S.; Matsumoto, S.; Miwatani, T.; Honda, T. A survey of urease-positive Vibrio parahaemolyticus strains isolated from traveller’s diarrhea, sea water and imported frozen sea foods. Eur. J. Epidemiol. 1992, 8, 861–864. [Google Scholar] [CrossRef]
- Huq, M.I.; Huber, D.; Kibryia, G. Isolation of urease producing Vibrio parahaemolyticus strains from cases of gastroenteritis. Indian J. Med. Res. 1979, 70, 549–553. [Google Scholar]
- Suthienkul, O.; Ishibashi, M.; Iida, T.; Nettip, N.; Supavej, S.; Eampokalap, B.; Makino, M.; Honda, T. Urease production correlates with possession of the trh gene in Vibrio parahaemolyticus strains isolated in Thailand. J. Infect. Dis. 1995, 172, 1405–1408. [Google Scholar] [CrossRef]
- Eko, F.O. Urease production in Vibrio parahaemolyticus: A potential marker for virulence. Eur. J. Epidemiol. 1992, 8, 627–628. [Google Scholar] [CrossRef] [PubMed]
- Osawa, R.; Okitsu, T.; Morozumi, H.; Yamai, S. Occurrence of urease-positive Vibrio parahaemolyticus in Kanagawa, Japan, with specific reference to presence of thermostable direct hemolysin (TDH) and the TDH-related-hemolysin genes. Appl. Environ. Microbiol. 1996, 62, 725–727. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.Y.; Na, E.J.; Lee, K.H.; Ryu, S.; Yoon, H.; Lee, J.H.; Kim, H.B.; Kim, H.; Choi, S.H.; Kim, B.S. Complete genome sequence of Vibrio parahaemolyticus FORC_023 isolated from raw fish storage water. Pathog. Dis. 2016, 74, ftw032. [Google Scholar] [CrossRef] [PubMed]
- Elexson, N.; Afsah-Hejri, L.; Rukayadi, Y.; Soopna, P.; Lee, H.Y.; Tuan Zainazor, T.C.; Son, R. Effect of detergents as antibacterial agents on biofilm of antibiotics-resistant Vibrio parahaemolyticus isolates. Food Control 2014, 35, 378–385. [Google Scholar] [CrossRef]
- Shishir, M.A.; Mamun, M.A.; Mian, M.M.; Ferdous, U.T.; Akter, N.J.; Suravi, R.S.; Datta, S.; Kabir, M.E. Prevalence of Vibrio cholerae in coastal alternative supplies of drinking water and association with Bacillus-like spore formers. Front. Public Health 2018, 6, 50. [Google Scholar] [CrossRef]
- Vezzulli, L.; Grande, C.; Reid, P.C.; Hélaouët, P.; Edwards, M.; Höfle, M.G.; Brettar, I.; Colwell, R.R.; Pruzzo, C. Climate influence on Vibrio and associated human diseases during the past half-century in the coastal North Atlantic. Proc. Natl. Acad. Sci. USA 2016, 113, E5062–E5071. [Google Scholar] [CrossRef]
- Ueda, Y.; Yamauchi, S.; Fukata, S.; Okuyama, H.; Morita, E.; Shelake, R.; Hayashi, H. Heterologous expression of thermolabile proteins enhances thermotolerance in Escherichia coli. Adv. Microbiol. 2016, 6, 602–612. [Google Scholar] [CrossRef][Green Version]
- Macaluso, A.; Best, E.A.; Bender, R.A. Role of the nac gene product in the nitrogen regulation of some NTR-regulated operons of Klebsiella aerogenes. J. Bacteriol. 1990, 172, 7249–7255. [Google Scholar] [CrossRef]
- Sissons, C.H.; Perinpanayagam, H.E.R.; Hancock, E.M.; Cutress, T.W. pH regulation of urease levels in Streptoccus salivarius. J. Dent. Res. 1990, 69, 1131–1137. [Google Scholar] [CrossRef]
- Nicholson, E.B.; Concaugh, E.A.; Foxall, P.A.; Island, M.D.; Mobley, H.L.T. Proteus mirabilis urease: Transcriptional regulation by ureR. J. Bacteriol. 1993, 175, 465–473. [Google Scholar] [CrossRef]
- Berutti, T.R.; Williams, R.E.; Shen, S.; Taylor, M.M.; Grimes, D.J. Prevalence of urease in Vibrio parahaemolyticus from the Mississippi Sound. Lett. Appl. Microbiol. 2014, 58, 624–628. [Google Scholar] [CrossRef] [PubMed]
- Blair, J.M.; Webber, M.A.; Baylay, A.J.; Ogbolu, D.O.; Piddock, L.J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 2015, 13, 42–51. [Google Scholar] [CrossRef] [PubMed]
- Miranda, C.D.; Godoy, F.A.; Lee, M.R. Current status of the use of antibiotics and the antimicrobial resistance in the Chilean Salmon Farms. Front. Microbiol. 2018, 9, 1284. [Google Scholar] [CrossRef] [PubMed]
- Tan, C.W.; Rukayadi, Y.; Hasan, H.; Thung, T.Y.; Lee, E.; Rollon, W.D.; Hara, H.; Kayali, A.Y.; Nishibuchi, M.; Radu, S. Prevalence and antibiotic resistance patterns of Vibrio parahaemolyticus isolated from different types of seafood in Selangor, Malaysia. Saudi J. Biol. Sci. 2020, 27, 1602–1608. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).